+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6379 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Electron Microscopy of Bg505 in complex with 9H3L antibody | |||||||||
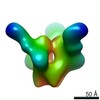
 Map data Map data | Bg505 SOSIP in complex with 9H3L | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  Simian-Human immunodeficiency virus / Simian-Human immunodeficiency virus /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 21.0 Å | |||||||||
 Authors Authors | Garces F / de Val N / Ward AB / Wilson IA | |||||||||
 Citation Citation |  Journal: Immunity / Year: 2015 Journal: Immunity / Year: 2015Title: Affinity Maturation of a Potent Family of HIV Antibodies Is Primarily Focused on Accommodating or Avoiding Glycans. Authors: Fernando Garces / Jeong Hyun Lee / Natalia de Val / Alba Torrents de la Pena / Leopold Kong / Cristina Puchades / Yuanzi Hua / Robyn L Stanfield / Dennis R Burton / John P Moore / Rogier W ...Authors: Fernando Garces / Jeong Hyun Lee / Natalia de Val / Alba Torrents de la Pena / Leopold Kong / Cristina Puchades / Yuanzi Hua / Robyn L Stanfield / Dennis R Burton / John P Moore / Rogier W Sanders / Andrew B Ward / Ian A Wilson /   Abstract: The high-mannose patch on the HIV-1 envelope (Env) glycoprotein is the epicenter for binding of the potent broadly neutralizing PGT121 family of antibodies, but strategies for generating such ...The high-mannose patch on the HIV-1 envelope (Env) glycoprotein is the epicenter for binding of the potent broadly neutralizing PGT121 family of antibodies, but strategies for generating such antibodies by vaccination have not been defined. We generated structures of inferred antibody intermediates by X-ray crystallography and electron microscopy to elucidate the molecular events that occurred during evolution of this family. Binding analyses revealed that affinity maturation was primarily focused on avoiding, accommodating, or binding the N137 glycan. The overall antibody approach angle to Env was defined very early in the maturation process, yet some variation evolved in the PGT121 family branches that led to differences in glycan specificities in their respective epitopes. Furthermore, we determined a crystal structure of the recombinant BG505 SOSIP.664 HIV-1 trimer with a PGT121 family member at 3.0 Å that, in concert with these antibody intermediate structures, provides insights to advance design of HIV vaccine candidates. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6379.map.gz emd_6379.map.gz | 1.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6379-v30.xml emd-6379-v30.xml emd-6379.xml emd-6379.xml | 10.6 KB 10.6 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6379.gif 400_6379.gif 80_6379.gif 80_6379.gif | 15.7 KB 2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6379 http://ftp.pdbj.org/pub/emdb/structures/EMD-6379 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6379 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6379 | HTTPS FTP |
-Validation report
| Summary document |  emd_6379_validation.pdf.gz emd_6379_validation.pdf.gz | 77.9 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_6379_full_validation.pdf.gz emd_6379_full_validation.pdf.gz | 77 KB | Display | |
| Data in XML |  emd_6379_validation.xml.gz emd_6379_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6379 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6379 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6379 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-6379 | HTTPS FTP |
-Related structure data
| Related structure data |  3092C  3093C  6380C  5cexC  5ceyC  5cezC C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6379.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6379.map.gz / Format: CCP4 / Size: 1.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Bg505 SOSIP in complex with 9H3L | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 4.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Bg505 SOSIP in complex with 9H3L
| Entire | Name: Bg505 SOSIP in complex with 9H3L |
|---|---|
| Components |
|
-Supramolecule #1000: Bg505 SOSIP in complex with 9H3L
| Supramolecule | Name: Bg505 SOSIP in complex with 9H3L / type: sample / ID: 1000 / Oligomeric state: one trimer of Bg505 bound to three Fabs / Number unique components: 2 |
|---|---|
| Molecular weight | Experimental: 570 KDa / Theoretical: 570 KDa / Method: Size exclusion chromatography |
-Macromolecule #1: Bg505 SOSIP
| Macromolecule | Name: Bg505 SOSIP / type: protein_or_peptide / ID: 1 / Oligomeric state: trimer / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Simian-Human immunodeficiency virus / synonym: HIV Simian-Human immunodeficiency virus / synonym: HIV |
| Molecular weight | Experimental: 570 KDa / Theoretical: 570 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK293F Homo sapiens (human) / Recombinant cell: HEK293F |
-Macromolecule #2: 9H3L antibody
| Macromolecule | Name: 9H3L antibody / type: protein_or_peptide / ID: 2 / Number of copies: 3 / Recombinant expression: Yes |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) / synonym: human Homo sapiens (human) / synonym: human |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant cell: HEK293F Homo sapiens (human) / Recombinant cell: HEK293F |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.03 mg/mL |
|---|---|
| Buffer | pH: 7.4 / Details: 50 mM Tris-HCl, 150 mM NaCl |
| Staining | Type: NEGATIVE Details: Grids were stained for 30 seconds using 2% uranyl formate. |
| Grid | Details: 400 Cu mesh grids were negatively glow discharged for 30 seconds at 20 mA. |
| Vitrification | Cryogen name: NONE / Instrument: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI SPIRIT |
|---|---|
| Date | May 8, 2015 |
| Image recording | Category: CCD / Film or detector model: TVIPS TEMCAM-F416 (4k x 4k) / Number real images: 126 / Average electron dose: 30.43 e/Å2 |
| Tilt angle min | 0 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Calibrated magnification: 52000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.0 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 46000 |
| Sample stage | Specimen holder model: OTHER / Tilt angle max: 50 |
| Experimental equipment |  Model: Tecnai Spirit / Image courtesy: FEI Company |
- Image processing
Image processing
| Details | The particles were selected and processed using Appion. |
|---|---|
| Final reconstruction | Algorithm: OTHER / Resolution.type: BY AUTHOR / Resolution: 21.0 Å / Resolution method: OTHER / Software - Name: sparx, EMAN2 / Number images used: 16946 |
| Final two d classification | Number classes: 147 |
-Atomic model buiding 1
| Initial model | PDB ID: |
|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera