2WRA
 
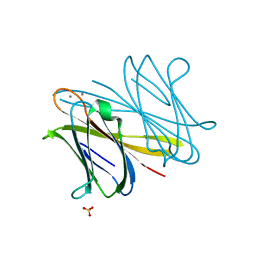 | | BclA lectin from Burkholderia cenocepacia complexed with aMan1(aMan1- 6)-3Man trisaccharide | | Descriptor: | CALCIUM ION, LECTIN, SULFATE ION, ... | | Authors: | Lameignere, E, Shiao, T.C, Roy, R, Wimmerova, M, Dubreuil, F, Varrot, A, Imberty, A. | | Deposit date: | 2009-09-01 | | Release date: | 2009-09-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Structural Basis of the Affinity for Oligomannosides and Analogs Displayed by Bc2L-A, a Burkholderia Cenocepacia Soluble Lectin.
Glycobiology, 20, 2010
|
|
7BIJ
 
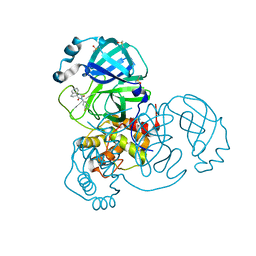 | | Crystal Structure of SARS-CoV-2 main protease (Nsp5) in complex with compound 13 | | Descriptor: | (3~{S})-3'-(5-fluoranylpyridin-3-yl)spiro[1,2-dihydroindene-3,5'-imidazolidine]-2',4'-dione, 3C-like proteinase nsp5, DIMETHYL SULFOXIDE | | Authors: | Talibov, V.O. | | Deposit date: | 2021-01-12 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.47 Å) | | Cite: | Ultralarge Virtual Screening Identifies SARS-CoV-2 Main Protease Inhibitors with Broad-Spectrum Activity against Coronaviruses.
J.Am.Chem.Soc., 144, 2022
|
|
2VCQ
 
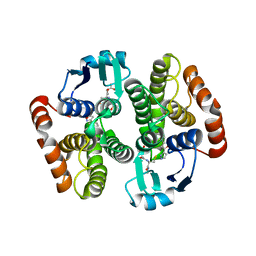 | | Complex structure of prostaglandin D2 synthase at 1.95A. | | Descriptor: | 3-phenyl-5-(1H-pyrazol-3-yl)isoxazole, GLUTATHIONE, GLUTATHIONE-REQUIRING PROSTAGLANDIN D SYNTHASE | | Authors: | Hohwy, M, Spadola, L, Lundquist, B, von Wachenfeldt, K, Persdotter, S, Hawtin, P, Dahmen, J, Groth-Clausen, I, Folmer, R.H.A, Edman, K. | | Deposit date: | 2007-09-26 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Novel Prostaglandin D Synthase Inhibitors Generated by Fragment-Based Drug Design.
J.Med.Chem., 51, 2008
|
|
7DOO
 
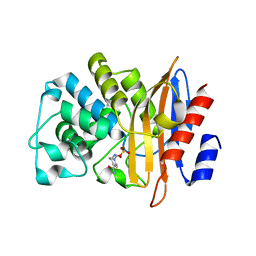 | | Crystal Structure of PenA beta-Lactamase-Avibactam Complex | | Descriptor: | (2S,5R)-1-formyl-5-[(sulfooxy)amino]piperidine-2-carboxamide, Beta-lactamase | | Authors: | Nukaga, M, Papp-Wallace, K.M, Bonomo, R.A. | | Deposit date: | 2020-12-15 | | Release date: | 2020-12-23 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Overcoming an Extremely Drug Resistant (XDR) Pathogen: Avibactam Restores Susceptibility to Ceftazidime for Burkholderia cepacia Complex Isolates from Cystic Fibrosis Patients.
Acs Infect Dis., 3, 2017
|
|
2VCX
 
 | | Complex structure of prostaglandin D2 synthase at 2.1A. | | Descriptor: | GLUTATHIONE, GLUTATHIONE-REQUIRING PROSTAGLANDIN D SYNTHASE, MAGNESIUM ION, ... | | Authors: | Hohwy, M, Spadola, L, Lundquist, B, von Wachenfeldt, K, Persdotter, S, Hawtin, P, Dahmen, J, Groth-Clausen, I, Folmer, R.H.A, Edman, K. | | Deposit date: | 2007-09-27 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Novel Prostaglandin D Synthase Inhibitors Generated by Fragment-Based Drug Design.
J.Med.Chem., 51, 2008
|
|
8HRD
 
