4CTK
 
 | | DENGUE 3 NS5 METHYLTRANSFERASE BOUND TO S-ADENOSYL METHIONINE AND FRAGMENT 2A4 | | Descriptor: | DIMETHYL SULFOXIDE, POLYPROTEIN, S-ADENOSYLMETHIONINE, ... | | Authors: | Barral, K, Bricogne, G, Sharff, A. | | Deposit date: | 2014-03-14 | | Release date: | 2014-04-16 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (1.53 Å) | | Cite: | Assessment of Dengue Virus Helicase and Methyltransferase as Targets for Fragment-Based Drug Discovery.
Antiviral Res., 106, 2014
|
|
3AT7
 | | Crystal structure of bacterial cell-surface alginate-binding protein Algp7 | | Descriptor: | Alginate-binding flagellin | | Authors: | Maruyama, Y, Ochiai, A, Mikami, B, Hashimoto, W, Murata, K. | | Deposit date: | 2010-12-27 | | Release date: | 2011-02-23 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of bacterial cell-surface alginate-binding protein with an M75 peptidase motif.
Biochem.Biophys.Res.Commun., 405, 2011
|
|
3R3A
 
 | | Crystal structure of Arthrobacter sp. strain SU 4-hydroxybenzoyl CoA thioesterase mutant Q58A complexed with 4-hydroxybenzoic acid and CoA | | Descriptor: | 4-hydroxybenzoyl-CoA thioesterase, COENZYME A, P-HYDROXYBENZOIC ACID | | Authors: | Holden, H.M, Thoden, J.B, Song, F, Zhuang, Z, Trujillo, M, Dunaway-Mariano, D. | | Deposit date: | 2011-03-15 | | Release date: | 2012-03-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Catalytic Mechanism of the Hotdog-fold Enzyme Superfamily 4-Hydroxybenzoyl-CoA Thioesterase from Arthrobacter sp. Strain SU.
Biochemistry, 51, 2012
|
|
4EK0
 
 | | Initial Thaumatin Structure for Radiation Damage Experiment at 25 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-08 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4EKH
 
 | | Final Thaumatin Structure for Radiation Damage Experiment at 100 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-09 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4ELA
 
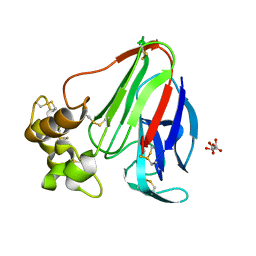 | | Final Thaumatin Structure for Radiation Damage Experiment at 300 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-10 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4EPD
 
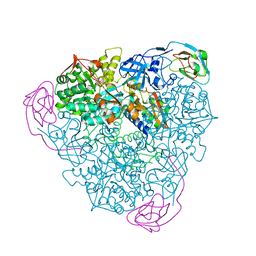 | | Initial Urease Structure for Radiation Damage Experiment at 300 K | | Descriptor: | NICKEL (II) ION, Urease subunit alpha, Urease subunit beta, ... | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-17 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3SGS
 | |
3SGO
 | |
4EKO
 
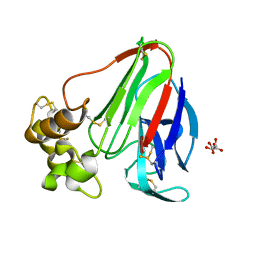 | | Initial Thaumatin Structure for Radiation Damage Experiment at 180 K | | Descriptor: | L(+)-TARTARIC ACID, Thaumatin-1 | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-09 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
4EPB
 
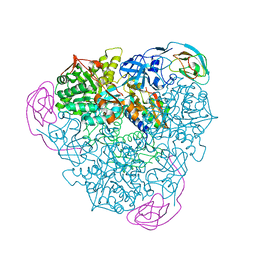 | | Final Urease Structure for Radiation Damage Experiment at 100 K | | Descriptor: | NICKEL (II) ION, Urease subunit alpha, Urease subunit beta, ... | | Authors: | Warkentin, M, Badeau, R, Hopkins, J.B, Thorne, R.E. | | Deposit date: | 2012-04-17 | | Release date: | 2012-08-29 | | Last modified: | 2013-01-23 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Spatial distribution of radiation damage to crystalline proteins at 25-300 K.
Acta Crystallogr.,Sect.D, 68, 2012
|
|
3TTA
 
 | |
1OGZ
 
 | | Crystal Structure Of 5-3-Ketosteroid Isomerase Mutants P39A Complexed With Equilenin From Pseudomonas Testosteroni | | Descriptor: | EQUILENIN, STEROID DELTA-ISOMERASE | | Authors: | Nam, G.H, Cha, S.-S, Yun, Y.S, Oh, Y.H, Hong, B.H, Lee, H.-S, Choi, K.Y. | | Deposit date: | 2003-05-20 | | Release date: | 2003-09-04 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Conserved Cis-Pro39 Residue Plays a Crucial Role in the Proper Positioning of the Catalytic Base Asp38 in Ketosteroid Isomerase from Comamonas Testosteroni.
Biochem.J., 375, 2003
|
|
1A8J
 
 | |
1A8H
 
 | | METHIONYL-TRNA SYNTHETASE FROM THERMUS THERMOPHILUS | | Descriptor: | METHIONYL-TRNA SYNTHETASE, ZINC ION | | Authors: | Sugiura, I, Nureki, O, Ugaji, Y, Kuwabara, S, Lober, B, Giege, R, Moras, D, Yokoyama, S, Konno, M, RIKEN Structural Genomics/Proteomics Initiative (RSGI) | | Deposit date: | 1998-03-26 | | Release date: | 1999-05-04 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | The 2.0 A crystal structure of Thermus thermophilus methionyl-tRNA synthetase reveals two RNA-binding modules.
Structure, 8, 2000
|
|
2OR2
 
