5TWS
 
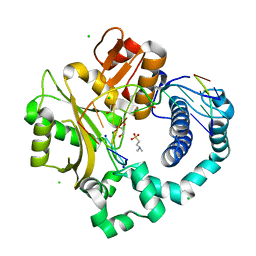 | | Post-catalytic complex of human Polymerase Mu (H329A) with newly incorporated UTP | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CHLORIDE ION, ... | | Authors: | Moon, A.F, Pryor, J.M, Ramsden, D.A, Kunkel, T.A, Bebenek, K, Pedersen, L.C. | | Deposit date: | 2016-11-14 | | Release date: | 2017-07-05 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Structural accommodation of ribonucleotide incorporation by the DNA repair enzyme polymerase Mu.
Nucleic Acids Res., 45, 2017
|
|
3LRN
 
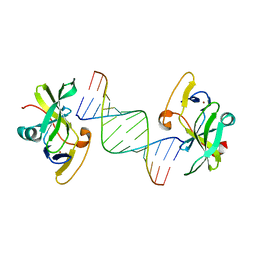 | | Crystal structure of human RIG-I CTD bound to a 14 bp GC 5' ppp dsRNA | | Descriptor: | Probable ATP-dependent RNA helicase DDX58, RNA (5'-R(*(GTP)P*GP*CP*GP*CP*GP*CP*GP*CP*GP*CP*GP*CP*C)-3'), ZINC ION | | Authors: | Li, P. | | Deposit date: | 2010-02-11 | | Release date: | 2010-06-02 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | The Structural Basis of 5' Triphosphate Double-Stranded RNA Recognition by RIG-I C-Terminal Domain.
Structure, 18, 2010
|
|
5TZV
 
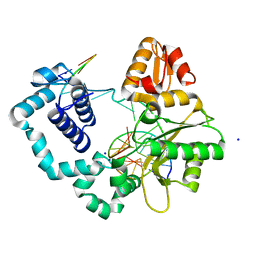 | |
5TWR
 
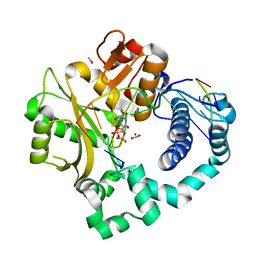 | | Pre-catalytic ternary complex of human Polymerase Mu (H329A) mutant with incoming nonhydrolyzable UMPNPP | | Descriptor: | 1,2-ETHANEDIOL, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, 5'-O-[(S)-hydroxy{[(S)-hydroxy(phosphonooxy)phosphoryl]amino}phosphoryl]uridine, ... | | Authors: | Moon, A.F, Pryor, J.M, Ramsden, D.A, Kunkel, T.A, Bebenek, K, Pedersen, L.C. | | Deposit date: | 2016-11-14 | | Release date: | 2017-07-05 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural accommodation of ribonucleotide incorporation by the DNA repair enzyme polymerase Mu.
Nucleic Acids Res., 45, 2017
|
|
4QDW
 
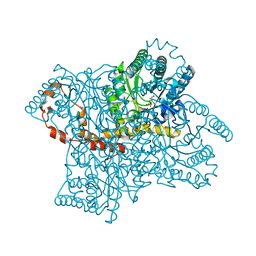 | | Joint X-ray and neutron structure of Streptomyces rubiginosus D-xylose isomerase in complex with two Ni2+ ions and linear L-arabinose | | Descriptor: | L-arabinose, NICKEL (II) ION, Xylose isomerase | | Authors: | Kovalevsky, A.Y, Langan, P. | | Deposit date: | 2014-05-14 | | Release date: | 2014-09-03 | | Last modified: | 2024-02-28 | | Method: | NEUTRON DIFFRACTION (1.8 Å), X-RAY DIFFRACTION | | Cite: | L-Arabinose Binding, Isomerization, and Epimerization by D-Xylose Isomerase: X-Ray/Neutron Crystallographic and Molecular Simulation Study.
Structure, 22, 2014
|
|
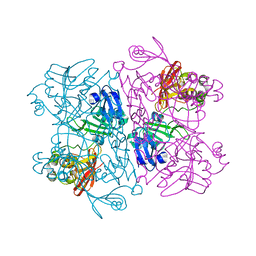
8D34
 
