7P0O
 
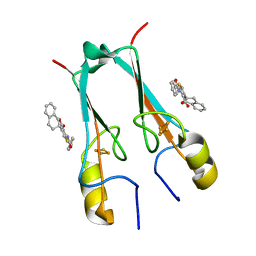 | | mitoNEET bound to M1 molecule | | Descriptor: | 2-benzamido-4-[(2~{R})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, 2-benzamido-4-[(2~{S})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, CDGSH iron-sulfur domain-containing protein 1, ... | | Authors: | Livnah, O, Eisenberg-Domovich, Y, Marjault, H.B, Nechushtai, R. | | Deposit date: | 2021-06-30 | | Release date: | 2022-05-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | An anti-diabetic drug targets NEET (CISD) proteins through destabilization of their [2Fe-2S] clusters.
Commun Biol, 5, 2022
|
|
4ND2
 
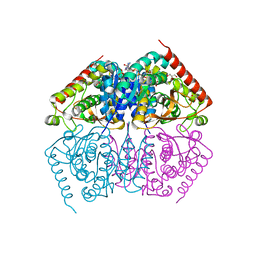 | |
4NDF
 
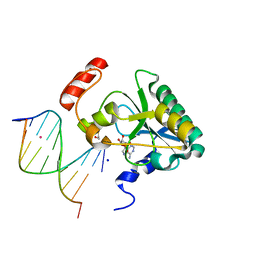 | | Human Aprataxin (Aptx) bound to RNA-DNA, AMP, and Zn - product complex | | Descriptor: | 5'-D(*GP*AP*AP*TP*CP*AP*TP*AP*AP*C)-3', 5'-R(P*G)-D(P*TP*TP*AP*TP*GP*AP*TP*TP*C)-3', ADENOSINE MONOPHOSPHATE, ... | | Authors: | Schellenberg, M.J, Tumbale, P.S, Williams, R.S. | | Deposit date: | 2013-10-26 | | Release date: | 2013-12-18 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.944 Å) | | Cite: | Aprataxin resolves adenylated RNA-DNA junctions to maintain genome integrity.
Nature, 506, 2013
|
|
4WGG
 
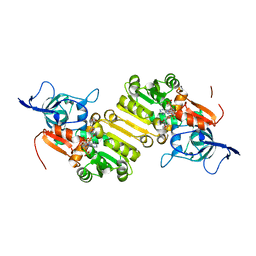 | | STRUCTURE OF THE TERNARY COMPLEX OF A ZINGIBER OFFICINALE DOUBLE BOND REDUCTASE IN COMPLEX WITH NADP AND CONIFERYL ALDEHYDE | | Descriptor: | (2E)-3-(4-hydroxy-3-methoxyphenyl)prop-2-enal, Double Bond Reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Collery, J, Langlois d'Estaintot, B, Buratto, J, Granier, T, Gallois, B, Willis, M.A, Sang, Y, Flores-Sanchez, I.J, Gang, D.R. | | Deposit date: | 2014-09-18 | | Release date: | 2015-09-30 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | STRUCTURE OF ZINGIBER OFFICINALE DOUBLE BOND REDUCTASE
to be published
|
|
4WHR
 
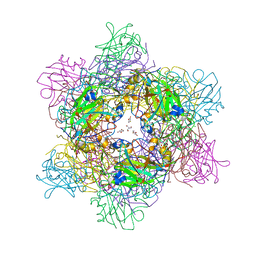 | | Anhydride reaction intermediate trapped in Protocatechuate 3,4-dioxygenase (pseudomonas putida) at pH 8.5 | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, 4-fluorobenzene-1,2-diol, 4-fluorooxepine-2,7-dione, ... | | Authors: | Knoot, C.J, Lipscomb, J.D. | | Deposit date: | 2014-09-23 | | Release date: | 2014-12-31 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystal structures of alkylperoxo and anhydride intermediates in an intradiol ring-cleaving dioxygenase.
Proc.Natl.Acad.Sci.USA, 112, 2015
|
|
7P0P
 
