+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
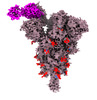
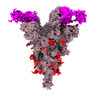
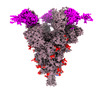
| Title | SARS-CoV-2 Spike (6P) in complex with 3 R1-32 Fabs and 3 ACE2 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords |  ACE2 / ACE2 /  VIRAL PROTEIN-IMMUNE SYSTEM COMPLEX VIRAL PROTEIN-IMMUNE SYSTEM COMPLEX | |||||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of amino acid transport /  angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane /  Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / regulation of systemic arterial blood pressure by renin-angiotensin / tryptophan transport / positive regulation of gap junction assembly / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / regulation of systemic arterial blood pressure by renin-angiotensin / tryptophan transport / positive regulation of gap junction assembly /  regulation of vasoconstriction / regulation of cardiac conduction ...positive regulation of amino acid transport / regulation of vasoconstriction / regulation of cardiac conduction ...positive regulation of amino acid transport /  angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane / angiotensin-converting enzyme 2 / positive regulation of L-proline import across plasma membrane /  Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / regulation of systemic arterial blood pressure by renin-angiotensin / tryptophan transport / positive regulation of gap junction assembly / Hydrolases; Acting on peptide bonds (peptidases); Metallocarboxypeptidases / angiotensin-mediated drinking behavior / regulation of systemic arterial blood pressure by renin-angiotensin / tryptophan transport / positive regulation of gap junction assembly /  regulation of vasoconstriction / regulation of cardiac conduction / peptidyl-dipeptidase activity / angiotensin maturation / maternal process involved in female pregnancy / regulation of vasoconstriction / regulation of cardiac conduction / peptidyl-dipeptidase activity / angiotensin maturation / maternal process involved in female pregnancy /  metallocarboxypeptidase activity / Metabolism of Angiotensinogen to Angiotensins / Attachment and Entry / metallocarboxypeptidase activity / Metabolism of Angiotensinogen to Angiotensins / Attachment and Entry /  carboxypeptidase activity / negative regulation of signaling receptor activity / positive regulation of cardiac muscle contraction / regulation of cytokine production / carboxypeptidase activity / negative regulation of signaling receptor activity / positive regulation of cardiac muscle contraction / regulation of cytokine production /  viral life cycle / blood vessel diameter maintenance / brush border membrane / regulation of transmembrane transporter activity / negative regulation of smooth muscle cell proliferation / viral life cycle / blood vessel diameter maintenance / brush border membrane / regulation of transmembrane transporter activity / negative regulation of smooth muscle cell proliferation /  cilium / negative regulation of ERK1 and ERK2 cascade / endocytic vesicle membrane / cilium / negative regulation of ERK1 and ERK2 cascade / endocytic vesicle membrane /  metallopeptidase activity / positive regulation of reactive oxygen species metabolic process / virus receptor activity / regulation of cell population proliferation / metallopeptidase activity / positive regulation of reactive oxygen species metabolic process / virus receptor activity / regulation of cell population proliferation /  regulation of inflammatory response / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release / regulation of inflammatory response / Maturation of spike protein / viral translation / Translation of Structural Proteins / Virion Assembly and Release /  endopeptidase activity / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / Potential therapeutics for SARS / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry / endopeptidase activity / host cell surface / host extracellular space / suppression by virus of host tetherin activity / Induction of Cell-Cell Fusion / Potential therapeutics for SARS / structural constituent of virion / host cell endoplasmic reticulum-Golgi intermediate compartment membrane / entry receptor-mediated virion attachment to host cell / receptor-mediated endocytosis of virus by host cell / Attachment and Entry /  membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell / membrane fusion / positive regulation of viral entry into host cell / receptor-mediated virion attachment to host cell /  receptor ligand activity / host cell surface receptor binding / symbiont entry into host cell / receptor ligand activity / host cell surface receptor binding / symbiont entry into host cell /  membrane raft / apical plasma membrane / fusion of virus membrane with host plasma membrane / membrane raft / apical plasma membrane / fusion of virus membrane with host plasma membrane /  endoplasmic reticulum lumen / fusion of virus membrane with host endosome membrane / endoplasmic reticulum lumen / fusion of virus membrane with host endosome membrane /  viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane / viral envelope / virion attachment to host cell / SARS-CoV-2 activates/modulates innate and adaptive immune responses / host cell plasma membrane / virion membrane /  cell surface / cell surface /  extracellular space / extracellular exosome / zinc ion binding / extracellular region / extracellular space / extracellular exosome / zinc ion binding / extracellular region /  membrane / identical protein binding / membrane / identical protein binding /  plasma membrane plasma membraneSimilarity search - Function | |||||||||
