1QE5
 
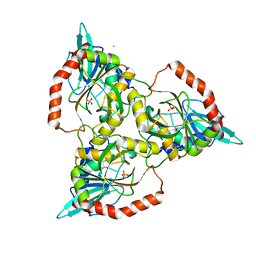 | | PURINE NUCLEOSIDE PHOSPHORYLASE FROM CELLULOMONAS SP. IN COMPLEX WITH PHOSPHATE | | Descriptor: | CALCIUM ION, PENTOSYLTRANSFERASE, PHOSPHATE ION | | Authors: | Tebbe, J, Bzowska, A, Wielgus-Kutrowska, B, Schroeder, W, Kazimierczuk, Z, Shugar, D, Saenger, W, Koellner, G. | | Deposit date: | 1999-07-13 | | Release date: | 1999-12-12 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structure of the purine nucleoside phosphorylase (PNP) from Cellulomonas sp. and its implication for the mechanism of trimeric PNPs.
J.Mol.Biol., 294, 1999
|
|
8BFH
 
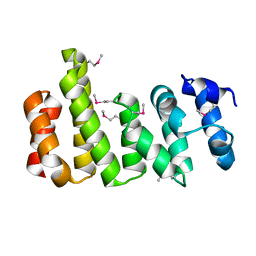 | | CNOT11 | | Descriptor: | CCR4-NOT transcription complex subunit 11 | | Authors: | Basquin, J, Ozgur, S, Conti, E. | | Deposit date: | 2022-10-26 | | Release date: | 2022-12-28 | | Last modified: | 2023-02-15 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | The human CNOT1-CNOT10-CNOT11 complex forms a structural platform for protein-protein interactions.
Cell Rep, 42, 2023
|
|
8JU9
 
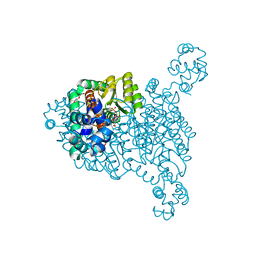 | |
1QSA
 
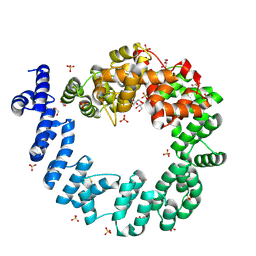 | | CRYSTAL STRUCTURE OF THE 70 KDA SOLUBLE LYTIC TRANSGLYCOSYLASE SLT70 FROM ESCHERICHIA COLI AT 1.65 ANGSTROMS RESOLUTION | | Descriptor: | ACETATE ION, GLYCEROL, PROTEIN (SOLUBLE LYTIC TRANSGLYCOSYLASE SLT70), ... | | Authors: | van Asselt, E.J, Thunnissen, A.-M.W.H, Dijkstra, B.W. | | Deposit date: | 1999-06-20 | | Release date: | 1999-09-01 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | High resolution crystal structures of the Escherichia coli lytic transglycosylase Slt70 and its complex with a peptidoglycan fragment.
J.Mol.Biol., 291, 1999
|
|
1APY
 
 | | HUMAN ASPARTYLGLUCOSAMINIDASE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, ASPARTYLGLUCOSAMINIDASE, beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | | Authors: | Rouvinen, J, Oinonen, C. | | Deposit date: | 1995-06-14 | | Release date: | 1996-12-23 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Three-dimensional structure of human lysosomal aspartylglucosaminidase.
Nat.Struct.Biol., 2, 1995
|
|
8BBV
 
 | | Coproporphyrin III - LmCpfC complex soaked 2min with Fe2+ | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, Coproporphyrin III ferrochelatase, ... | | Authors: | Gabler, T, Hofbauer, S. | | Deposit date: | 2022-10-14 | | Release date: | 2023-01-11 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Iron insertion into coproporphyrin III-ferrochelatase complex: Evidence for an intermediate distorted catalytic species.
Protein Sci., 32, 2023
|
|
8E89
 
 | | Human DNA polymerase eta-DNA-rU-ended primer-binary complex | | Descriptor: | DNA (5'-D(*CP*AP*TP*TP*AP*TP*GP*AP*CP*GP*CP*T)-3'), DNA polymerase eta, DNA/RNA (5'-D(*AP*GP*CP*GP*TP*CP*A)-R(P*U)-3') | | Authors: | Chang, C, Gao, Y. | | Deposit date: | 2022-08-25 | | Release date: | 2023-02-01 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Primer terminal ribonucleotide alters the active site dynamics of DNA polymerase eta and reduces DNA synthesis fidelity.
J.Biol.Chem., 299, 2023
|
|
6HZA
 