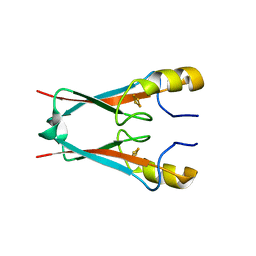 | | NAF-1 bound to M1 molecule | | Descriptor: | 2-benzamido-4-[(2~{R})-1,2,3,4-tetrahydronaphthalen-2-yl]thiophene-3-carboxylic acid, CDGSH iron-sulfur domain-containing protein 2, FE2/S2 (INORGANIC) CLUSTER | | Authors: | Livnah, O, Eisenberg-Domovich, Y, Marjault, H.B, Nechushtai, R. | | Deposit date: | 2021-06-30 | | Release date: | 2022-05-25 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.74 Å) | | Cite: | An anti-diabetic drug targets NEET (CISD) proteins through destabilization of their [2Fe-2S] clusters.
Commun Biol, 5, 2022
|
|
4NGL
 
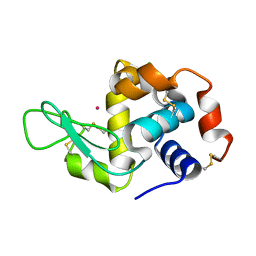 | | Previously de-ionized HEW lysozyme batch crystallized in 0.6 M CoCl2 | | Descriptor: | CHLORIDE ION, COBALT (II) ION, Lysozyme C | | Authors: | Benas, P, Legrand, L, Ries-Kautt, M. | | Deposit date: | 2013-11-02 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.52 Å) | | Cite: | Weak protein-cationic co-ion interactions addressed by X-ray crystallography and mass spectrometry.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
4WJB
 
 | | X-ray crystal structure of a putative amidohydrolase/peptidase from Burkholderia cenocepacia | | Descriptor: | 1,2-ETHANEDIOL, Putative amidohydrolase/peptidase, SULFATE ION, ... | | Authors: | Lukacs, C.M, Dranow, D.M, Edwards, T.E, Lorimer, D, Seattle Structural Genomics Center for Infectious Disease (SSGCID) | | Deposit date: | 2014-09-29 | | Release date: | 2014-10-08 | | Last modified: | 2023-09-27 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | X-ray crystal structure of a putative amidohydrolase/peptidase from Burkholderia cenocepacia
To Be Published
|
|
4WKU
 
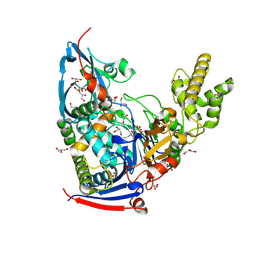 | | n-Alkylboronic Acid Inhibitors Reveal Determinants of Ligand Specificity in the Quorum-Quenching and Siderophore Biosynthetic Enzyme PvdQ | | Descriptor: | Acyl-homoserine lactone acylase PvdQ, GLYCEROL, hexyl(trihydroxy)borate(1-) | | Authors: | Wu, R, Clevenger, K.D, Fast, W, Liu, D. | | Deposit date: | 2014-10-03 | | Release date: | 2014-11-12 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | n-Alkylboronic Acid Inhibitors Reveal Determinants of Ligand Specificity in the Quorum-Quenching and Siderophore Biosynthetic Enzyme PvdQ.
Biochemistry, 53, 2014
|
|
4NHP
 
 | | X-ray structure of the complex between the hen egg white lysozyme and pentachlorocarbonyliridate (III) (4 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-05 | | Release date: | 2014-09-17 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.42 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4NHT
 
 | | X-ray structure of the complex between hen egg white lysozyme and pentachlorocarbonyliridate(III) (6 days) | | Descriptor: | CHLORIDE ION, Lysozyme C, SODIUM ION, ... | | Authors: | Petruk, A.A, Bikiel, D.E, Vergara, A, Merlino, A. | | Deposit date: | 2013-11-05 | | Release date: | 2014-09-17 | | Last modified: | 2015-06-24 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Interaction between proteins and Ir based CO releasing molecules: mechanism of adduct formation and CO release.
Inorg.Chem., 53, 2014
|
|
4WLD
 
 | |
4NMD
 
 | |
4WOA
 
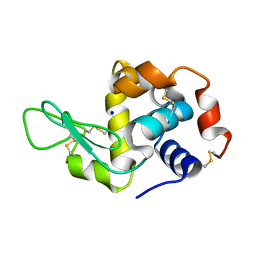 | |
4NSJ
 | | Carboplatin binding to HEWL in 2M NH4formate, 0.1M HEPES at pH 7.5 | | Descriptor: | DIMETHYL SULFOXIDE, FORMIC ACID, Lysozyme C, ... | | Authors: | Tanley, S.W.M, Diederichs, K, Kroon-Batenburg, L.M.J, Levy, C, Schreurs, A.M.M, Helliwell, J.R. | | Deposit date: | 2013-11-28 | | Release date: | 2014-10-15 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Carboplatin binding to histidine.
Acta Crystallogr F Struct Biol Commun, 70, 2014
|
|
4NJO
 
