4C75
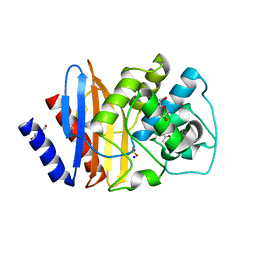 
 | | Consensus (ALL-CON) beta-lactamase class A | | Descriptor: | ACETATE ION, BETA-LACTAMASE, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Gavira, J.A, Risso, V.A, Sanchez-Ruiz, J.M. | | Deposit date: | 2013-09-19 | | Release date: | 2014-04-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Phenotypic Comparisons of Consensus Variants Versus Laboratory Resurrections of Precambrian Proteins.
Proteins, 82, 2014
|
|
7Y0D
 
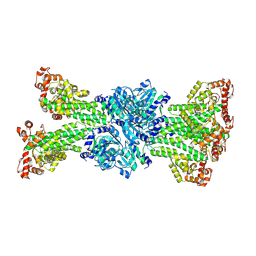 | |
6GW7
 
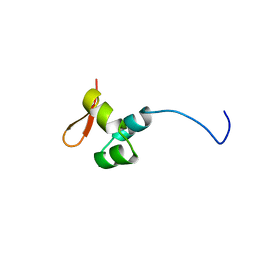 | | The CTD of HpDprA, a DNA binding Winged Helix domain which do not bind dsDNA | | Descriptor: | DNA protecting protein DprA | | Authors: | Lisboa, J, Celma, L, Sanchez, D, Marquis, M, Andreani, J, Guerois, R, Ochsenbein, F, Durand, D, Marsin, S, Cuniasse, P, Radicella, J.P, Quevillon-Cheruel, S. | | Deposit date: | 2018-06-22 | | Release date: | 2019-04-24 | | Last modified: | 2024-06-19 | | Method: | SOLUTION NMR | | Cite: | The C-terminal domain of HpDprA is a DNA-binding winged helix domain that does not bind double-stranded DNA.
Febs J., 286, 2019
|
|
3QBU
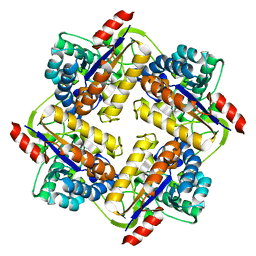 
 | | Crystal structure of putative peptidoglycan deactelyase (HP0310) from Helicobacter pylori | | Descriptor: | Putative uncharacterized protein, ZINC ION | | Authors: | Shaik, M.M, Cendron, L, Percudani, R, Zanotti, G. | | Deposit date: | 2011-01-14 | | Release date: | 2011-05-25 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.5701 Å) | | Cite: | The Structure of Helicobacter pylori HP0310 Reveals an Atypical Peptidoglycan Deacetylase.
Plos One, 6, 2011
|
|
1NYC
 
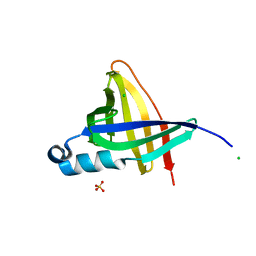 | | Staphostatins resemble lipocalins, not cystatins in fold. | | Descriptor: | CHLORIDE ION, SULFATE ION, cysteine protease inhibitor | | Authors: | Rzychon, M, Filipek, R, Sabat, A, Kosowska, K, Dubin, A, Potempa, J, Bochtler, M. | | Deposit date: | 2003-02-12 | | Release date: | 2003-09-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Staphostatins resemble lipocalins, not cystatins in fold.
Protein Sci., 12, 2003
|
|
4NJP
 
