2XBW
 
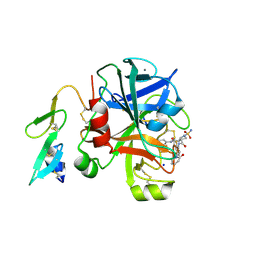 | | Factor Xa in complex with a pyrrolidine-3,4-dicarboxylic acid inhibitor | | Descriptor: | (3R,4R)-1-SULFAMOYL-PYRROLIDINE-3,4-DICARBOXYLIC ACID 3-[(4-CHLORO-PHENYL)-AMIDE] 4-{[2-FLUORO-4-(2-OXO-2H-PYRIDIN-1-YL)-PHENYL]-AMIDE}, ACTIVATED FACTOR XA HEAVY CHAIN, CALCIUM ION, ... | | Authors: | Banner, D.W, Benz, J, Schlatter, D, Thomi, S, Haap, W. | | Deposit date: | 2010-04-15 | | Release date: | 2010-08-04 | | Last modified: | 2024-05-01 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | Discovery of a Factor Xa Inhibitor (3R,4R)-1-(2,2-Difluoro-Ethyl)-Pyrrolidine-3,4-Dicarboxylic Acid 3-[(5-Chloro-Pyridin-2-Yl)-Amide] 4-{[2-Fluoro-4-(2-Oxo-2H-Pyridin-1-Yl)-Phenyl]-Amide} as a Clinical Candidate.
Bioorg.Med.Chem., 20, 2010
|
|
3LLN
 
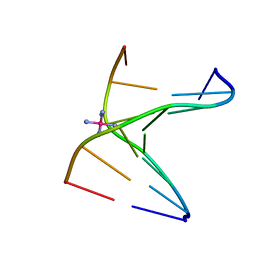 | | Comparison between the orthorhombic an tetragonal form of the heptamer sequence d(GCG(xT)GCG)/d(CGCACGC). | | Descriptor: | COBALT HEXAMMINE(III), DNA (5'-D(*CP*GP*CP*AP*CP*GP*C)-3'), DNA (5'-D(*GP*CP*GP*(XTR)P*GP*CP*G)-3') | | Authors: | Robeyns, K, Herdewijn, P, Van Meervelt, L. | | Deposit date: | 2010-01-29 | | Release date: | 2010-08-18 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Comparison between the orthorhombic and tetragonal forms of the heptamer sequence d[GCG(xT)GCG]/d(CGCACGC)
Acta Crystallogr.,Sect.F, 66, 2010
|
|
3LGP
 
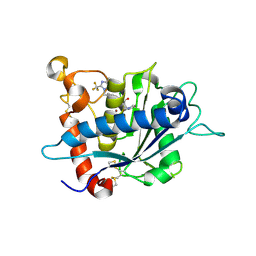 | | Crystal structure of catalytic domain of tace with benzimidazolyl-thienyl-tartrate based inhibitor | | Descriptor: | (2R,3R)-4-[(2R)-2-(3-chlorophenyl)pyrrolidin-1-yl]-2,3-dihydroxy-4-oxo-N-[(5-{[2-(trifluoromethyl)-1H-benzimidazol-1-yl]methyl}thiophen-2-yl)methyl]butanamide, Disintegrin and metalloproteinase domain-containing protein 17, ZINC ION | | Authors: | Orth, P. | | Deposit date: | 2010-01-21 | | Release date: | 2010-07-28 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure and activity relationships of tartrate-based TACE inhibitors.
Bioorg.Med.Chem.Lett., 20, 2010
|
|
3LAQ
 
