+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23708 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
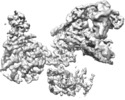
| Title | ATP-bound AMP-activated protein kinase | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationAMPK inhibits chREBP transcriptional activation activity / Lipophagy / import into nucleus /  cAMP-dependent protein kinase regulator activity / protein kinase regulator activity / Energy dependent regulation of mTOR by LKB1-AMPK / Carnitine metabolism / nucleotide-activated protein kinase complex / Activation of PPARGC1A (PGC-1alpha) by phosphorylation / regulation of glycolytic process ...AMPK inhibits chREBP transcriptional activation activity / Lipophagy / import into nucleus / cAMP-dependent protein kinase regulator activity / protein kinase regulator activity / Energy dependent regulation of mTOR by LKB1-AMPK / Carnitine metabolism / nucleotide-activated protein kinase complex / Activation of PPARGC1A (PGC-1alpha) by phosphorylation / regulation of glycolytic process ...AMPK inhibits chREBP transcriptional activation activity / Lipophagy / import into nucleus /  cAMP-dependent protein kinase regulator activity / protein kinase regulator activity / Energy dependent regulation of mTOR by LKB1-AMPK / Carnitine metabolism / nucleotide-activated protein kinase complex / Activation of PPARGC1A (PGC-1alpha) by phosphorylation / regulation of glycolytic process / cAMP-dependent protein kinase regulator activity / protein kinase regulator activity / Energy dependent regulation of mTOR by LKB1-AMPK / Carnitine metabolism / nucleotide-activated protein kinase complex / Activation of PPARGC1A (PGC-1alpha) by phosphorylation / regulation of glycolytic process /  cAMP-dependent protein kinase activity / cAMP-dependent protein kinase activity /  Macroautophagy / AMP binding / cellular response to nutrient levels / carbohydrate transmembrane transporter activity / positive regulation of protein kinase activity / Activation of AMPK downstream of NMDARs / Translocation of SLC2A4 (GLUT4) to the plasma membrane / Macroautophagy / AMP binding / cellular response to nutrient levels / carbohydrate transmembrane transporter activity / positive regulation of protein kinase activity / Activation of AMPK downstream of NMDARs / Translocation of SLC2A4 (GLUT4) to the plasma membrane /  ADP binding / TP53 Regulates Metabolic Genes / fatty acid biosynthetic process / positive regulation of cold-induced thermogenesis / ADP binding / TP53 Regulates Metabolic Genes / fatty acid biosynthetic process / positive regulation of cold-induced thermogenesis /  spermatogenesis / Regulation of TP53 Activity through Phosphorylation / spermatogenesis / Regulation of TP53 Activity through Phosphorylation /  periplasmic space / periplasmic space /  protein kinase activity / protein kinase activity /  protein phosphorylation / positive regulation of gene expression / protein phosphorylation / positive regulation of gene expression /  protein kinase binding / protein kinase binding /  signal transduction / signal transduction /  nucleoplasm / nucleoplasm /  ATP binding / ATP binding /  membrane / membrane /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |   Homo sapiens (human) / Homo sapiens (human) /   Escherichia coli (E. coli) / Escherichia coli (E. coli) /   Lama glama (llama) Lama glama (llama) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.93 Å cryo EM / Resolution: 3.93 Å | |||||||||
