6K83
 
 | | Structure of RGLG1 mutant-D207G | | Descriptor: | CALCIUM ION, E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.39 Å) | | Cite: | RGLG1 mutant-D207G
To Be Published
|
|
6K86
 
 | | The closed state of RGLG1 mutant-E378A | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION, SODIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.593 Å) | | Cite: | The closed state of RGLG1 mutant-E378A
To Be Published
|
|
6K8B
 
 | | The open state of RGLG1 VWA domain | | Descriptor: | CALCIUM ION, E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (2.21 Å) | | Cite: | The open state of RGLG1 VWA domain
To Be Published
|
|
6K82
 
 | | RGLG1 mutant-D338A E378A | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION, SODIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.402 Å) | | Cite: | RGLG1 mutant-D338A E378A
To Be Published
|
|
6K89
 
 | | The closed state of RGLG1 VWA domain | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION, SODIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.689 Å) | | Cite: | The closed state of RGLG1 VWA domain
To Be Published
|
|
6K85
 
 | | The closed state of RGLG1 mutant-D338A | | Descriptor: | E3 ubiquitin-protein ligase RGLG1, MAGNESIUM ION | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.606 Å) | | Cite: | The closed state of RGLG1 mutant-D338A
To Be Published
|
|
6K8A
 
 | |
6K88
 
 | | RGLG1 MIDAS binds calcium ion | | Descriptor: | CALCIUM ION, E3 ubiquitin-protein ligase RGLG1 | | Authors: | Wang, Q, Wu, Y. | | Deposit date: | 2019-06-11 | | Release date: | 2020-06-24 | | Last modified: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.788 Å) | | Cite: | RGLG1 MIDAS binds calcium ion
To Be Published
|
|
6LUO
 
 | | Structure of nurse shark beta-2-microglobulin | | Descriptor: | Beta-2-microglobulin | | Authors: | Xia, C, Wu, Y. | | Deposit date: | 2020-01-30 | | Release date: | 2021-04-28 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.302 Å) | | Cite: | The Structure of a Peptide-Loaded Shark MHC Class I Molecule Reveals Features of the Binding between beta 2 -Microglobulin and H Chain Conserved in Evolution.
J Immunol., 2021
|
|
6MHN
 
 | | Photoactive Yellow Protein with covalently bound 3-chloro-4-hydroxycinnamic acid chromophore | | Descriptor: | (2E)-3-(3-chloro-4-hydroxyphenyl)prop-2-enoic acid, Photoactive yellow protein | | Authors: | Thomson, B.D, Both, J, Wu, Y, Parrish, R.M, Martinez, T, Boxer, S.G. | | Deposit date: | 2018-09-18 | | Release date: | 2019-05-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Perturbation of Short Hydrogen Bonds in Photoactive Yellow Protein via Noncanonical Amino Acid Incorporation.
J.Phys.Chem.B, 123, 2019
|
|
6MHI
 
 | | Photoactive Yellow Protein with covalently bound 3,5-dichloro-4-hydroxycinnamic acid chromophore | | Descriptor: | (2E)-3-(3,5-dichloro-4-hydroxyphenyl)prop-2-enoic acid, Photoactive yellow protein | | Authors: | Thomson, B.D, Both, J, Wu, Y, Parrish, R.M, Martinez, T, Boxer, S.G. | | Deposit date: | 2018-09-18 | | Release date: | 2019-05-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Perturbation of Short Hydrogen Bonds in Photoactive Yellow Protein via Noncanonical Amino Acid Incorporation.
J.Phys.Chem.B, 123, 2019
|
|
6MKT
 
 | | Photoactive Yellow Protein with 3-chlorotyrosine substituted at position 42 | | Descriptor: | 4'-HYDROXYCINNAMIC ACID, Photoactive yellow protein | | Authors: | Thomson, B.D, Both, J, Wu, Y, Parrish, R.M, Martinez, T, Boxer, S.G. | | Deposit date: | 2018-09-26 | | Release date: | 2019-05-29 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Perturbation of Short Hydrogen Bonds in Photoactive Yellow Protein via Noncanonical Amino Acid Incorporation.
J.Phys.Chem.B, 123, 2019
|
|
7YR7
 
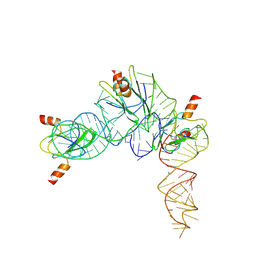 | | Cryo-EM structure of Pseudomonas aeruginosa RsmZ RNA in complex with three RsmA protein dimers | | Descriptor: | RsmZ RNA (118-MER), Translational regulator CsrA | | Authors: | Jia, X, Pan, Z, Yuan, Y, Luo, B, Luo, Y, Mukherjee, S, Jia, G, Liu, L, Ling, X, Yang, X, Wu, Y, Liu, T, Miao, Z, Wei, X, Bujnicki, J.M, Zhao, K, Su, Z. | | Deposit date: | 2022-08-09 | | Release date: | 2023-05-17 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (3.8 Å) | | Cite: | Structural basis of sRNA RsmZ regulation of Pseudomonas aeruginosa virulence.
Cell Res., 33, 2023
|
|
7YR6
 
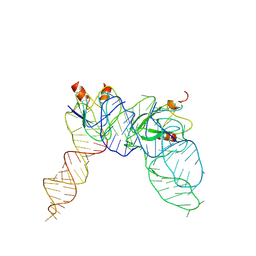 | | Cryo-EM structure of Pseudomonas aeruginosa RsmZ RNA in complex with two RsmA protein dimers | | Descriptor: | RsmZ RNA, Translational regulator CsrA | | Authors: | Jia, X, Pan, Z, Yuan, Y, Luo, B, Luo, Y, Mukherjee, S, Jia, G, Ling, X, Yang, X, Wu, Y, Liu, T, Wei, X, Bujnick, J.M, Zhao, K, Su, Z. | | Deposit date: | 2022-08-09 | | Release date: | 2023-05-17 | | Last modified: | 2024-07-03 | | Method: | ELECTRON MICROSCOPY (4.8 Å) | | Cite: | Structural basis of sRNA RsmZ regulation of Pseudomonas aeruginosa virulence.
Cell Res., 33, 2023
|
|
2JS4
 
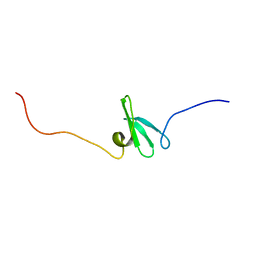 | | Solution NMR Structure of Bordetella bronchiseptica protein BB2007. Northeast Structural Genomics Consortium target BoR54 | | Descriptor: | UPF0434 protein BB2007 | | Authors: | Eletsky, A, Sukumaran, D, Wu, Y, Singarapu, K, Parish, D, Xu, D, Wang, D, Nwosu, C, Cunningham, K, Xiao, R, Liu, J, Baran, M.C, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2007-06-29 | | Release date: | 2007-07-10 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | NMR Solution Structure of Bordetella bronchiseptica protein BB2007.
To be Published
|
|