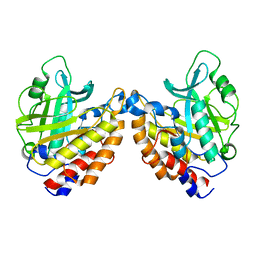 | | Structure of the W47A/W242A Mutant of Bacterial Phosphatidylinositol-Specific Phospholipase C | | Descriptor: | 1-phosphatidylinositol phosphodiesterase | | Authors: | Shao, C, Shi, X, Wehbi, H, Zambonelli, C, Head, J.F, Seaton, B.A, Roberts, M.F. | | Deposit date: | 2007-02-01 | | Release date: | 2007-02-13 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Dimer structure of an interfacially impaired phosphatidylinositol-specific phospholipase C.
J.Biol.Chem., 282, 2007
|
|
3R3C
 
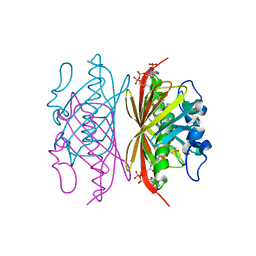 | | Crystal structure of Arthrobacter sp. strain SU 4-hydroxybenzoyl CoA thioesterase mutant H64A complexed with 4-hydroxyphenacyl CoA | | Descriptor: | 4-HYDROXYPHENACYL COENZYME A, 4-hydroxybenzoyl-CoA thioesterase | | Authors: | Holden, H.M, Thoden, J.B, Song, F, Zhuang, Z, Trujillo, M, Dunaway-Mariano, D. | | Deposit date: | 2011-03-15 | | Release date: | 2012-03-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Catalytic Mechanism of the Hotdog-fold Enzyme Superfamily 4-Hydroxybenzoyl-CoA Thioesterase from Arthrobacter sp. Strain SU.
Biochemistry, 51, 2012
|
|
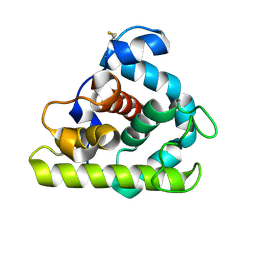
153L
 
 | |
2O6R
 | |
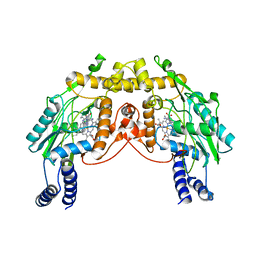
2FLQ
 
 | |
3R37
 
 | | Crystal structure of Arthrobacter sp. strain SU 4-hydroxybenzoyl CoA thioesterase mutant E73Q complexed with 4-hydroxyphenacyl CoA | | Descriptor: | 4-HYDROXYPHENACYL COENZYME A, 4-hydroxybenzoyl-CoA thioesterase | | Authors: | Holden, H.M, Thoden, J.B, Song, F, Zhuang, Z, Trujillo, M, Dunaway-Mariano, D. | | Deposit date: | 2011-03-15 | | Release date: | 2012-03-28 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | The Catalytic Mechanism of the Hotdog-fold Enzyme Superfamily 4-Hydroxybenzoyl-CoA Thioesterase from Arthrobacter sp. Strain SU.
Biochemistry, 51, 2012
|
|
2FYS
 
 | | Crystal structure of Erk2 complex with KIM peptide derived from MKP3 | | Descriptor: | Dual specificity protein phosphatase 6, Mitogen-activated protein kinase 1 | | Authors: | Liu, S, Sun, J.P, Zhou, B, Zhang, Z.Y. | | Deposit date: | 2006-02-08 | | Release date: | 2006-04-11 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structural basis of docking interactions between ERK2 and MAP kinase phosphatase 3
Proc.Natl.Acad.Sci.Usa, 103, 2006
|
|
3TRN
 
 | |
3TPN
 
 | | Crystal structure of M-PMV dUTPASE complexed with dUPNPP, substrate | | Descriptor: | 2'-DEOXYURIDINE 5'-ALPHA,BETA-IMIDO-TRIPHOSPHATE, 2,3-DIHYDROXY-1,4-DITHIOBUTANE, 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, ... | | Authors: | Nemeth, V, Barabas, O, Vertessy, G.B. | | Deposit date: | 2011-09-08 | | Release date: | 2011-10-12 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structural Snapshots of Enzyme-Catalysed Phosphate Ester Hydrolysis Directly Visualize In-line Attack and Inversion
To be Published
|
|
3TSL
 
 | |
