+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2j28 | ||||||
|---|---|---|---|---|---|---|---|
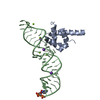
| Title | MODEL OF E. COLI SRP BOUND TO 70S RNCS | ||||||
 Components Components |
| ||||||
 Keywords Keywords | RIBOSOME / PROTEIN-RNA COMPLEX / SIGNAL RECOGNITION PARTICLE | ||||||
| Function / homology |  Function and homology information Function and homology informationabsorption of visible light / G protein-coupled opsin signaling pathway / signal recognition particle / photoreceptor inner segment membrane / 11-cis retinal binding / G protein-coupled photoreceptor activity / signal-recognition-particle GTPase / 7S RNA binding / SRP-dependent cotranslational protein targeting to membrane / protein targeting to membrane ...absorption of visible light / G protein-coupled opsin signaling pathway / signal recognition particle / photoreceptor inner segment membrane / 11-cis retinal binding / G protein-coupled photoreceptor activity / signal-recognition-particle GTPase / 7S RNA binding / SRP-dependent cotranslational protein targeting to membrane / protein targeting to membrane / negative regulation of cytoplasmic translational initiation / photoreceptor outer segment membrane / stringent response / transcriptional attenuation / endoribonuclease inhibitor activity / RNA-binding transcription regulator activity / positive regulation of ribosome biogenesis / negative regulation of cytoplasmic translation / translational termination / DnaA-L2 complex / translation repressor activity / negative regulation of DNA-templated DNA replication initiation / visual perception / mRNA regulatory element binding translation repressor activity / ribosome assembly / assembly of large subunit precursor of preribosome / cytosolic ribosome assembly / ribosomal large subunit assembly / response to reactive oxygen species / translational initiation / regulation of cell growth / DNA-templated transcription termination / response to radiation / mRNA 5'-UTR binding / photoreceptor disc membrane / large ribosomal subunit / ribosome binding / transferase activity / 5S rRNA binding / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / rRNA binding / negative regulation of translation / ribosome / structural constituent of ribosome / translation / ribonucleoprotein complex / response to antibiotic / negative regulation of DNA-templated transcription / GTPase activity / mRNA binding / GTP binding / ATP hydrolysis activity / DNA binding / RNA binding / zinc ion binding / membrane / metal ion binding / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |   | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 9.5 Å | ||||||
 Authors Authors | Halic, M. / Blau, M. / Becker, T. / Mielke, T. / Pool, M.R. / Wild, K. / Sinning, I. / Beckmann, R. | ||||||
 Citation Citation |  Journal: Nature / Year: 2006 Journal: Nature / Year: 2006Title: Following the signal sequence from ribosomal tunnel exit to signal recognition particle. Authors: Mario Halic / Michael Blau / Thomas Becker / Thorsten Mielke / Martin R Pool / Klemens Wild / Irmgard Sinning / Roland Beckmann /  Abstract: Membrane and secretory proteins can be co-translationally inserted into or translocated across the membrane. This process is dependent on signal sequence recognition on the ribosome by the signal ...Membrane and secretory proteins can be co-translationally inserted into or translocated across the membrane. This process is dependent on signal sequence recognition on the ribosome by the signal recognition particle (SRP), which results in targeting of the ribosome-nascent-chain complex to the protein-conducting channel at the membrane. Here we present an ensemble of structures at subnanometre resolution, revealing the signal sequence both at the ribosomal tunnel exit and in the bacterial and eukaryotic ribosome-SRP complexes. Molecular details of signal sequence interaction in both prokaryotic and eukaryotic complexes were obtained by fitting high-resolution molecular models. The signal sequence is presented at the ribosomal tunnel exit in an exposed position ready for accommodation in the hydrophobic groove of the rearranged SRP54 M domain. Upon ribosome binding, the SRP54 NG domain also undergoes a conformational rearrangement, priming it for the subsequent docking reaction with the NG domain of the SRP receptor. These findings provide the structural basis for improving our understanding of the early steps of co-translational protein sorting. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2j28.cif.gz 2j28.cif.gz | 2.2 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2j28.ent.gz pdb2j28.ent.gz | 1.7 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2j28.json.gz 2j28.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2j28_validation.pdf.gz 2j28_validation.pdf.gz | 1.3 MB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2j28_full_validation.pdf.gz 2j28_full_validation.pdf.gz | 2.1 MB | Display | |
| Data in XML |  2j28_validation.xml.gz 2j28_validation.xml.gz | 259.8 KB | Display | |
| Data in CIF |  2j28_validation.cif.gz 2j28_validation.cif.gz | 394.9 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/j2/2j28 https://data.pdbj.org/pub/pdb/validation_reports/j2/2j28 ftp://data.pdbj.org/pub/pdb/validation_reports/j2/2j28 ftp://data.pdbj.org/pub/pdb/validation_reports/j2/2j28 | HTTPS FTP |
-Related structure data
| Related structure data |  1261MC  1262MC  1263MC  1264C  2j37C M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
+50S ribosomal protein ... , 29 types, 29 molecules 01234CDEFGHIJKLMNOPQRSTUVWXYZ
-RNA chain , 3 types, 3 molecules 8AB
| #7: RNA chain | Mass: 23968.311 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #9: RNA chain | Mass: 37848.555 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
| #10: RNA chain | Mass: 941612.375 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
-Protein/peptide / Protein , 2 types, 2 molecules 79
| #6: Protein/peptide | Mass: 2121.542 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|---|
| #8: Protein | Mass: 47362.168 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Strain: K12 / Gene: FAZ83_22145 / Production host:  |
-Non-polymers , 2 types, 623 molecules 


| #35: Chemical | ChemComp-MG / #36: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: SRP BOUND TO 70S RNCS / Type: RIBOSOME |
|---|---|
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Specimen support | Details: CARBON |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Tecnai F30 / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TECNAI F30 |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD |
| Image recording | Film or detector model: KODAK SO-163 FILM |
| Radiation wavelength | Relative weight: 1 |
- Processing
Processing
| EM software | Name: SPIDER / Category: 3D reconstruction | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symmetry | Point symmetry: C1 (asymmetric) | ||||||||||||
| 3D reconstruction | Resolution: 9.5 Å / Resolution method: FSC 0.5 CUT-OFF / Symmetry type: POINT | ||||||||||||
| Refinement | Highest resolution: 9.5 Å | ||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 8 Å
|
 Movie
Movie Controller
Controller


















 PDBj
PDBj






