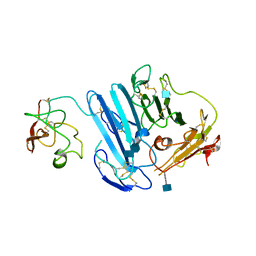 | | Structure-based engineering of species selectivity in the uPA-uPAR interaction | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Urokinase plasminogen activator surface receptor, Urokinase-type plasminogen activator | | Authors: | Huang, M. | | Deposit date: | 2010-01-06 | | Release date: | 2010-02-02 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Structure-based engineering of species selectivity in the interaction between urokinase and its receptor: implication for preclinical cancer therapy.
J.Biol.Chem., 285, 2010
|
|
2YBI
 
 | |
1DGN
 
 | | SOLUTION STRUCTURE OF ICEBERG, AN INHIBITOR OF INTERLEUKIN-1BETA GENERATION | | Descriptor: | ICEBERG (PROTEASE INHIBITOR) | | Authors: | Humke, E.W, Shriver, S.K, Starovasnik, M.A, Fairbrother, W.J, Dixit, V.M. | | Deposit date: | 1999-11-24 | | Release date: | 2000-10-09 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | ICEBERG: a novel inhibitor of interleukin-1beta generation.
Cell(Cambridge,Mass.), 103, 2000
|
|
3LJZ
 
 | | Crystal Structure of Human MMP-13 complexed with an Amino-2-indanol compound | | Descriptor: | (2R)-2-[4-(1,3-benzodioxol-5-yl)benzyl]-N~4~-hydroxy-N~1~-[(1S,2R)-2-hydroxy-2,3-dihydro-1H-inden-1-yl]butanediamide, 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, CALCIUM ION, ... | | Authors: | Shieh, H.-S, Kiefer, J.R. | | Deposit date: | 2010-01-26 | | Release date: | 2011-02-02 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Structure analysis reveals the flexibility of the ADAMTS-5 active site.
Protein Sci., 20, 2011
|
|
1DQZ
 
 | |
3LJ6
 
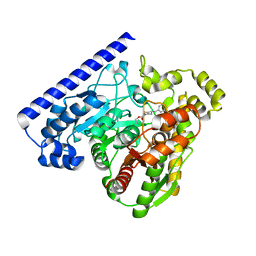 | |
2YD0
 
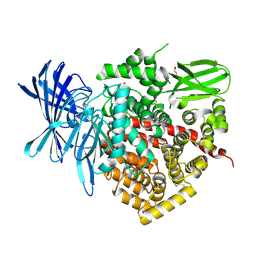 | | Crystal structure of the soluble domain of human endoplasmic reticulum aminopeptidase 1 ERAP1 | | Descriptor: | 1,2-ETHANEDIOL, 2-(3-AMINO-2-HYDROXY-4-PHENYL-BUTYRYLAMINO)-4-METHYL-PENTANOIC ACID, 2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Vollmar, M, Kochan, G, Krojer, T, Ugochukwu, E, Muniz, J.R.C, Raynor, J, Chaikuad, A, Allerston, C, von Delft, F, Bountra, C, Arrowsmith, C.H, Weigelt, J, Edwards, A, Knapp, S. | | Deposit date: | 2011-03-17 | | Release date: | 2011-04-13 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal Structures of the Endoplasmic Reticulum Aminopeptidase-1 (Erap1) Reveal the Molecular Basis for N-Terminal Peptide Trimming.
Proc.Natl.Acad.Sci.USA, 108, 2011
|
|
1ATN
 
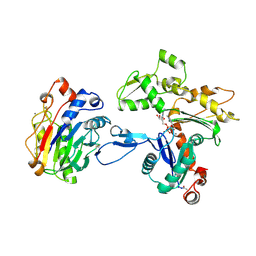 | | Atomic structure of the actin:DNASE I complex | | Descriptor: | ACTIN, ADENOSINE-5'-TRIPHOSPHATE, CALCIUM ION, ... | | Authors: | Kabsch, W, Mannherz, H.G, Suck, D, Pai, E, Holmes, K.C. | | Deposit date: | 1991-03-08 | | Release date: | 1992-07-15 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Atomic structure of the actin:DNase I complex.
Nature, 347, 1990
|
|
1BIM
 