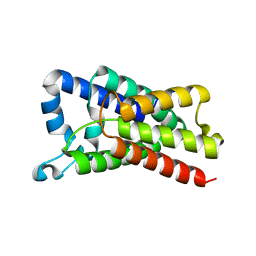 | | Proteolysis inside the membrane is a rate-governed reaction not Driven by substrate affinity | | Descriptor: | Rhomboid protease GlpG | | Authors: | Dickey, S.W, Baker, R.P, Cho, S, Urban, S. | | Deposit date: | 2013-11-11 | | Release date: | 2013-12-25 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Proteolysis inside the Membrane Is a Rate-Governed Reaction Not Driven by Substrate Affinity.
Cell(Cambridge,Mass.), 155, 2013
|
|
1L35
 
 | |
5JME
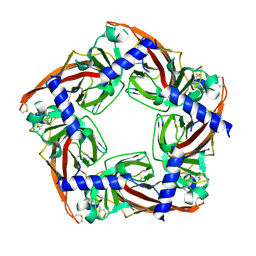 
 | |
1L30
 
 | |
1L28
 
 | |
1L32
 
 | |
1L27
 
 | |
1L25
 
 | |
1L31
 
 | |
3NIG
 
 | | The Closed Headpiece of Integrin IIb 3 and its Complex with an IIb 3 -Specific Antagonist That Does Not Induce Opening | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Zhu, J.H, Zhu, J.Q, Springer, T.A. | | Deposit date: | 2010-06-15 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Closed headpiece of integrin {alpha}IIb{beta}3 and its complex with an {alpha}IIb{beta}3-specific antagonist that does not induce opening.
Blood, 116, 2010
|
|
3NIF
 
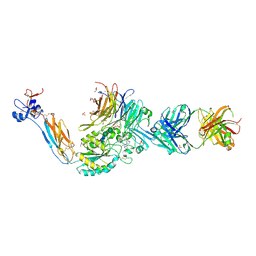 | | The Closed Headpiece of Integrin IIb 3 and its Complex with an IIb 3 -Specific Antagonist That Does Not Induce Opening | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2-ethyl-7-piperazin-1-yl-5H-[1,3,4]thiadiazolo[3,2-a]pyrimidin-5-one, ... | | Authors: | Zhu, J.H, Zhu, J.Q, Springer, T.A. | | Deposit date: | 2010-06-15 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Closed headpiece of integrin {alpha}IIb{beta}3 and its complex with an {alpha}IIb{beta}3-specific antagonist that does not induce opening.
Blood, 116, 2010
|
|
1L26
 
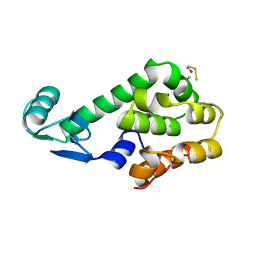 | |
5Z5O
 
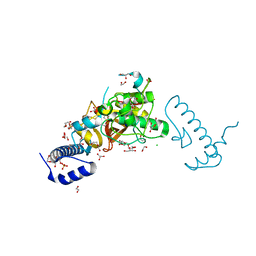 | | Structure of Pycnonodysostosis disease related I249T mutant of human cathepsin K | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, 1,2-ETHANEDIOL, CHLORIDE ION, ... | | Authors: | Biswas, S, Roy, S. | | Deposit date: | 2018-01-19 | | Release date: | 2018-09-26 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | Not all pycnodysostosis-related mutants of human cathepsin K are inactive - crystal structure and biochemical studies of an active mutant I249T.
FEBS J., 285, 2018
|
|
1L29
 