 | |
7Y8A
 
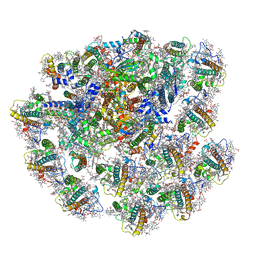 | | Cryo-EM structure of cryptophyte photosystem I | | Descriptor: | (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E})-3,7,12,16-tetramethyl-18-[(4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohexen-1-yl]octadeca-3,5,7,9,11,13,15-heptaen-1,17-diynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-(2,6,6-trimethylcyclohexen-1-yl)octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, (1~{R})-3,5,5-trimethyl-4-[(3~{E},5~{E},7~{E},9~{E},11~{E},13~{E},15~{E},17~{E})-3,7,12,16-tetramethyl-18-[(1~{R},4~{R})-2,6,6-trimethyl-4-oxidanyl-cyclohex-2-en-1-yl]octadeca-3,5,7,9,11,13,15,17-octaen-1-ynyl]cyclohex-3-en-1-ol, ... | | Authors: | Zhao, L.S, Zhang, Y.Z, Liu, L.N, Li, K. | | Deposit date: | 2022-06-23 | | Release date: | 2023-04-12 | | Last modified: | 2023-07-12 | | Method: | ELECTRON MICROSCOPY (2.71 Å) | | Cite: | Structural basis and evolution of the photosystem I-light-harvesting supercomplex of cryptophyte algae.
Plant Cell, 35, 2023
|
|
1QE1
 
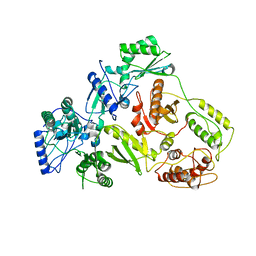 | | CRYSTAL STRUCTURE OF 3TC-RESISTANT M184I MUTANT OF HIV-1 REVERSE TRANSCRIPTASE | | Descriptor: | REVERSE TRANSCRIPTASE, SUBUNIT P51, SUBUNIT P66 | | Authors: | Sarafianos, S.G, Das, K, Ding, J, Hughes, S.H, Arnold, E. | | Deposit date: | 1999-07-12 | | Release date: | 1999-08-30 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.85 Å) | | Cite: | Lamivudine (3TC) resistance in HIV-1 reverse transcriptase involves steric hindrance with beta-branched amino acids.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
8BNQ
 
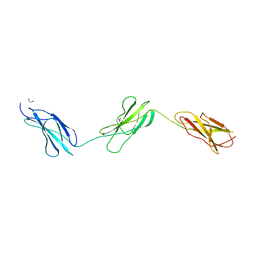 | |
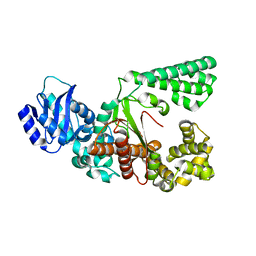
8JRB
 
 | |
5CP2
 
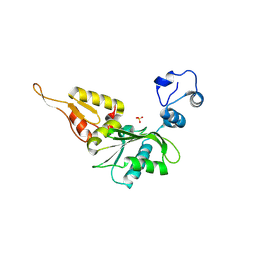 | |
8K6P
 
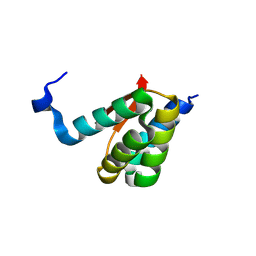 | | Crystal structure of SHARPIN LTM motif | | Descriptor: | Sharpin | | Authors: | Yan, Z, Pan, L.F. | | Deposit date: | 2023-07-25 | | Release date: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.86 Å) | | Cite: | Mechanistic insights into the homo-dimerization of HOIL-1L and SHARPIN.
Biochem.Biophys.Res.Commun., 689, 2023
|
|
8K6Q
 
 | | Crystal structure of HOIL-1L LTM domain | | Descriptor: | RanBP-type and C3HC4-type zinc finger-containing protein 1 | | Authors: | Yan, Z, Pan, L.F. | | Deposit date: | 2023-07-25 | | Release date: | 2024-07-03 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | Mechanistic insights into the homo-dimerization of HOIL-1L and SHARPIN.
Biochem.Biophys.Res.Commun., 689, 2023
|
|
7WU0
 
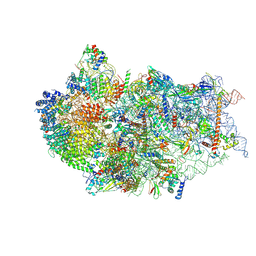 | | Cryo-EM structure of a human pre-40S ribosomal subunit - State RRP12-B3 | | Descriptor: | 18S rRNA, 40S ribosomal protein S11, 40S ribosomal protein S12, ... | | Authors: | Cheng, J, Lau, B, Thoms, M, Ameismeier, M, Berninghausen, O, Hurt, E, Beckmann, R. | | Deposit date: | 2022-02-05 | | Release date: | 2022-10-19 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.3 Å) | | Cite: | The nucleoplasmic phase of pre-40S formation prior to nuclear export.
Nucleic Acids Res., 50, 2022
|
|
8B2U
 
