5UAG
 
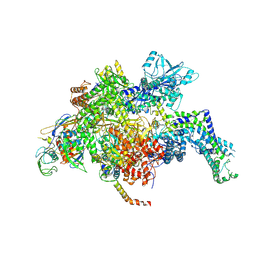 | | Escherichia coli RNA polymerase mutant - RpoB D516V | | Descriptor: | DNA-directed RNA polymerase subunit alpha, DNA-directed RNA polymerase subunit beta, DNA-directed RNA polymerase subunit beta', ... | | Authors: | Molodtsov, V, Scharf, N.T, Stefan, M.A, Garcia, G.A, Murakami, K.S. | | Deposit date: | 2016-12-19 | | Release date: | 2017-02-08 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (3.399 Å) | | Cite: | Structural basis for rifamycin resistance of bacterial RNA polymerase by the three most clinically important RpoB mutations found in Mycobacterium tuberculosis.
Mol. Microbiol., 103, 2017
|
|
1A2D
 
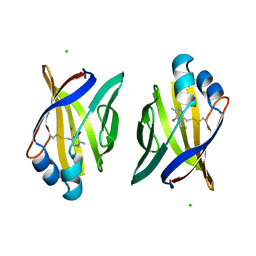 | | PYRIDOXAMINE MODIFIED MURINE ADIPOCYTE LIPID BINDING PROTEIN | | Descriptor: | ADIPOCYTE LIPID BINDING PROTEIN, CHLORIDE ION | | Authors: | Ory, J, Mazhary, A, Kuang, H, Davies, R, Distefano, M, Banaszak, L. | | Deposit date: | 1997-12-29 | | Release date: | 1998-07-01 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural characterization of two synthetic catalysts based on adipocyte lipid-binding protein.
Protein Eng., 11, 1998
|
|
1A18
 
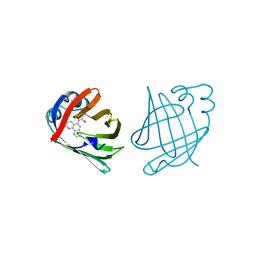 | | PHENANTHROLINE MODIFIED MURINE ADIPOCYTE LIPID BINDING PROTEIN | | Descriptor: | ADIPOCYTE LIPID BINDING PROTEIN | | Authors: | Ory, J, Mazhary, A, Kuang, H, Davies, R, Distefano, M, Banaszak, L. | | Deposit date: | 1997-12-23 | | Release date: | 1998-07-01 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Structural characterization of two synthetic catalysts based on adipocyte lipid-binding protein.
Protein Eng., 11, 1998
|
|
2MYG
 
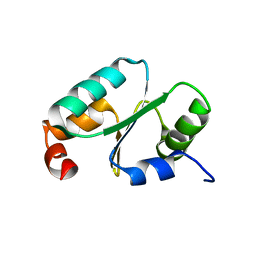 | | Solution structure of the dithiolic glutaredoxin 2-C-Grx1 from the pathogen Trypanosoma brucei brucei | | Descriptor: | Dithiol glutaredoxin 1 | | Authors: | Sturlese, M, Stefani, M, Manta, B, Mammi, S, Comini, M, Bellanda, M. | | Deposit date: | 2015-01-22 | | Release date: | 2016-02-10 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the dithiolic glutaredoxin 2-C-Grx1 from the pathogen Trypanosoma brucei brucei
To be Published
|
|
2J4H
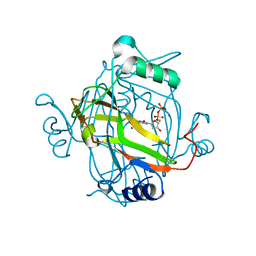 
 | | Crystal structure of a H121A Escherichia coli dCTP deaminase mutant enzyme | | Descriptor: | DEOXYCYTIDINE DIPHOSPHATE, DEOXYCYTIDINE TRIPHOSPHATE DEAMINASE, MAGNESIUM ION | | Authors: | Johansson, E, Thymark, M, Bynck, J.H, Fanoe, M, Larsen, S, Willemoes, M. | | Deposit date: | 2006-08-31 | | Release date: | 2007-08-07 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Regulation of Dctp Deaminase from Escherichia Coli by Nonallosteric Dttp Binding to an Inactive Form of the Enzyme
FEBS J., 274, 2007
|
|
3KSL
 