 | |
8D3S
 
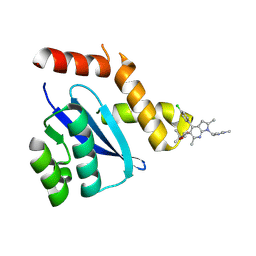 | |
8DEI
 
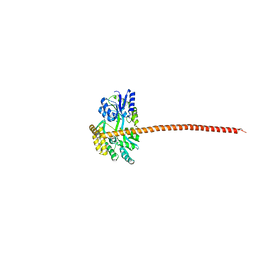 | | Structure of the Cac1 KER domain | | Descriptor: | DI(HYDROXYETHYL)ETHER, GLYCEROL, Maltodextrin-binding protein,Chromatin assembly factor 1 subunit p90 fusion, ... | | Authors: | Rosas, R, Churchill, M.E.A. | | Deposit date: | 2022-06-20 | | Release date: | 2023-07-05 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.81 Å) | | Cite: | A novel single alpha-helix DNA-binding domain in CAF-1 promotes gene silencing and DNA damage survival through tetrasome-length DNA selectivity and spacer function.
Elife, 12, 2023
|
|
3M1D
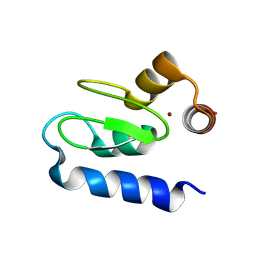 
 | | Structure of BIR1 from cIAP1 | | Descriptor: | Baculoviral IAP repeat-containing protein 2, ZINC ION | | Authors: | Mace, P.D, Day, C.L. | | Deposit date: | 2010-03-04 | | Release date: | 2010-05-26 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Asymmetric recruitment of cIAPs by TRAF2
J.Mol.Biol., 400, 2010
|
|
3M0K
 
 | |
3MFQ
 
 | | A Glance into the Metal Binding Specificity of TroA: Where Elaborate Behaviors Occur in the Active Center | | Descriptor: | High-affinity zinc uptake system protein znuA, ZINC ION | | Authors: | Gao, G.F, Zheng, B, Zhang, Q, Gao, J, Han, H, Li, M. | | Deposit date: | 2010-04-03 | | Release date: | 2011-04-13 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.598 Å) | | Cite: | Insight into the interaction of metal ions with TroA from Streptococcus suis
Plos One, 6, 2011
|
|
4Q7E
 
 | | Non-phosphorylated HemR Receiver Domain from Leptospira biflexa | | Descriptor: | GLYCEROL, Response regulator of a two component regulatory system, SULFATE ION | | Authors: | Morero, N.R, Buschiazzo, A. | | Deposit date: | 2014-04-24 | | Release date: | 2014-05-14 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.441 Å) | | Cite: | HemR is an OmpR/PhoB-like response regulator from Leptospira, which simultaneously effects transcriptional activation and repression of key haem metabolism genes.
Mol.Microbiol., 94, 2014
|
|
3MGT
 
 | | Crystal structure of a H5-specific CTL epitope variant derived from H5N1 influenza virus in complex with HLA-A*0201 | | Descriptor: | 10-meric peptide from Hemagglutinin, Beta-2-microglobulin, HLA class I histocompatibility antigen, ... | | Authors: | Sun, Y, Liu, J, Yang, M, Gao, F, Zhou, J, Kitamura, Y. | | Deposit date: | 2010-04-07 | | Release date: | 2010-05-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.197 Å) | | Cite: | Identification and structural definition of H5-specific CTL epitopes restricted by HLA-A*0201 derived from the H5N1 subtype of influenza A viruses
J.Gen.Virol., 91, 2010
|
|
7F84
 
 | | Crystal structure of CRISPR-associated Cas2c of Leptospira interrogans | | Descriptor: | CRISPR-associated endoribonuclease Cas2, GLYCEROL | | Authors: | Gogoi, P, Anand, V, Prabhakaran, H.S, Kumar, M, Kanaujia, S.P. | | Deposit date: | 2021-07-01 | | Release date: | 2022-07-06 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural and functional characterization of Cas2 of CRISPR-Cas subtype I-C lacking the CRISPR component.
Front Mol Biosci, 9, 2022
|
|
8CQH
 