 | | Crystal structure of the receptor binding domain of SARS-CoV-2 Delta variant in complex with IMCAS74 Fab and W14 Fab | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, IMCAS74 Fab heavy chain, IMCAS74 Fab light chain, ... | | Authors: | Zhao, R.C, Wu, L.L, Han, P. | | Deposit date: | 2022-12-15 | | Release date: | 2023-12-20 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Defining a de novo non-RBM antibody as RBD-8 and its synergistic rescue of immune-evaded antibodies to neutralize Omicron SARS-CoV-2.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
2VCW
 
 | | Complex structure of prostaglandin D2 synthase at 1.95A. | | Descriptor: | 1-PHENYL-1H-PYRAZOLE-4-CARBOXYLIC ACID, GLUTATHIONE, GLUTATHIONE-REQUIRING PROSTAGLANDIN D SYNTHASE | | Authors: | Hohwy, M, Spadola, L, Lundquist, B, von Wachenfeldt, K, Persdotter, S, Hawtin, P, Dahmen, J, Groth-Clausen, I, Folmer, R.H.A, Edman, K. | | Deposit date: | 2007-09-27 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Novel Prostaglandin D Synthase Inhibitors Generated by Fragment-Based Drug Design.
J.Med.Chem., 51, 2008
|
|
2VD1
 
 | | Complex structure of prostaglandin D2 synthase at 2.25A. | | Descriptor: | 4-{[4-(4-fluoro-3-methylphenyl)-1,3-thiazol-2-yl]amino}-2-hydroxybenzoic acid, GLUTATHIONE, GLUTATHIONE-REQUIRING PROSTAGLANDIN D SYNTHASE, ... | | Authors: | Hohwy, M, Spadola, L, Lundquist, B, von Wachenfeldt, K, Persdotter, S, Hawtin, P, Dahmen, J, Groth-Clausen, I, Folmer, R.H.A, Edman, K. | | Deposit date: | 2007-09-28 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Novel Prostaglandin D Synthase Inhibitors Generated by Fragment-Based Drug Design.
J.Med.Chem., 51, 2008
|
|
2VCZ
 
 | | Complex structure of prostaglandin D2 synthase at 1.95A. | | Descriptor: | 3-(4-nitrophenyl)-1H-pyrazole, GLUTATHIONE, GLUTATHIONE-REQUIRING PROSTAGLANDIN D SYNTHASE | | Authors: | Hohwy, M, Spadola, L, Lundquist, B, von Wachenfeldt, K, Persdotter, S, Hawtin, P, Dahmen, J, Groth-Clausen, I, Folmer, R.H.A, Edman, K. | | Deposit date: | 2007-09-28 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Novel Prostaglandin D Synthase Inhibitors Generated by Fragment-Based Drug Design.
J.Med.Chem., 51, 2008
|
|
2VD0
 
 | | Complex structure of prostaglandin D2 synthase at 2.2A. | | Descriptor: | 2-{[(2E)-3-(3,4-dimethoxyphenyl)prop-2-enoyl]amino}benzoic acid, GLUTATHIONE, GLUTATHIONE-REQUIRING PROSTAGLANDIN D SYNTHASE, ... | | Authors: | Hohwy, M, Spadola, L, Lundquist, B, von Wachenfeldt, K, Persdotter, S, Hawtin, P, Dahmen, J, Groth-Clausen, I, Folmer, R.H.A, Edman, K. | | Deposit date: | 2007-09-28 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Novel Prostaglandin D Synthase Inhibitors Generated by Fragment-Based Drug Design.
J.Med.Chem., 51, 2008
|
|
6FR2
 
 | | Soluble epoxide hydrolase in complex with LK864 | | Descriptor: | 1-[(4~{S})-9-propan-2-ylsulfonyl-1-oxa-9-azaspiro[5.5]undecan-4-yl]-3-[[4-(trifluoromethyloxy)phenyl]methyl]urea, Bifunctional epoxide hydrolase 2, MAGNESIUM ION | | Authors: | Kramer, J.S, Pogoryelov, D, Krasavin, M, Proschak, E. | | Deposit date: | 2018-02-15 | | Release date: | 2018-09-12 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.262 Å) | | Cite: | Discovery of polar spirocyclic orally bioavailable urea inhibitors of soluble epoxide hydrolase.
Bioorg. Chem., 80, 2018
|
|
4HR3
 
 | |
7O46
 
 | | Crystal Structure of SARS-CoV-2 main protease (Nsp5) in complex with compound 17 | | Descriptor: | 2-cyclobutyl-7-isoquinolin-4-yl-5,7-diazaspiro[3.4]octane-6,8-dione, 3C-like proteinase nsp5 | | Authors: | Talibov, V.O. | | Deposit date: | 2021-04-05 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.23 Å) | | Cite: | Ultralarge Virtual Screening Identifies SARS-CoV-2 Main Protease Inhibitors with Broad-Spectrum Activity against Coronaviruses.
J.Am.Chem.Soc., 144, 2022
|
|
7QBB
 