 | | Structure of FPT bound to DATFP-DH-GPP | | Descriptor: | (2S,6E)-8-{[(R)-hydroxy(phosphonooxy)phosphoryl]oxy}-2,6-dimethyloct-6-en-1-yl (2S)-3,3,3-trifluoro-2-hydrazinopropanoate, Farnesyltransferase, CAAX box, ... | | Authors: | Hovlid, M.L, Edelstein, R.L, Henry, O, Ochocki, J, DeGraw, A, Lenevich, S, Talbot, T, Young, V, Hruza, A.W, Lopez-Gallego, F, Labello, N.P, Strickland, C.L, Schmidt-Dannert, C, Distefano, M.D. | | Deposit date: | 2009-11-23 | | Release date: | 2009-12-22 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Synthesis, properties, and applications of diazotrifluropropanoyl-containing photoactive analogs of farnesyl diphosphate containing modified linkages for enhanced stability.
Chem.Biol.Drug Des., 75, 2010
|
|
1PHR
 
 | | THE CRYSTAL STRUCTURE OF A LOW MOLECULAR PHOSPHOTYROSINE PROTEIN PHOSPHATASE | | Descriptor: | LOW MOLECULAR WEIGHT PHOSPHOTYROSINE PROTEIN PHOSPHATASE, SULFATE ION | | Authors: | Su, X.-D, Taddei, N, Stefani, M, Ramponi, G, Nordlund, P. | | Deposit date: | 1994-07-05 | | Release date: | 1995-07-31 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The crystal structure of a low-molecular-weight phosphotyrosine protein phosphatase.
Nature, 370, 1994
|
|
1O1R
 
 | | Structure of FPT bound to GGPP | | Descriptor: | GERANYLGERANYL DIPHOSPHATE, Protein farnesyltransferase alpha subunit, Protein farnesyltransferase beta subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights
into Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
1O1S
 
 | | Structure of FPT bound to isoprenoid analog 3b | | Descriptor: | (2E,6E)-8-[(3-BENZOYLBENZYL)OXY]-3,7-DIMETHYLOCTA-2,6-DIENYL TRIHYDROGEN DIPHOSPHATE, Protein farnesyltransferase alpha subunit, Protein farnesyltransferase beta subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights into
Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
1O1T
 
 | | Structure of FPT bound to the CVIM-FPP product | | Descriptor: | MAGNESIUM ION, N-ACETYL-S-[(2E,6E)-3,7,11-TRIMETHYLDODECA-2,6,10-TRIENYL]-L-CYSTEINYL-D-VALYL-L-ISOLEUCYL-L-METHIONINE, Protein farnesyltransferase alpha subunit, ... | | Authors: | Turek-Etienne, T.C, Strickland, C.L, Distefano, M.D. | | Deposit date: | 2003-01-14 | | Release date: | 2003-05-06 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Biochemical and Structural Studies with Prenyl Diphosphate Analogues Provide Insights into
Isoprenoid Recognition by Protein Farnesyl Transferase
Biochemistry, 42, 2003
|
|
1Z6F
 
 | | Crystal structure of penicillin-binding protein 5 from E. coli in complex with a boronic acid inhibitor | | Descriptor: | GLYCEROL, N1-[(1R)-1-(DIHYDROXYBORYL)ETHYL]-N2-[(TERT-BUTOXYCARBONYL)-D-GAMMA-GLUTAMYL]-N6-[(BENZYLOXY)CARBONYL-L-LYSINAMIDE, Penicillin-binding protein 5 | | Authors: | Nicola, G, Peddi, S, Stefanova, M, Nicholas, R.A, Gutheil, W.G, Davies, C. | | Deposit date: | 2005-03-22 | | Release date: | 2005-06-21 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal Structure of Escherichia coli Penicillin-Binding Protein 5 Bound to a Tripeptide Boronic Acid Inhibitor: A Role for Ser-110 in Deacylation.
Biochemistry, 44, 2005
|
|