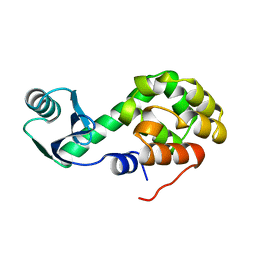 | |
3NID
 
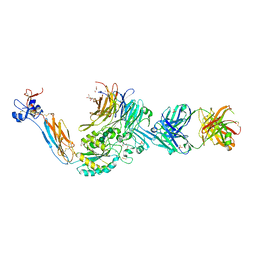 | | The Closed Headpiece of Integrin alphaIIB beta3 and its Complex with an alpahIIB beta3 -Specific Antagonist That Does Not Induce Opening | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, CALCIUM ION, ... | | Authors: | Zhu, J.H, Zhu, J.Q, Springer, T.A. | | Deposit date: | 2010-06-15 | | Release date: | 2010-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Closed headpiece of integrin {alpha}IIb{beta}3 and its complex with an {alpha}IIb{beta}3-specific antagonist that does not induce opening.
Blood, 116, 2010
|
|
1JN0
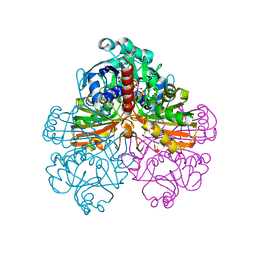 
 | | Crystal structure of the non-regulatory A4 isoform of spinach chloroplast glyceraldehyde-3-phosphate dehydrogenase complexed with NADP | | Descriptor: | GLYCERALDEHYDE-3-PHOSPHATE DEHYDROGENASE A, NADPH DIHYDRO-NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SULFATE ION | | Authors: | Fermani, S, Ripamonti, A, Sabatino, P, Zanotti, G, Scagliarini, S, Sparla, F, Trost, P, Pupillo, P. | | Deposit date: | 2001-07-21 | | Release date: | 2001-11-30 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Crystal structure of the non-regulatory A(4 )isoform of spinach chloroplast glyceraldehyde-3-phosphate dehydrogenase complexed with NADP.
J.Mol.Biol., 314, 2001
|
|
5BJ4
 
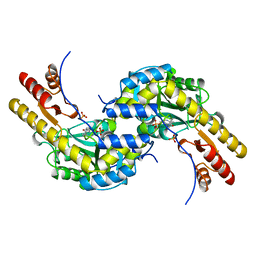 | | THERMUS THERMOPHILUS ASPARTATE AMINOTRANSFERASE TETRA MUTANT 2 | | Descriptor: | PHOSPHATE ION, PROTEIN (ASPARTATE AMINOTRANSFERASE), PYRIDOXAL-5'-PHOSPHATE | | Authors: | Ura, H, Nakai, T, Kawaguchi, S.I, Miyahara, I, Hirotsu, K, Kuramitsu, S. | | Deposit date: | 1999-01-11 | | Release date: | 2003-09-02 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Substrate recognition mechanism of thermophilic dual-substrate enzyme
J.BIOCHEM.(TOKYO), 130, 2001
|
|
6NBS
 
 | | WT ERK2 with compound 2507-8 | | Descriptor: | (5S)-5-benzyl-4,5-dihydro-1H-imidazol-2-amine, GLYCEROL, Mitogen-activated protein kinase 1, ... | | Authors: | Sammons, R.M, Perry, N.A, Cho, E.J, Kaoud, T.S, Zamora-Olivares, D.P, Piserchio, A, Houghten, R.A, Giulianotti, M, Li, Y, Debevec, G, Gurevich, V.V, Ghose, R, Iverson, T.M, Dalby, K.N. | | Deposit date: | 2018-12-10 | | Release date: | 2019-07-31 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | A Novel Class of Common Docking Domain Inhibitors That Prevent ERK2 Activation and Substrate Phosphorylation.
Acs Chem.Biol., 14, 2019
|
|
8ETQ
 
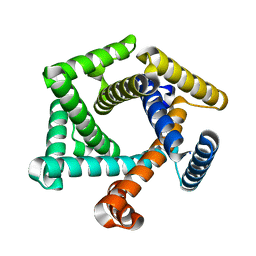 | |
6U1R
 
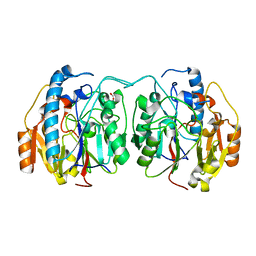 | | SxtG an amidinotransferase from the Microseira wollei in Saxitoxin biosynthetic pathway | | Descriptor: | FORMIC ACID, SxtG | | Authors: | Mallik, L, Lukowski, A.L, Narayan, A.R.H, Koutmos, M. | | Deposit date: | 2019-08-16 | | Release date: | 2020-03-18 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.792 Å) | | Cite: | Substrate Promiscuity of a Paralytic Shellfish Toxin Amidinotransferase.
Acs Chem.Biol., 15, 2020
|
|