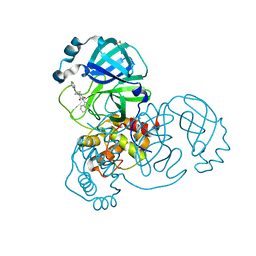 | | Crystal Structure of SARS-CoV-2 main protease (Nsp5) in complex with compound 18 | | Descriptor: | 3C-like proteinase nsp5, 7-isoquinolin-4-yl-2-phenyl-5,7-diazaspiro[3.4]octane-6,8-dione, DIMETHYL SULFOXIDE | | Authors: | Talibov, V.O. | | Deposit date: | 2021-11-18 | | Release date: | 2022-02-23 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Ultralarge Virtual Screening Identifies SARS-CoV-2 Main Protease Inhibitors with Broad-Spectrum Activity against Coronaviruses.
J.Am.Chem.Soc., 144, 2022
|
|
4PUB
 
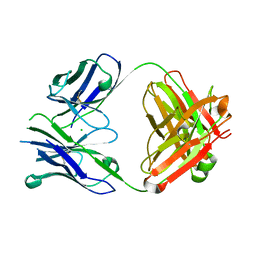 | | Crystal structure of Fab DX-2930 | | Descriptor: | CHLORIDE ION, DX-2930 HEAVY CHAIN, DX-2930 LIGHT CHAIN | | Authors: | Abendroth, J, Edwards, T.E, Nixon, A, Ladner, R. | | Deposit date: | 2014-03-12 | | Release date: | 2014-07-09 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Inhibition of plasma kallikrein by a highly specific active site blocking antibody.
J.Biol.Chem., 289, 2014
|
|
6H6R
 
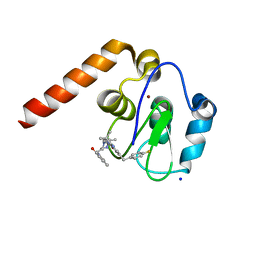 | | Fragment Derived XIAP inhibitor | | Descriptor: | 2-[[(2~{R},5~{R})-1-[2-[6-[(4-fluorophenyl)methyl]-3,3-dimethyl-2~{H}-pyrrolo[3,2-b]pyridin-1-yl]-2-oxidanylidene-ethyl]-5-methyl-piperazin-2-yl]methyl]-3~{H}-isoindol-1-one, E3 ubiquitin-protein ligase XIAP, SODIUM ION, ... | | Authors: | Williams, P.A. | | Deposit date: | 2018-07-30 | | Release date: | 2018-08-22 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.03 Å) | | Cite: | A Fragment-Derived Clinical Candidate for Antagonism of X-Linked and Cellular Inhibitor of Apoptosis Proteins: 1-(6-[(4-Fluorophenyl)methyl]-5-(hydroxymethyl)-3,3-dimethyl-1 H,2 H,3 H-pyrrolo[3,2- b]pyridin-1-yl)-2-[(2 R,5 R)-5-methyl-2-([(3R)-3-methylmorpholin-4-yl]methyl)piperazin-1-yl]ethan-1-one (ASTX660).
J. Med. Chem., 61, 2018
|
|
4HS3
 
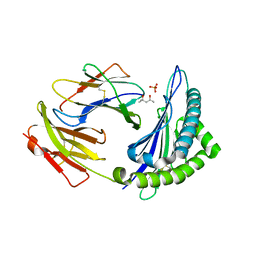 | | Crystal structure of H-2Kb with a disulfide stabilized F pocket in complex with the LCMV derived peptide GP34 | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, Beta-2-microglobulin, Envelope glycoprotein, ... | | Authors: | Uchtenhagen, H, Boulanger, B, Hein, Z, Abualrous, E.T, Zacharias, M, Werner, J, Elliott, T, Springer, S, Achour, A. | | Deposit date: | 2012-10-29 | | Release date: | 2014-05-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Peptide-independent stabilization of MHC class I molecules breaches cellular quality control.
J.Cell.Sci., 127, 2014
|
|
7OO2
 
