+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2xkv | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
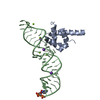
| Title | Atomic Model of the SRP-FtsY Early Conformation | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationendoplasmic reticulum signal peptide binding / signal recognition particle binding / signal recognition particle / signal-recognition-particle GTPase / 7S RNA binding / SRP-dependent cotranslational protein targeting to membrane, translocation / SRP-dependent cotranslational protein targeting to membrane / protein targeting to membrane / ribonucleoprotein complex / GTPase activity ...endoplasmic reticulum signal peptide binding / signal recognition particle binding / signal recognition particle / signal-recognition-particle GTPase / 7S RNA binding / SRP-dependent cotranslational protein targeting to membrane, translocation / SRP-dependent cotranslational protein targeting to membrane / protein targeting to membrane / ribonucleoprotein complex / GTPase activity / GTP binding / ATP hydrolysis activity / plasma membrane / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 13.5 Å | |||||||||
 Authors Authors | Estrozi, L.F. / Boehringer, D. / Shan, S.-o. / Ban, N. / Schaffitzel, C. | |||||||||
 Citation Citation |  Journal: Nat Struct Mol Biol / Year: 2011 Journal: Nat Struct Mol Biol / Year: 2011Title: Cryo-EM structure of the E. coli translating ribosome in complex with SRP and its receptor. Authors: Leandro F Estrozi / Daniel Boehringer / Shu-Ou Shan / Nenad Ban / Christiane Schaffitzel /  Abstract: We report the 'early' conformation of the Escherichia coli signal recognition particle (SRP) and its receptor FtsY bound to the translating ribosome, as determined by cryo-EM. FtsY binds to the ...We report the 'early' conformation of the Escherichia coli signal recognition particle (SRP) and its receptor FtsY bound to the translating ribosome, as determined by cryo-EM. FtsY binds to the tetraloop of the SRP RNA, whereas the NG domains of the SRP protein and FtsY interact weakly in this conformation. Our results suggest that optimal positioning of the SRP RNA tetraloop and the Ffh NG domain leads to FtsY recruitment. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2xkv.cif.gz 2xkv.cif.gz | 172.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2xkv.ent.gz pdb2xkv.ent.gz | 130.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2xkv.json.gz 2xkv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2xkv_validation.pdf.gz 2xkv_validation.pdf.gz | 816.1 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2xkv_full_validation.pdf.gz 2xkv_full_validation.pdf.gz | 826.9 KB | Display | |
| Data in XML |  2xkv_validation.xml.gz 2xkv_validation.xml.gz | 23.6 KB | Display | |
| Data in CIF |  2xkv_validation.cif.gz 2xkv_validation.cif.gz | 36.2 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xk/2xkv https://data.pdbj.org/pub/pdb/validation_reports/xk/2xkv ftp://data.pdbj.org/pub/pdb/validation_reports/xk/2xkv ftp://data.pdbj.org/pub/pdb/validation_reports/xk/2xkv | HTTPS FTP |
-Related structure data
| Related structure data |  1762MC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 32300.371 Da / Num. of mol.: 1 / Fragment: NG DOMAIN, RESIDUES 1-294 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   References: UniProt: O07347, signal-recognition-particle GTPase |
|---|---|
| #2: RNA chain | Mass: 36856.863 Da / Num. of mol.: 1 / Source method: isolated from a natural source Details: ONLY THE PART OF THE 4.5S RNA THAT IS VISIBLE IN THE EM RECONSTRUCTION IS INCLUDED Source: (natural)  |
| #3: Protein | Mass: 8160.762 Da / Num. of mol.: 1 / Fragment: M DOMAIN, RESIDUES 329-430 / Source method: isolated from a natural source Details: ONLY THE PART OF THE M DOMAIN THAT IS VISIBLE IN THE EM RECONSTRUCTION IS INCLUDED Source: (natural)  |
| #4: Protein | Mass: 32972.230 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: RNC-SRP-FTSY / Type: RIBOSOME |
|---|---|
| Buffer solution | pH: 7.5 / Details: 50 mM Hepes-KOH, 100 mM KOAc, 8 mM Mg(OAc)2 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Details: HOLEY CARBON |
| Vitrification | Instrument: HOMEMADE PLUNGER / Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TECNAI F20 |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 200 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD / Nominal magnification: 50000 X / Nominal defocus max: 3500 nm / Nominal defocus min: 1500 nm |
| Image recording | Film or detector model: KODAK SO-163 FILM |
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| EM software |
| |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symmetry | Point symmetry: C1 (asymmetric) | |||||||||||||||||||||
| 3D reconstruction | Resolution: 13.5 Å / Num. of particles: 28822 / Symmetry type: POINT | |||||||||||||||||||||
| Atomic model building | Space: REAL | |||||||||||||||||||||
| Atomic model building |
| |||||||||||||||||||||
| Refinement | Highest resolution: 13.5 Å | |||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 13.5 Å
|
 Movie
Movie Controller
Controller
































 PDBj
PDBj





























