9IYF
 
 | | Structure of Phosphopantetheine adenylyltransferase (PPAT) from Enterobacter spp. with the expression tag bound in the substrate binding site of a neighbouring molecule at 2.37 A resolution. | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, PHOSPHONOACETIC ACID, ... | | Authors: | Ahmad, N, Sharma, P, Sharma, S, Singh, T.P. | | Deposit date: | 2024-07-30 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Structure of Phosphopantetheine adenylyltransferase (PPAT) from Enterobacter spp. with the expression tag bound in the substrate binding site of a neighbouring molecule at 2.37 A resolution.
To Be Published
|
|
9IYE
 
 | | Structure of Phosphopantetheine adenylyltransferase (PPAT) from Enterobacter spp. with the expression tag bound in the substrate binding site of a neighbouring molecule at 2.39 A resolution. | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, PHOSPHONOACETIC ACID, ... | | Authors: | Ahmad, N, Sharma, P, Sharma, S, Singh, T.P. | | Deposit date: | 2024-07-30 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | Structure of Phosphopantetheine adenylyltransferase (PPAT) from Enterobacter spp. with the expression tag bound in the substrate binding site of a neighbouring molecule at 2.39 A resolution.
To Be Published
|
|
9IY2
 
 | | Immune complex of HEV-E2s, nAb 8C11 and nAb 8H3 | | Descriptor: | Heavy Chain of mAb 8C11, Heavy Chain of mAb 8H3, Light Chain of mAb 8C11, ... | | Authors: | Minghua, Z, Lizhi, Z, Ying, G, Shaowei, L. | | Deposit date: | 2024-07-29 | | Release date: | 2024-08-28 | | Last modified: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (3.476 Å) | | Cite: | Structural basis for the synergetic neutralization of hepatitis E virus by antibody-antibody interaction
To Be Published
|
|
9IY0
 
 | | anti-HEV mAb 8H3 | | Descriptor: | Heavy chain of 8H3 Fab, Light chain of 8H3 Fab, MAGNESIUM ION | | Authors: | Minghua, Z, Lizhi, Z, Yang, H, Ying, G, Shaowei, L. | | Deposit date: | 2024-07-29 | | Release date: | 2024-09-04 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Structural basis for the synergetic neutralization of hepatitis E virus by antibody-antibody interaction
To Be Published
|
|
9IWQ
 
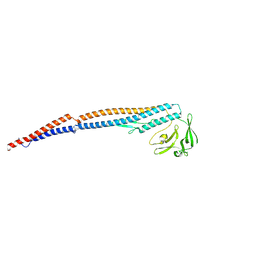 | | Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament | | Descriptor: | Flagellin | | Authors: | Waraich, K, Makino, F, Miyata, T, Kinoshita, M, Minamino, T, Namba, K. | | Deposit date: | 2024-07-25 | | Release date: | 2024-08-07 | | Method: | ELECTRON MICROSCOPY (2.08 Å) | | Cite: | Salmonella enterica serovar Typhimurium FliC(G426A)delta(204-292) forming the L-type straight filament
To Be Published
|
|
9IUY
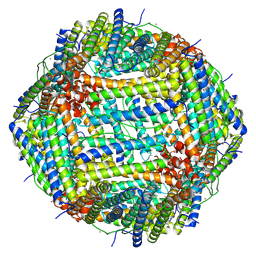 
 | |
9IUM
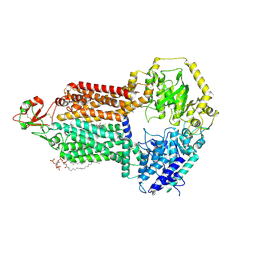 
 | |
9IUL
 
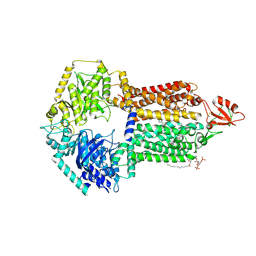 | | The structure of Candida albicans Cdr1 in fluconazole-bound state | | Descriptor: | 2-(2,4-DIFLUOROPHENYL)-1,3-DI(1H-1,2,4-TRIAZOL-1-YL)PROPAN-2-OL, Pip2(20:4/18:0), Pleiotropic ABC efflux transporter of multiple drugs CDR1 | | Authors: | Peng, Y, Sun, H, Yan, Z.F. | | Deposit date: | 2024-07-23 | | Release date: | 2024-09-18 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | Cryo-EM structures of Candida albicans Cdr1 reveal azole-substrate recognition and inhibitor blocking mechanisms.
Nat Commun, 15, 2024
|
|
9IUK
 