 | | Crystal structure of an antibody targeting the capsular polysaccharide of serogroup X Neisseria meningitidis (MenX) | | Descriptor: | THIOCYANATE ION, anti-MenX Fab heavy chain, anti-MenX Fab light chain | | Authors: | Pietri, G.P, de Ruyck, J, Lenac, T, Adamo, R, Bouckaert, J. | | Deposit date: | 2021-05-26 | | Release date: | 2021-10-06 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.16 Å) | | Cite: | Elucidating the Structural and Minimal Protective Epitope of the Serogroup X Meningococcal Capsular Polysaccharide.
Front Mol Biosci, 8, 2021
|
|
4IV6
 
 | |
6H6Q
 
 | | Fragment Derived XIAP inhibitor | | Descriptor: | 2-[(2~{R},5~{R})-2-[[(3~{R},5~{R})-3,5-dimethylmorpholin-4-yl]methyl]-5-methyl-piperazin-1-yl]-1-[6-[(~{S})-(4-fluorophenyl)-oxidanyl-methyl]-3,3-dimethyl-2~{H}-pyrrolo[3,2-b]pyridin-1-yl]ethanone, E3 ubiquitin-protein ligase XIAP, ZINC ION | | Authors: | Williams, P.A. | | Deposit date: | 2018-07-30 | | Release date: | 2018-08-22 | | Last modified: | 2024-05-15 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | A Fragment-Derived Clinical Candidate for Antagonism of X-Linked and Cellular Inhibitor of Apoptosis Proteins: 1-(6-[(4-Fluorophenyl)methyl]-5-(hydroxymethyl)-3,3-dimethyl-1 H,2 H,3 H-pyrrolo[3,2- b]pyridin-1-yl)-2-[(2 R,5 R)-5-methyl-2-([(3R)-3-methylmorpholin-4-yl]methyl)piperazin-1-yl]ethan-1-one (ASTX660).
J. Med. Chem., 61, 2018
|
|
4IZ9
 
 | |
4J3J
 
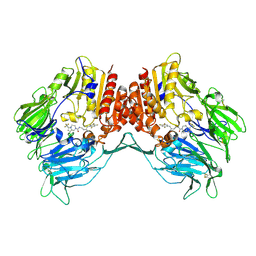 | | Crystal Structure of DPP-IV with Compound C3 | | Descriptor: | Dipeptidyl peptidase 4, N-[(3R)-3-amino-4-(2,4,5-trifluorophenyl)butyl]-6-(trifluoromethyl)-3,4-dihydropyrrolo[1,2-a]pyrazine-2(1H)-carboxamide | | Authors: | Xiong, B, Zhu, L.R, Chen, D.Q, Zhao, Y.L, Jiang, F, Shen, J.K. | | Deposit date: | 2013-02-05 | | Release date: | 2014-02-05 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Design and synthesis of 4-(2,4,5-trifluorophenyl)butane-1,3-diamines as dipeptidyl peptidase IV inhibitors
Chemmedchem, 8, 2013
|
|
7P3I
 
 | | Crystal structure of human CD40/TNFRSF5 in complex with the anti-CD40 DARPin protein | | Descriptor: | Darpin, SODIUM ION, Tumor necrosis factor receptor superfamily member 5 | | Authors: | Malvezzi, F, Mangold, S, Hospodarsch, T, Reichen, C, Iss, C, Lammens, A, Krapp, S, Domke, C. | | Deposit date: | 2021-07-07 | | Release date: | 2022-04-06 | | Last modified: | 2022-05-11 | | Method: | X-RAY DIFFRACTION (2.29 Å) | | Cite: | A Multispecific Anti-CD40 DARPin Construct Induces Tumor-Selective CD40 Activation and Tumor Regression.
Cancer Immunol Res, 10, 2022
|
|
7R98
 
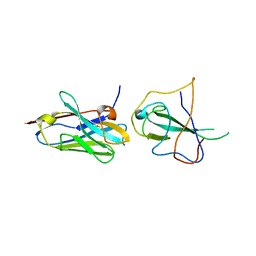 | |
8B48
 
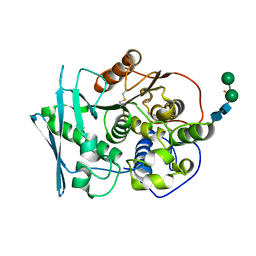 | | Structure of Lentithecium fluviatile carbohydrate esterase from the CE15 family (LfCE15C) | | Descriptor: | Carbohydrate esterase family 15 protein, FORMIC ACID, alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Scholzen, K, Mazurkewich, S, Poulsen, J.C.N, Larsbrink, J, Lo Leggio, L. | | Deposit date: | 2022-09-20 | | Release date: | 2023-06-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Structural and functional investigation of a fungal member of carbohydrate esterase family 15 with potential specificity for rare xylans.
Acta Crystallogr D Struct Biol, 79, 2023
|
|
