[English] 日本語
 Yorodumi
Yorodumi- EMDB-1024: Six molecules of SV40 large T antigen assemble in a propeller-sha... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-1024 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
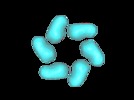
| Title | Six molecules of SV40 large T antigen assemble in a propeller-shaped particle around a channel. | |||||||||
 Map data Map data | SV40 T antigen | |||||||||
 Sample Sample |
| |||||||||
| Function / homology | T antigen, Ori-binding / DNA replication origin binding Function and homology information Function and homology information | |||||||||
| Biological species |  Simian virus 40 Simian virus 40 | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 29.0 Å | |||||||||
 Authors Authors | San Martin C | |||||||||
 Citation Citation |  Journal: J Mol Biol / Year: 1997 Journal: J Mol Biol / Year: 1997Title: Six molecules of SV40 large T antigen assemble in a propeller-shaped particle around a channel. Authors: M C San Martín / C Gruss / J M Carazo /  Abstract: The large T antigen of simian virus 40 (SV40) is a multifunctional regulatory protein, responsible for both the control of viral infection and the required alterations of cellular processes. T ...The large T antigen of simian virus 40 (SV40) is a multifunctional regulatory protein, responsible for both the control of viral infection and the required alterations of cellular processes. T antigen is the only viral protein required for viral DNA replication. It binds specifically to the viral origin and as a helicase unwinds the SV40 DNA bidirectionally. The functional complex is a double hexameric oligomer. In the absence of DNA, but in the presence of ATP or a non-hydrolyzable analog, T antigen assembles into hexamers, which are active as a helicase when a partially single-stranded (3') entry site exists on the substrate. We have used negative staining electron microscopy, single particle image processing and three-dimensional reconstruction with a new algebraic reconstruction techniques (ART) algorithm to study the structure of these hexameric particles in the presence of different nucleotide cofactors (ATP, ADP, and the non-hydrolyzable analogs ATPgammaS and AMP-PNP). In every case a strong 6-fold structure was found, with the six density maxima arranged in a ring-like particle around a channel, and a well-defined vorticity. Because these structural features have recently been found in other prokaryotic helicases, they seem to be strongly related to the activity of the protein, which suggests a general functional model conserved through evolution. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_1024.map.gz emd_1024.map.gz | 443.4 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-1024-v30.xml emd-1024-v30.xml emd-1024.xml emd-1024.xml | 7.7 KB 7.7 KB | Display Display |  EMDB header EMDB header |
| Images |  1024.gif 1024.gif | 9.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-1024 http://ftp.pdbj.org/pub/emdb/structures/EMD-1024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1024 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-1024 | HTTPS FTP |
-Validation report
| Summary document |  emd_1024_validation.pdf.gz emd_1024_validation.pdf.gz | 207.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_1024_full_validation.pdf.gz emd_1024_full_validation.pdf.gz | 206.3 KB | Display | |
| Data in XML |  emd_1024_validation.xml.gz emd_1024_validation.xml.gz | 4.9 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1024 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1024 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1024 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-1024 | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_1024.map.gz / Format: CCP4 / Size: 478.5 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_1024.map.gz / Format: CCP4 / Size: 478.5 KB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | SV40 T antigen | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.8 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : SV40 T antigen
| Entire | Name: SV40 T antigen |
|---|---|
| Components |
|
-Supramolecule #1000: SV40 T antigen
| Supramolecule | Name: SV40 T antigen / type: sample / ID: 1000 / Oligomeric state: homohexamer / Number unique components: 1 |
|---|---|
| Molecular weight | Theoretical: 500 KDa |
-Macromolecule #1: Large T antigen
| Macromolecule | Name: Large T antigen / type: protein_or_peptide / ID: 1 / Name.synonym: T antigen / Number of copies: 6 / Oligomeric state: homohexamer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Simian virus 40 / synonym: SV40 Simian virus 40 / synonym: SV40 |
| Molecular weight | Theoretical: 500 KDa |
| Recombinant expression | Organism: AcNPV |
| Sequence | GO: DNA replication origin binding / InterPro: T antigen, Ori-binding |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | .06 mg/mL |
|---|---|
| Buffer | pH: 7.5 Details: 20 mM Tris.HCl pH 7.5 5 mM KCl 1.5 mM MgCl2 0.1 mM DTT 4 mM ADP |
| Staining | Type: NEGATIVE / Details: 2% Uranyl Acetate |
| Vitrification | Cryogen name: NONE |
- Electron microscopy
Electron microscopy
| Microscope | JEOL 1200EXII |
|---|---|
| Image recording | Category: FILM / Film or detector model: KODAK SO-163 FILM / Digitization - Scanner: EIKONIX IEEE 488 / Number real images: 8 / Average electron dose: 10 e/Å2 / Bits/pixel: 8 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: TUNGSTEN HAIRPIN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 5.6 mm / Nominal magnification: 60000 |
| Sample stage | Specimen holder: Jeol / Specimen holder model: JEOL / Tilt angle max: 55 |
- Image processing
Image processing
| Final reconstruction | Applied symmetry - Point group: C6 (6 fold cyclic) / Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 29.0 Å / Resolution method: OTHER / Software - Name: Xmipp / Number images used: 1472 |
|---|
 Movie
Movie Controller
Controller


 UCSF Chimera
UCSF Chimera