 | | Ntaya virus methyltransferase in complex with GTP and SAH | | Descriptor: | GUANOSINE-5'-TRIPHOSPHATE, Genome polyprotein, MAGNESIUM ION, ... | | Authors: | Boura, E, Krejcova, K. | | Deposit date: | 2023-03-06 | | Release date: | 2024-03-13 | | Last modified: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structural and functional insights in flavivirus NS5 proteins gained by the structure of Ntaya virus polymerase and methyltransferase.
Structure, 32, 2024
|
|
4Q2K
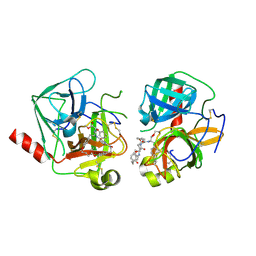 
 | | Bovine alpha chymotrypsin bound to a cyclic peptide inhibitor, 5b | | Descriptor: | (11S)-4,9-dioxo-N-[(2S)-1-oxo-3-phenylpropan-2-yl]-17,22-dioxa-10,30-diazatetracyclo[21.2.2.2~13,16~.1~5,8~]triaconta-1(25),5,7,13,15,23,26,28-octaene-11-carboxamide, Chymotrypsinogen A | | Authors: | Chan, H.Y, Bruning, J.B, Abell, A.D. | | Deposit date: | 2014-04-09 | | Release date: | 2014-07-23 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Macrocyclic protease inhibitors with reduced peptide character.
Angew.Chem.Int.Ed.Engl., 53, 2014
|
|
3MP4
 
 | | Crystal structure of Human lyase R41M mutant | | Descriptor: | Hydroxymethylglutaryl-CoA lyase | | Authors: | Fu, Z, Runquist, J.A, Montgomery, C, Miziorko, H.M, Kim, J.-J.P. | | Deposit date: | 2010-04-24 | | Release date: | 2010-06-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Functional insights into human HMG-CoA lyase from structures of Acyl-CoA-containing ternary complexes.
J.Biol.Chem., 285, 2010
|
|
3M0J
 
 | | Structure of oxaloacetate acetylhydrolase in complex with the inhibitor 3,3-difluorooxalacetate | | Descriptor: | 2,2-difluoro-3,3-dihydroxybutanedioic acid, CALCIUM ION, MANGANESE (II) ION, ... | | Authors: | Herzberg, O, Chen, C. | | Deposit date: | 2010-03-03 | | Release date: | 2010-06-16 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Structure of oxalacetate acetylhydrolase, a virulence factor of the chestnut blight fungus.
J.Biol.Chem., 285, 2010
|
|
3MDA
 
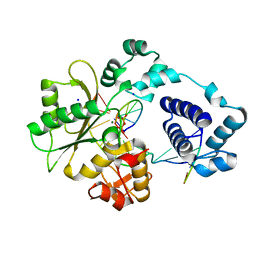 | | DNA polymerase lambda in complex with araC | | Descriptor: | DNA (5'-D(*CP*AP*GP*TP*AP*CP*(CAR))-3'), DNA (5'-D(*CP*GP*GP*CP*GP*GP*TP*AP*CP*TP*G)-3'), DNA (5'-D(P*GP*CP*CP*G)-3'), ... | | Authors: | Garcia-Diaz, M, Murray, M, Kunkel, T, Chou, K.M. | | Deposit date: | 2010-03-30 | | Release date: | 2010-04-28 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.031 Å) | | Cite: | Interaction between DNA Polymerase lambda and anticancer nucleoside analogs.
J.Biol.Chem., 285, 2010
|
|
3MGI
 