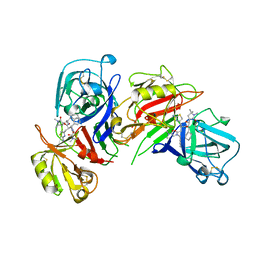 | | CRYSTALLOGRAPHIC STUDIES ON THE BINDING MODES OF P2-P3 BUTANEDIAMIDE RENIN INHIBITORS | | Descriptor: | (2S)-2-[(2-amino-1,3-thiazol-4-yl)methyl]-N~1~-{(1S,2S)-1-(cyclohexylmethyl)-2-hydroxy-2-[(3R)-1,5,5-trimethyl-2-oxopyrrolidin-3-yl]ethyl}-N~4~-[2-(dimethylamino)-2-oxoethyl]-N~4~-[(1S)-1-phenylethyl]butanediamide, Renin | | Authors: | Tong, L. | | Deposit date: | 1995-09-27 | | Release date: | 1996-01-29 | | Last modified: | 2022-03-09 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Crystallographic studies on the binding modes of P2-P3 butanediamide renin inhibitors.
J.Biol.Chem., 270, 1995
|
|
1AWJ
 
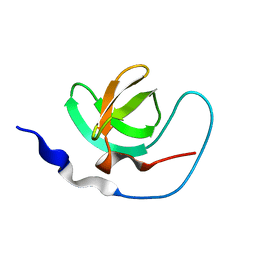 | | INTRAMOLECULAR ITK-PROLINE COMPLEX, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | ITK | | Authors: | Andreotti, A.H, Bunnell, S.C, Feng, S, Berg, L.J, Schreiber, S.L. | | Deposit date: | 1997-10-02 | | Release date: | 1998-01-14 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Regulatory intramolecular association in a tyrosine kinase of the Tec family.
Nature, 385, 1997
|
|
1BGI
 
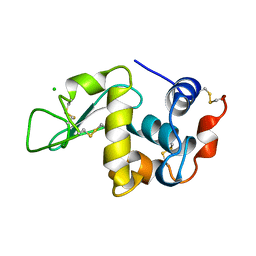 | | ORTHORHOMBIC LYSOZYME CRYSTALLIZED AT HIGH TEMPERATURE (310K) | | Descriptor: | CHLORIDE ION, LYSOZYME | | Authors: | Oki, H, Matsuura, Y, Komatsu, H, Chernov, A.A. | | Deposit date: | 1998-05-28 | | Release date: | 1998-10-28 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Refined structure of orthorhombic lysozyme crystallized at high temperature: correlation between morphology and intermolecular contacts.
Acta Crystallogr.,Sect.D, 55, 1999
|
|
1AOW
 
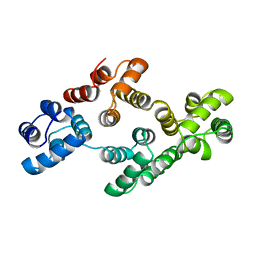 | | ANNEXIN IV | | Descriptor: | ANNEXIN IV | | Authors: | Zanotti, G, Malpeli, G, Gliubich, F, Folli, C, Stoppini, M, Olivi, L, Savoia, A, Berni, R. | | Deposit date: | 1997-07-11 | | Release date: | 1998-01-14 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure of the trigonal crystal form of bovine annexin IV.
Biochem.J., 329, 1998
|
|
1DNK
 
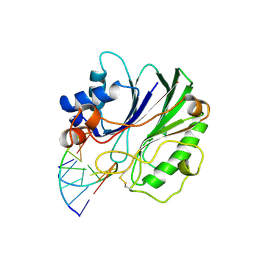 | | THE X-RAY STRUCTURE OF THE DNASE I-D(GGTATACC)2 COMPLEX AT 2.3 ANGSTROMS RESOLUTION | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DNA (5'-D(*GP*GP*TP*AP*TP*AP*C)-3'), DNA (5'-D(*GP*GP*TP*AP*TP*AP*CP*C)-3'), ... | | Authors: | Weston, S.A, Lahm, A, Suck, D. | | Deposit date: | 1992-08-10 | | Release date: | 1994-01-31 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | X-ray structure of the DNase I-d(GGTATACC)2 complex at 2.3 A resolution.
J.Mol.Biol., 226, 1992
|
|
1DQY
 