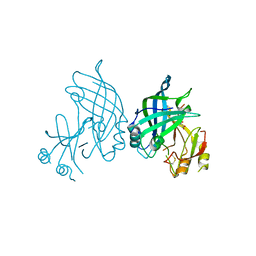 | | Crystal structure of SUDV VP40 in complex with salicylic acid | | Descriptor: | 2-HYDROXYBENZOIC ACID, Matrix protein VP40 | | Authors: | Werner, A.-D, Krapoth, N, Norris, M.J, Heine, A, Klebe, G, Ollmann Saphire, E, Becker, S. | | Deposit date: | 2022-09-14 | | Release date: | 2023-09-27 | | Last modified: | 2024-08-14 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Development of a Crystallographic Screening to Identify Sudan Virus VP40 Ligands.
Acs Omega, 9, 2024
|
|
7WTS
 
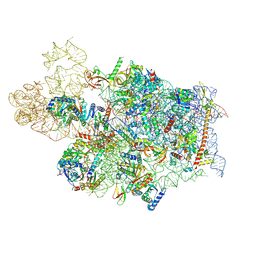 | | Cryo-EM structure of a human pre-40S ribosomal subunit - State UTP14 | | Descriptor: | 18S rRNA, 40S ribosomal protein S11, 40S ribosomal protein S13, ... | | Authors: | Cheng, J, Lau, B, Thoms, M, Ameismeier, M, Berninghausen, O, Hurt, E, Beckmann, R. | | Deposit date: | 2022-02-05 | | Release date: | 2022-10-19 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.2 Å) | | Cite: | The nucleoplasmic phase of pre-40S formation prior to nuclear export.
Nucleic Acids Res., 50, 2022
|
|
7PCY
 
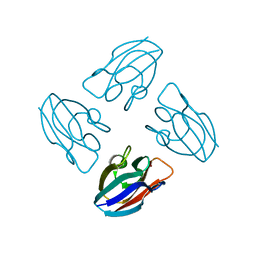 | | THE CRYSTAL STRUCTURE OF PLASTOCYANIN FROM A GREEN ALGA, ENTEROMORPHA PROLIFERA | | Descriptor: | COPPER (II) ION, PLASTOCYANIN | | Authors: | Collyer, C.A, Guss, J.M, Freeman, H.C. | | Deposit date: | 1989-09-22 | | Release date: | 1990-07-15 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of plastocyanin from a green alga, Enteromorpha prolifera.
J.Mol.Biol., 211, 1990
|
|
7WTX
 
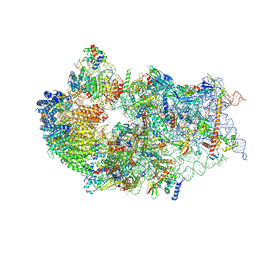 | | Cryo-EM structure of a human pre-40S ribosomal subunit - State RRP12-B1 | | Descriptor: | 18S rRNA, 40S ribosomal protein S11, 40S ribosomal protein S12, ... | | Authors: | Cheng, J, Lau, B, Thoms, M, Ameismeier, M, Berninghausen, O, Hurt, E, Beckmann, R. | | Deposit date: | 2022-02-05 | | Release date: | 2022-10-19 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3.1 Å) | | Cite: | The nucleoplasmic phase of pre-40S formation prior to nuclear export.
Nucleic Acids Res., 50, 2022
|
|
5CHA
 
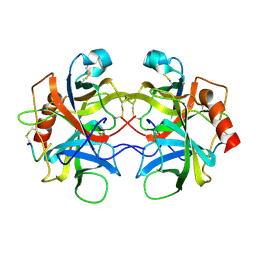 | |
7WTZ
 
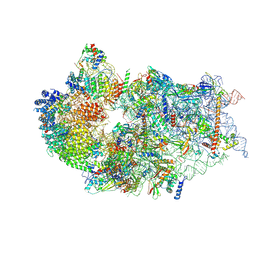 | | Cryo-EM structure of a human pre-40S ribosomal subunit - State RRP12-B2 | | Descriptor: | 18S rRNA, 40S ribosomal protein S11, 40S ribosomal protein S12, ... | | Authors: | Cheng, J, Lau, B, Thoms, M, Ameismeier, M, Berninghausen, O, Hurt, E, Beckmann, R. | | Deposit date: | 2022-02-05 | | Release date: | 2022-10-19 | | Last modified: | 2024-06-26 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | The nucleoplasmic phase of pre-40S formation prior to nuclear export.
Nucleic Acids Res., 50, 2022
|
|
8K3G
 
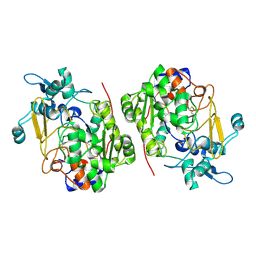 | |
5CSC
 
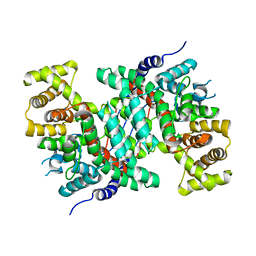 | |
7D42
 
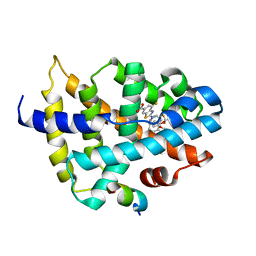 | |