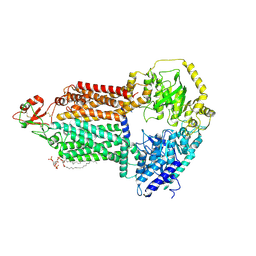 | | The structure of Candida albicans Cdr1 in apo state | | Descriptor: | Pip2(20:4/18:0), Pleiotropic ABC efflux transporter of multiple drugs CDR1 | | Authors: | Peng, Y, Sun, H, Yan, Z.F. | | Deposit date: | 2024-07-22 | | Release date: | 2024-09-18 | | Method: | ELECTRON MICROSCOPY (3.38 Å) | | Cite: | Cryo-EM structures of Candida albicans Cdr1 reveal azole-substrate recognition and inhibitor blocking mechanisms.
Nat Commun, 15, 2024
|
|
9IUH
 
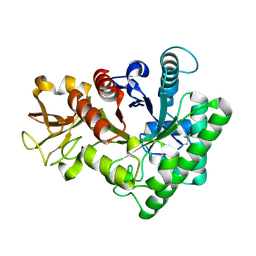 | |
9IT8
 
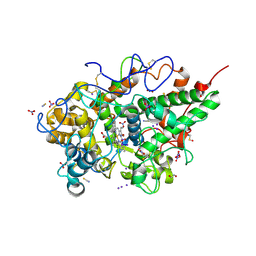 | | Crystal structure of the ternary complex of lactoperoxidase with nitric oxide and nitrite ion at 1.95 A resolution | | Descriptor: | 1,2-ETHANEDIOL, 1-(OXIDOSULFANYL)METHANAMINE, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Maurya, A, Ahmad, N, Sharma, P, Sharma, S, Singh, T.P. | | Deposit date: | 2024-07-19 | | Release date: | 2024-09-11 | | Method: | X-RAY DIFFRACTION (1.954 Å) | | Cite: | Crystal structure of the ternary complex of lactoperoxidase with nitric oxide and nitrite ion at 1.95 A resolution
To Be Published
|
|
9IT1
 
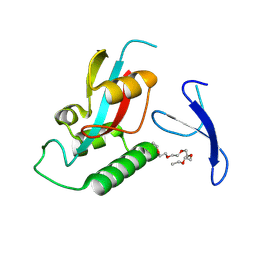 | | Crystal structure of Pin1 using laue diffraction | | Descriptor: | 3,6,9,12,15-PENTAOXAHEPTADECANE, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1 | | Authors: | Sun, B, Qi, Q, Xiao, Q.J, Wang, Z.J. | | Deposit date: | 2024-07-19 | | Release date: | 2024-08-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of Prolyl Isomerase NIMA-interacting 1 (Pin1) using laue diffraction
To Be Published
|
|
9IQ9
 