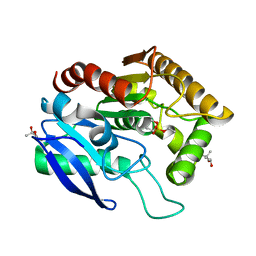 | | CRYSTAL STRUCTURE OF ANTIGEN 85C FROM MYCOBACTERIUM TUBERCULOSIS WITH DIETHYL PHOSPHATE INHIBITOR | | Descriptor: | (4R)-2-METHYLPENTANE-2,4-DIOL, DIETHYL PHOSPHONATE, PROTEIN (ANTIGEN 85-C) | | Authors: | Ronning, D.R, Klabunde, T, Sacchettini, J.C, TB Structural Genomics Consortium (TBSGC) | | Deposit date: | 2000-01-05 | | Release date: | 2000-07-12 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Crystal structure of the secreted form of antigen 85C reveals potential targets for mycobacterial drugs and vaccines.
Nat.Struct.Biol., 7, 2000
|
|
1DTG
 
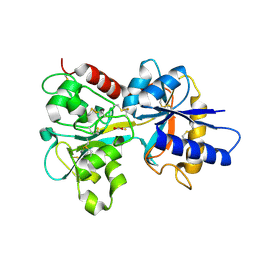 | | HUMAN TRANSFERRIN N-LOBE MUTANT H249E | | Descriptor: | CARBONATE ION, FE (III) ION, TRANSFERRIN | | Authors: | MacGillivray, R.T, Bewley, M.C, Smith, C.A, He, Q.Y, Mason, A.B. | | Deposit date: | 2000-01-12 | | Release date: | 2000-01-21 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Mutation of the iron ligand His 249 to Glu in the N-lobe of human transferrin abolishes the dilysine "trigger" but does not significantly affect iron release.
Biochemistry, 39, 2000
|
|
1D3Q
 
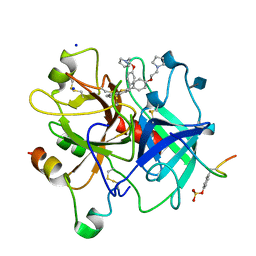 | | CRYSTAL STRUCTURE OF HUMAN ALPHA THROMBIN IN COMPLEX WITH BENZO[B]THIOPHENE INHIBITOR 2 | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 3-[4-(2-PYRROLIDIN-1-YL-ETHOXY)-BENZYL]-2-4-(2-PYRROLIDIN-1-YL-ETHOXY)-PHENYL] -BENZO[B]THIOPHENE, ALPHA-THROMBIN, ... | | Authors: | Chirgadze, N.Y. | | Deposit date: | 1999-09-30 | | Release date: | 2000-10-04 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | The crystal structures of human alpha-thrombin complexed with active site-directed diamino benzo[b]thiophene derivatives: a binding mode for a structurally novel class of inhibitors.
Protein Sci., 9, 2000
|
|
1CUA
 
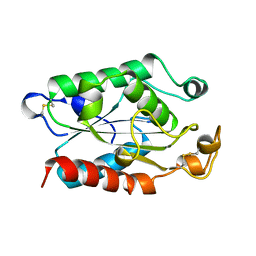 | | CUTINASE, N172K MUTANT | | Descriptor: | CUTINASE | | Authors: | Longhi, S, Cambillau, C. | | Deposit date: | 1995-11-16 | | Release date: | 1996-07-11 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Dynamics of Fusarium solani cutinase investigated through structural comparison among different crystal forms of its variants.
Proteins, 26, 1996
|
|
1CUI
 