| Biological species |   Severe acute respiratory syndrome coronavirus 2 / Severe acute respiratory syndrome coronavirus 2 /   Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.73 Å cryo EM / Resolution: 3.73 Å | |||||||||
 Authors Authors | Liu B / Gao X / Li Z / Chen X / He J / Chen L / Xiong X | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2022 Journal: Nat Microbiol / Year: 2022Title: SARS-CoV-2 Delta and Omicron variants evade population antibody response by mutations in a single spike epitope. Authors: Ping He / Banghui Liu / Xijie Gao / Qihong Yan / Rongjuan Pei / Jing Sun / Qiuluan Chen / Ruitian Hou / Zimu Li / Yanjun Zhang / Jincun Zhao / Hao Sun / Bo Feng / Qian Wang / Haisu Yi / ...Authors: Ping He / Banghui Liu / Xijie Gao / Qihong Yan / Rongjuan Pei / Jing Sun / Qiuluan Chen / Ruitian Hou / Zimu Li / Yanjun Zhang / Jincun Zhao / Hao Sun / Bo Feng / Qian Wang / Haisu Yi / Peiyu Hu / Pingchao Li / Yudi Zhang / Zhilong Chen / Xuefeng Niu / Xiaolin Zhong / Liang Jin / Xiaofeng Liu / Kun Qu / Katarzyna A Ciazynska / Andrew P Carter / John A G Briggs / Jizheng Chen / Jinsong Liu / Xinwen Chen / Jun He / Ling Chen / Xiaoli Xiong /     Abstract: Population antibody response is thought to be important in selection of virus variants. We report that SARS-CoV-2 infection elicits a population immune response that is mediated by a lineage of VH1- ...Population antibody response is thought to be important in selection of virus variants. We report that SARS-CoV-2 infection elicits a population immune response that is mediated by a lineage of VH1-69 germline antibodies. A representative antibody R1-32 from this lineage was isolated. By cryo-EM, we show that it targets a semi-cryptic epitope in the spike receptor-binding domain. Binding to this non-ACE2 competing epitope results in spike destruction, thereby inhibiting virus entry. On the basis of epitope location, neutralization mechanism and analysis of antibody binding to spike variants, we propose that recurrent substitutions at 452 and 490 are associated with immune evasion of the identified population antibody response. These substitutions, including L452R (present in the Delta variant), disrupt interactions mediated by the VH1-69-specific hydrophobic HCDR2 to impair antibody-antigen association, enabling variants to escape. The first Omicron variants were sensitive to antibody R1-32 but subvariants that harbour L452R quickly emerged and spread. Our results provide insights into how SARS-CoV-2 variants emerge and evade host immune responses. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_33772.map.gz emd_33772.map.gz | 96.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-33772-v30.xml emd-33772-v30.xml emd-33772.xml emd-33772.xml | 23 KB 23 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_33772_fsc.xml emd_33772_fsc.xml | 13.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_33772.png emd_33772.png | 137.6 KB | ||
| Others |  emd_33772_half_map_1.map.gz emd_33772_half_map_1.map.gz emd_33772_half_map_2.map.gz emd_33772_half_map_2.map.gz | 81 MB 81 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-33772 http://ftp.pdbj.org/pub/emdb/structures/EMD-33772 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33772 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-33772 | HTTPS FTP |
-Related structure data
| Related structure data |  7yegMC  7ydiC  7ydyC  7ye5C  7ye9C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_33772.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_33772.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.408 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #1
| File | emd_33772_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_33772_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
+Entire : SARS-COV-2 Spike_GSAS_6P bound with three R1-32 Fabs and three ACE2
+Supramolecule #1: SARS-COV-2 Spike_GSAS_6P bound with three R1-32 Fabs and three ACE2
+Supramolecule #2: SARS-COV-2 Spike_GSAS_6P
+Supramolecule #3: ACE2
+Supramolecule #4: Fabs
+Macromolecule #1: Spike glycoprotein
+Macromolecule #2: Angiotensin-converting enzyme 2
+Macromolecule #3: heavy chain of R1-32 Fab
+Macromolecule #4: light chain of R1-32 Fab
+Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
+Macromolecule #7: ZINC ION
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 45000 Bright-field microscopy / Nominal defocus max: 2.5 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 45000 |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Number real images: 6989 / Average exposure time: 1.6 sec. / Average electron dose: 63.0 e/Å2 |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z
Z Y
Y X
X




