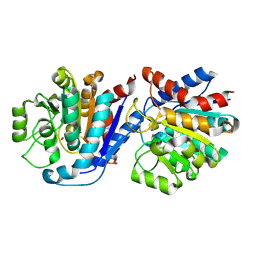 | |
9IOZ
 
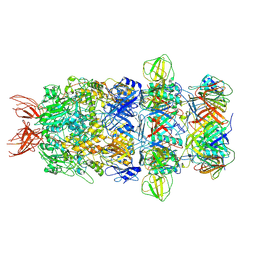 | | Structure of the bacteriophage T5 tail tip complex | | Descriptor: | Baseplate hub protein pb3, Baseplate tube protein p140, Distal tail protein pb9 | | Authors: | Peng, Y.N, Liu, H.R. | | Deposit date: | 2024-07-10 | | Release date: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (3.9 Å) | | Cite: | Structures of Mature and Urea-Treated Empty Bacteriophage T5: Insights into Siphophage Infection and DNA Ejection.
Int J Mol Sci, 25, 2024
|
|
9INY
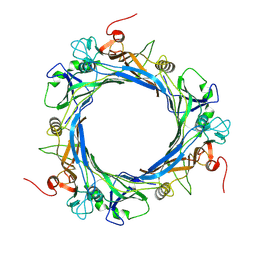 
 | | Structure of bacteriophage T5 tail tube | | Descriptor: | Tail tube protein pb6 | | Authors: | Peng, Y.N, Liu, H.R. | | Deposit date: | 2024-07-08 | | Release date: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (3.6 Å) | | Cite: | Structures of Mature and Urea-Treated Empty Bacteriophage T5: Insights into Siphophage Infection and DNA Ejection.
Int J Mol Sci, 25, 2024
|
|
9INT
 
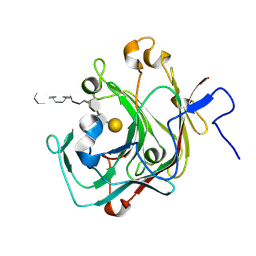 | | Crystal structure of the complex of the beta,kappa-carrageenase Cgbk16A from Wenyingzhuangia fucanilytica with an oligosaccharide of furcellaran | | Descriptor: | 3,6-anhydro-alpha-D-galactopyranose-(1-3)-4-O-sulfo-beta-D-galactopyranose-(1-4)-3,6-anhydro-alpha-D-galactopyranose-(1-3)-4-O-sulfo-beta-D-galactopyranose-(1-4)-3,6-anhydro-alpha-D-galactopyranose-(1-3)-beta-D-galactopyranose, GH16 domain-containing protein | | Authors: | Chang, Y, Chen, F. | | Deposit date: | 2024-07-08 | | Release date: | 2024-07-17 | | Last modified: | 2024-09-25 | | Method: | X-RAY DIFFRACTION (1.56 Å) | | Cite: | Structural Insights into the Substrate Recognition and Catalytic Mechanism of a GH16 beta kappa-Carrageenase from Wenyingzhuangia fucanilytica.
J.Agric.Food Chem., 72, 2024
|
|
9INS
 
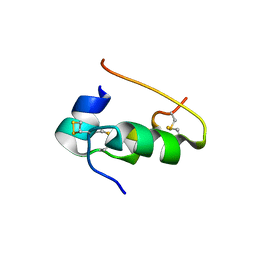 | |
9INR
 
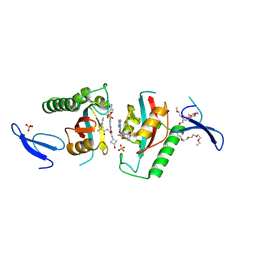 | | Crystal structure of PIN1 in complex with inhibitor C3 | | Descriptor: | 3,6,9,12,15,18,21,24,27,30,33,36,39-TRIDECAOXAHENTETRACONTANE-1,41-DIOL, Peptidyl-prolyl cis-trans isomerase NIMA-interacting 1, SULFATE ION, ... | | Authors: | Zhang, L.Y. | | Deposit date: | 2024-07-08 | | Release date: | 2024-09-25 | | Method: | X-RAY DIFFRACTION (1.93 Å) | | Cite: | Re-Evaluating PIN1 as a Therapeutic Target in Oncology Using Neutral Inhibitors and PROTACs.
J.Med.Chem., 67, 2024
|
|
9INK
 
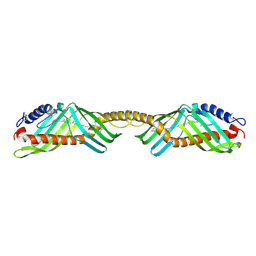 | | Crystal structure of beta-carotene-binding protein (BBP) from Schistocerca gregaria complexed with beta-carotene | | Descriptor: | BETA-CAROTENE, Yellow protein of the takeout family | | Authors: | Boyko, K.M, Varfolomeeva, L.A, Egorkin, N.A, Popov, V.O, Sluchanko, N.N. | | Deposit date: | 2024-07-08 | | Release date: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the locust beta-carotene-binding protein (BBP) complexed with beta-carotene
To Be Published
|
|
9IN6
 
 | | Capsid of Vibrio cholerae phage mature VP1 | | Descriptor: | SHP of VP1, major capsid of VP1 | | Authors: | Liu, H.R, Pang, H. | | Deposit date: | 2024-07-05 | | Release date: | 2024-08-14 | | Method: | ELECTRON MICROSCOPY (2.8 Å) | | Cite: | Three-dimensional structures of Vibrio cholerae typing podophage VP1 in two states
to be published
|
|
9IMP
 