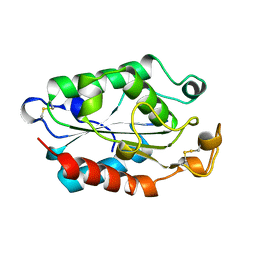 | | CUTINASE, S120A MUTANT | | Descriptor: | CUTINASE | | Authors: | Martinez, C, Cambillau, C. | | Deposit date: | 1995-11-16 | | Release date: | 1996-07-11 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Dynamics of Fusarium solani cutinase investigated through structural comparison among different crystal forms of its variants.
Proteins, 26, 1996
|
|
1CP7
 
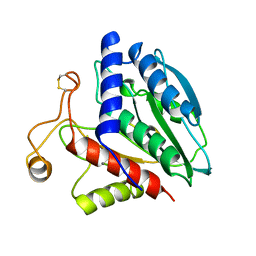 | | AMINOPEPTIDASE FROM STREPTOMYCES GRISEUS | | Descriptor: | AMINOPEPTIDASE, CALCIUM ION, ZINC ION | | Authors: | Gilboa, R, Greenblatt, H.M, Perach, M, Spungin-Bialik, A, Lessel, U, Schomburg, D, Blumberg, S, Shoham, G. | | Deposit date: | 1999-06-10 | | Release date: | 2000-05-03 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Interactions of Streptomyces griseus aminopeptidase with a methionine product analogue: a structural study at 1.53 A resolution.
Acta Crystallogr.,Sect.D, 56, 2000
|
|
1CJ1
 
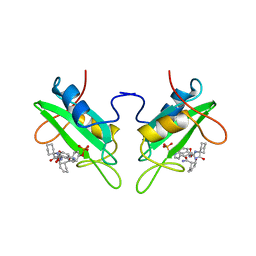 | | GROWTH FACTOR RECEPTOR BINDING PROTEIN SH2 DOMAIN (HUMAN) COMPLEXED WITH A PHOSPHOTYROSYL DERIVATIVE | | Descriptor: | PROTEIN (GROWTH FACTOR RECEPTOR-BOUND PROTEIN 2), [1-[1-(6-CARBAMOYL-CYCLOHEX-2-ENYLCARBAMOYL)-CYCLOHEXYLCARBAMOYL]-2-(4-PHOSPHONOOXY-PHENYL)- ETHYL]-CARBAMIC ACID 3-AMINOBENZYLESTER | | Authors: | Rahuel, J. | | Deposit date: | 1999-04-21 | | Release date: | 1999-12-22 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structure-based design, synthesis, and X-ray crystallography of a high-affinity antagonist of the Grb2-SH2 domain containing an asparagine mimetic.
J.Med.Chem., 42, 1999
|
|
1DPO
 
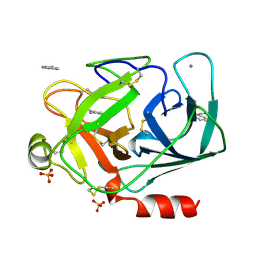 | | STRUCTURE OF RAT TRYPSIN | | Descriptor: | BENZAMIDINE, CALCIUM ION, SULFATE ION, ... | | Authors: | Stroud, R.M. | | Deposit date: | 1997-03-31 | | Release date: | 1997-07-07 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (1.59 Å) | | Cite: | 1.59 A structure of trypsin at 120 K: comparison of low temperature and room temperature structures.
Proteins, 10, 1991
|
|
1A60
 
 | | NMR STRUCTURE OF A CLASSICAL PSEUDOKNOT: INTERPLAY OF SINGLE-AND DOUBLE-STRANDED RNA, 24 STRUCTURES | | Descriptor: | TYMV PSEUDOKNOT | | Authors: | Kolk, M.H, Van Der Graaf, M, Wijmenga, S.S, Pleij, C.W.A, Heus, H.A, Hilbers, C.W. | | Deposit date: | 1998-03-04 | | Release date: | 1998-05-27 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR structure of a classical pseudoknot: interplay of single- and double-stranded RNA.
Science, 280, 1998
|
|