 Authors Authors | Yan Y / Mukherjee S / Harikumar KG / Strutzenberg T / Zhou XE / Powell SK / Xu T / Sheldon R / Lamp J / Brunzelle JS ...Yan Y / Mukherjee S / Harikumar KG / Strutzenberg T / Zhou XE / Powell SK / Xu T / Sheldon R / Lamp J / Brunzelle JS / Radziwon K / Ellis A / Novick SJ / Vega IE / Jones R / Miller LJ / Xu HE / Griffin PR / Kossiakoff AA / Melcher K | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2021 Journal: Science / Year: 2021Title: Structure of an AMPK complex in an inactive, ATP-bound state. Authors: Yan Yan / Somnath Mukherjee / Kaleeckal G Harikumar / Timothy S Strutzenberg / X Edward Zhou / Kelly Suino-Powell / Ting-Hai Xu / Ryan D Sheldon / Jared Lamp / Joseph S Brunzelle / Katarzyna ...Authors: Yan Yan / Somnath Mukherjee / Kaleeckal G Harikumar / Timothy S Strutzenberg / X Edward Zhou / Kelly Suino-Powell / Ting-Hai Xu / Ryan D Sheldon / Jared Lamp / Joseph S Brunzelle / Katarzyna Radziwon / Abigail Ellis / Scott J Novick / Irving E Vega / Russell G Jones / Laurence J Miller / H Eric Xu / Patrick R Griffin / Anthony A Kossiakoff / Karsten Melcher /   Abstract: Adenosine monophosphate (AMP)-activated protein kinase (AMPK) regulates metabolism in response to the cellular energy states. Under energy stress, AMP stabilizes the active AMPK conformation, in ...Adenosine monophosphate (AMP)-activated protein kinase (AMPK) regulates metabolism in response to the cellular energy states. Under energy stress, AMP stabilizes the active AMPK conformation, in which the kinase activation loop (AL) is protected from protein phosphatases, thus keeping the AL in its active, phosphorylated state. At low AMP:ATP (adenosine triphosphate) ratios, ATP inhibits AMPK by increasing AL dynamics and accessibility. We developed conformation-specific antibodies to trap ATP-bound AMPK in a fully inactive, dynamic state and determined its structure at 3.5-angstrom resolution using cryo-electron microscopy. A 180° rotation and 100-angstrom displacement of the kinase domain fully exposes the AL. On the basis of the structure and supporting biophysical data, we propose a multistep mechanism explaining how adenine nucleotides and pharmacological agonists modulate AMPK activity by altering AL phosphorylation and accessibility. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23708.map.gz emd_23708.map.gz | 8.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23708-v30.xml emd-23708-v30.xml emd-23708.xml emd-23708.xml | 21.1 KB 21.1 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23708.png emd_23708.png | 99.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23708 http://ftp.pdbj.org/pub/emdb/structures/EMD-23708 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23708 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23708 | HTTPS FTP |
-Related structure data
| Related structure data |  7m74MC  7jhgC  7jhhC  7jijC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23708.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23708.map.gz / Format: CCP4 / Size: 83.7 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.828 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
+Entire : MBP-fused ATP bound AMPK in complex with C-compound stabilized by...
+Supramolecule #1: MBP-fused ATP bound AMPK in complex with C-compound stabilized by...
+Macromolecule #1: 5'-AMP-activated protein kinase catalytic subunit alpha-1
+Macromolecule #2: 5'-AMP-activated protein kinase subunit beta-2
+Macromolecule #3: 5'-AMP-activated protein kinase subunit gamma-1
+Macromolecule #4: Maltose/maltodextrin ABC transporter substrate-binding protein MalE
+Macromolecule #5: Fab light chain
+Macromolecule #6: Fab heavy chain
+Macromolecule #7: Nanobody
+Macromolecule #9: 6-[4-(2-piperidin-1-ylethoxy)phenyl]-3-pyridin-4-ylpyrazolo[1,5-a...
+Macromolecule #10: ADENOSINE-5'-TRIPHOSPHATE
+Macromolecule #11: ADENOSINE-5'-DIPHOSPHATE
+Macromolecule #12: ADENOSINE MONOPHOSPHATE
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 200 / Pretreatment - Type: PLASMA CLEANING |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm Bright-field microscopy / Cs: 2.7 mm |
| Sample stage | Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 66.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 3918314 |
|---|---|
| CTF correction | Software - Name: CTFFIND |
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.93 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 569379 |
 Movie
Movie Controller
Controller