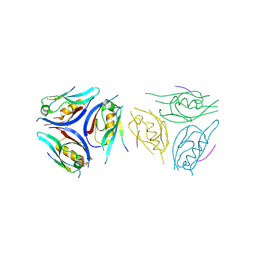 | | The complex of PDZ3 and PBM | | Descriptor: | INSC spindle orientation adaptor protein, Partitioning defective 3 homolog | | Authors: | Huang, S.J. | | Deposit date: | 2024-07-04 | | Release date: | 2024-09-11 | | Last modified: | 2024-09-18 | | Method: | X-RAY DIFFRACTION (2.87 Å) | | Cite: | mInsc coordinates Par3 and NuMA condensates for assembly of the spindle orientation machinery in asymmetric cell division.
Int.J.Biol.Macromol., 279, 2024
|
|
9IMJ
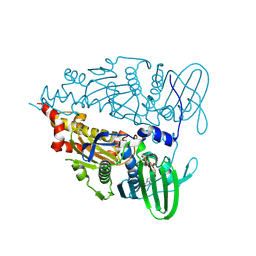 
 | |
9IMA
 
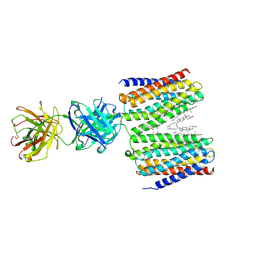 | | Cryo-EM structure for the GPRC5D complexed with Talquetamab Fab | | Descriptor: | CHOLESTEROL, G-protein coupled receptor family C group 5 member D, Talquetamab Fab (anti-GPRC5D) Heavy chain, ... | | Authors: | Jeong, J, Shin, J, Park, J, Cho, Y. | | Deposit date: | 2024-07-02 | | Release date: | 2024-09-11 | | Method: | ELECTRON MICROSCOPY (2.65 Å) | | Cite: | Structural Basis for the Recognition of GPRC5D by Talquetamab, a Bispecific Antibody for Multiple Myeloma.
J.Mol.Biol., 436, 2024
|
|
9ILV
 
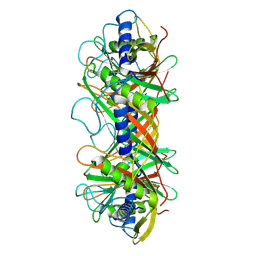 | | Structure of the bacteriophage T5 connector complex | | Descriptor: | Tail tube terminator protein p142 | | Authors: | Peng, Y.N, Liu, H.R. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-04 | | Method: | ELECTRON MICROSCOPY (4.8 Å) | | Cite: | Structures of Mature and Urea-Treated Empty Bacteriophage T5: Insights into Siphophage Infection and DNA Ejection.
Int J Mol Sci, 25, 2024
|
|
9ILP
 
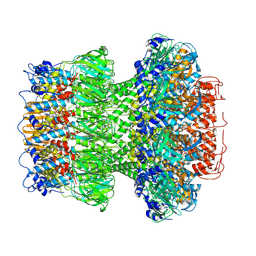 | | Structure of the bacteriophage T5 portal complex | | Descriptor: | Head completion protein, Portal protein pb7 | | Authors: | Peng, Y.N, Liu, H.R. | | Deposit date: | 2024-07-01 | | Release date: | 2024-09-18 | | Method: | ELECTRON MICROSCOPY (3.4 Å) | | Cite: | Structures of Mature and Urea-Treated Empty Bacteriophage T5: Insights into Siphophage Infection and DNA Ejection.
Int J Mol Sci, 25, 2024
|
|