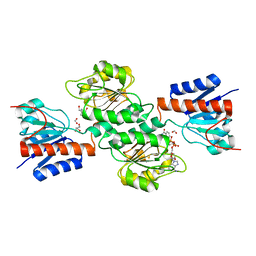 | |
4WQ0
 
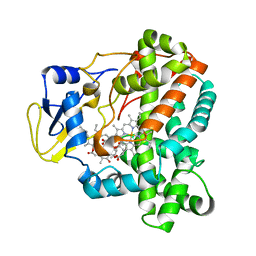 | |
4NUY
 
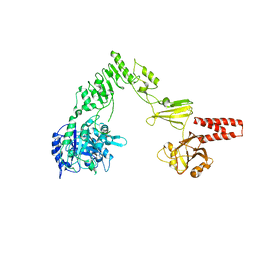 | | Crystal structure of EndoS, an endo-beta-N-acetyl-glucosaminidase from Streptococcus pyogenes | | Descriptor: | CALCIUM ION, Endo-beta-N-acetylglucosaminidase F2 | | Authors: | Trastoy, B, Guenther, S, Snyder, G.A, Sundberg, E.J. | | Deposit date: | 2013-12-04 | | Release date: | 2014-04-09 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.609 Å) | | Cite: | Crystal structure of Streptococcus pyogenes EndoS, an immunomodulatory endoglycosidase specific for human IgG antibodies.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
4NWE
 
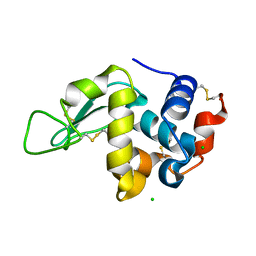 | | Lysozyme UNDER 30 BAR PRESSURE OF NITROUS OXIDE | | Descriptor: | CHLORIDE ION, Lysozyme C, NITROUS OXIDE | | Authors: | Colloc'h, N, Prange, T. | | Deposit date: | 2013-12-06 | | Release date: | 2014-09-24 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystallographic Studies with Xenon and Nitrous Oxide Provide Evidence for Protein-dependent Processes in the Mechanisms of General Anesthesia
Anesthesiology, 121, 2014
|
|
4NMC
 
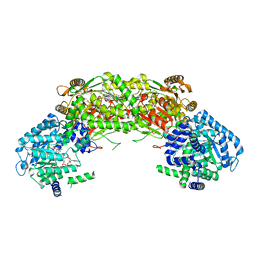 | |
476D
 
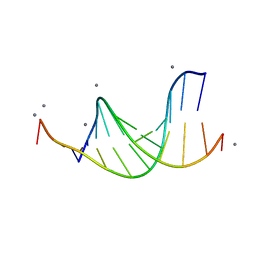 | | CALCIUM FORM OF B-DNA UNDECAMER GCGAATTCGCG | | Descriptor: | 5'-D(*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3', CALCIUM ION | | Authors: | Minasov, G, Tereshko, V, Egli, M. | | Deposit date: | 1999-06-25 | | Release date: | 1999-07-19 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Atomic-resolution crystal structures of B-DNA reveal specific influences of divalent metal ions on conformation and packing.
J.Mol.Biol., 291, 1999
|
|
4X4S
 
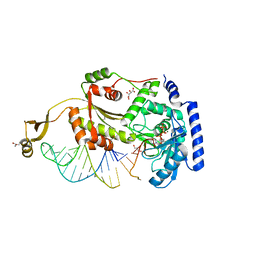 | |
4X4O
 
 | |
4ND1
 
 | |
4NGV
 
 | | Previously de-ionized HEW lysozyme batch crystallized in 0.5 M YbCl3 | | Descriptor: | CHLORIDE ION, Lysozyme C, YTTERBIUM (III) ION | | Authors: | Benas, P, Legrand, L, Ries-Kautt, M. | | Deposit date: | 2013-11-03 | | Release date: | 2014-05-28 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Weak protein-cationic co-ion interactions addressed by X-ray crystallography and mass spectrometry.
Acta Crystallogr.,Sect.D, 70, 2014
|
|