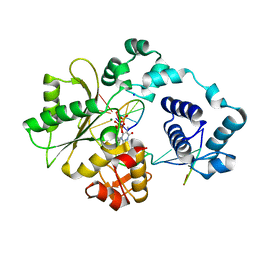 | | Ternary complex of a DNA polymerase lambda loop mutant | | Descriptor: | 2',3'-DIDEOXY-THYMIDINE-5'-TRIPHOSPHATE, DNA, DNA (5'-D(*CP*AP*GP*TP*AP*T)-3'), ... | | Authors: | Garcia-Diaz, M, Bebenek, K, Zhou, R.Z, Povirk, L.F, Kunkel, T. | | Deposit date: | 2010-04-06 | | Release date: | 2010-05-19 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.603 Å) | | Cite: | Loop 1 modulates the fidelity of DNA polymerase lambda
Nucleic Acids Res., 38, 2010
|
|
3MGO
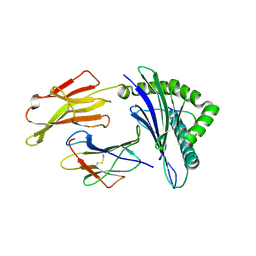 
 | | Crystal structure of a H5-specific CTL epitope derived from H5N1 influenza virus in complex with HLA-A*0201 | | Descriptor: | 10-meric peptide from Hemagglutinin, Beta-2-microglobulin, HLA class I histocompatibility antigen, ... | | Authors: | Sun, Y, Liu, J, Yang, M, Gao, F, Zhou, J, Kitamura, Y. | | Deposit date: | 2010-04-07 | | Release date: | 2010-05-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.297 Å) | | Cite: | Identification and structural definition of H5-specific CTL epitopes restricted by HLA-A*0201 derived from the H5N1 subtype of influenza A viruses
J.Gen.Virol., 91, 2010
|
|
3FHP
 
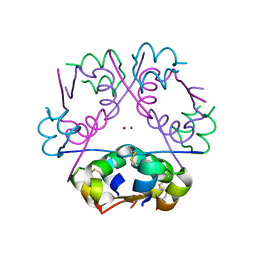 | | A neutron crystallographic analysis of a porcine 2Zn insulin at 2.0 A resolution | | Descriptor: | Insulin, ZINC ION | | Authors: | Iwai, W, Kurihara, K, Yamada, T, Kobayashi, Y, Ohnishi, Y, Tanaka, I, Takahashi, H, Niimura, N. | | Deposit date: | 2008-12-09 | | Release date: | 2009-10-20 | | Last modified: | 2023-11-01 | | Method: | NEUTRON DIFFRACTION (2 Å) | | Cite: | A neutron crystallographic analysis of T6 porcine insulin at 2.1 A resolution
Acta Crystallogr.,Sect.D, 65, 2009
|
|
4QHU
 
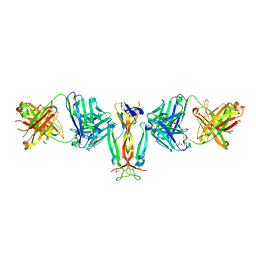 | | Crystal Structure of IL-17A/Fab6785 complex | | Descriptor: | CHLORIDE ION, Fab6785 heavy chain, Fab6785 light chain, ... | | Authors: | Luo, J, Gilliland, G.L, Malia, T, Obmolova, G, Teplyakov, A. | | Deposit date: | 2014-05-29 | | Release date: | 2015-08-05 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | IL-17A Asymmetry in a Complex with a Neutralizing Antibody Fab6785 Reveals a Potential Mechanism of Receptor Differentiation
To be Published
|
|
8E19
 
 | | Crystal structure of TnmK1 complexed with TNM H | | Descriptor: | (1R,8S,13S)-8-[(4-hydroxy-9,10-dioxo-9,10-dihydroanthracen-1-yl)amino]-12-methoxy-10-methylbicyclo[7.3.1]trideca-9,11-diene-2,6-diyne-13-carbaldehyde, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, SUCCINIC ACID, ... | | Authors: | Liu, Y.-C, Gui, C, Shen, B. | | Deposit date: | 2022-08-10 | | Release date: | 2022-11-09 | | Last modified: | 2023-10-18 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | Intramolecular C-C Bond Formation Links Anthraquinone and Enediyne Scaffolds in Tiancimycin Biosynthesis.
J.Am.Chem.Soc., 144, 2022
|
|
8E4R
 
 | |
