3NNJ
 
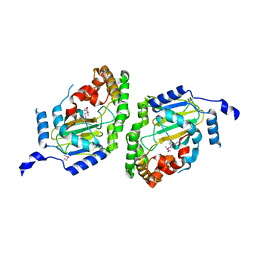 | | Halogenase domain from CurA module (apo Hal) | | Descriptor: | CurA | | Authors: | Khare, D, Smith, J.L. | | Deposit date: | 2010-06-23 | | Release date: | 2010-07-28 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.601 Å) | | Cite: | Conformational switch triggered by alpha-ketoglutarate in a halogenase of curacin A biosynthesis
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2J4B
 
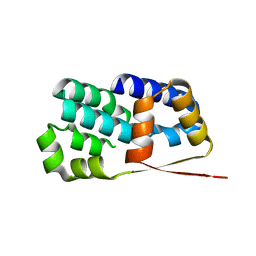 | | Crystal structure of Encephalitozoon cuniculi TAF5 N-terminal domain | | Descriptor: | TRANSCRIPTION INITIATION FACTOR TFIID SUBUNIT 72/90-100 KDA | | Authors: | Romier, C, James, N, Birck, C, Cavarelli, J, Vivares, C, Collart, M.A, Moras, D. | | Deposit date: | 2006-08-28 | | Release date: | 2007-04-10 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal Structure, Biochemical and Genetic Characterization of Yeast and E. Cuniculi Taf(II)5 N-Terminal Domain: Implications for TFIID Assembly.
J.Mol.Biol., 368, 2007
|
|
2IXO
 
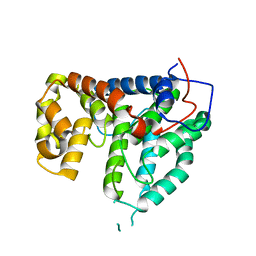 | | CRYSTAL STRUCTURE OF THE PP2A PHOSPHATASE ACTIVATOR Ypa1 PTPA1 | | Descriptor: | SERINE/THREONINE-PROTEIN PHOSPHATASE 2A ACTIVATOR 1 | | Authors: | Leulliot, N, Vicentini, G, Jordens, J, Quevillon-Cheruel, S, Schiltz, M, Barford, D, Van Tilbeurgh, H, Goris, J. | | Deposit date: | 2006-07-09 | | Release date: | 2006-07-31 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal Structure of the Pp2A Phosphatase Activator: Implications for its Pp2A-Specific Ppiase Activity.
Mol.Cell, 23, 2006
|
|
2IZY
 
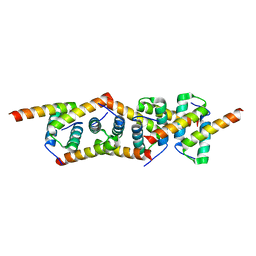 | | Molecular Basis of AKAP Specificity for PKA Regulatory Subunits | | Descriptor: | CAMP-DEPENDENT PROTEIN KINASE REGULATORY SUBUNIT II | | Authors: | Gold, M.G, Lygren, B, Dokurno, P, Hoshi, N, McConnachie, G, Tasken, K, Carlson, C.R, Scott, J.D, Barford, D. | | Deposit date: | 2006-07-27 | | Release date: | 2006-11-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular Basis of Akap Specificity for Pka Regululatory Subunits
Mol.Cell, 24, 2006
|
|
2KC0
 
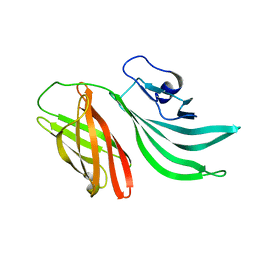 | | Solution structure of the factor H binding protein | | Descriptor: | lipoprotein | | Authors: | Cantini, F, Veggi, D, Dragonetti, S, Savino, S, Scarselli, M, Romagnoli, G, Pizza, M, Banci, L, Rappuoli, R. | | Deposit date: | 2008-12-13 | | Release date: | 2009-02-17 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of the Factor H-binding Protein, a Survival Factor and Protective Antigen of Neisseria meningitidis
J.Biol.Chem., 284, 2009
|
|
2KBE
 
 | |
3O4P
 
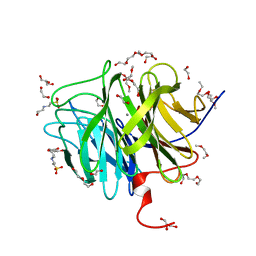 | | DFPase at 0.85 Angstrom resolution (H atoms included) | | Descriptor: | 1,2-DIMETHOXYETHANE, 1,2-ETHANEDIOL, 1-ETHOXY-2-(2-METHOXYETHOXY)ETHANE, ... | | Authors: | Liebschner, D, Elias, M, Koepke, J, Lecomte, C, Guillot, B, Jelsch, C, Chabriere, E. | | Deposit date: | 2010-07-27 | | Release date: | 2011-08-17 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (0.85 Å) | | Cite: | Hydrogen atoms in protein structures: high-resolution X-ray diffraction structure of the DFPase.
BMC Res Notes, 6, 2013
|
|
2KL8
 
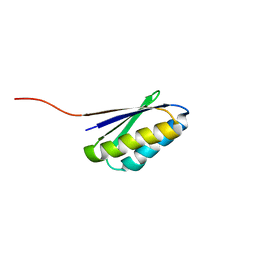 | | Solution NMR Structure of de novo designed ferredoxin-like fold protein, Northeast Structural Genomics Consortium Target OR15 | | Descriptor: | OR15 | | Authors: | Liu, G, Koga, N, Jiang, M, Koga, R, Xiao, R, Ciccosanti, C, Baker, D, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-30 | | Release date: | 2009-07-28 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Principles for designing ideal protein structures.
Nature, 491, 2012
|
|
3NNL
 
 | | Halogenase domain from CurA module (crystal form III) | | Descriptor: | 2-OXOGLUTARIC ACID, CHLORIDE ION, CurA, ... | | Authors: | Khare, D, Smith, J.L. | | Deposit date: | 2010-06-23 | | Release date: | 2010-07-28 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.883 Å) | | Cite: | Conformational switch triggered by alpha-ketoglutarate in a halogenase of curacin A biosynthesis
Proc.Natl.Acad.Sci.USA, 107, 2010
|
|
2KBY
 
 | | The Tetramerization Domain of Human p73 | | Descriptor: | Tumor protein p73 | | Authors: | Coutandin, D, Ikeya, T, Loehr, F, Guntert, P, Ou, H.D, Doetsch, V. | | Deposit date: | 2008-12-12 | | Release date: | 2009-09-29 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Conformational stability and activity of p73 require a second helix in the tetramerization domain.
Cell Death Differ., 16, 2009
|
|
2KE6
 
 | |
2K69
 
 | |
2KHH
 
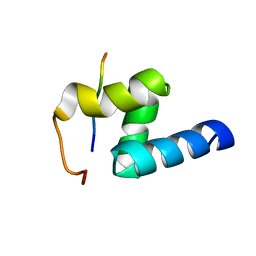 | | Structural requirements for the UBA domain of the mRNA export factor Mex67 to bind its specific targets, the transcription elongation Tho complex component Hpr1 and nucleoporin FxFG repeats | | Descriptor: | FxFG, mRNA export factor MEX67 | | Authors: | Hobeika, M, Brockmann, C, Gruessing, F, Neuhaus, D, Divita, G, Stewart, M, Dargemont, C. | | Deposit date: | 2009-04-06 | | Release date: | 2009-05-12 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Structural requirements for the ubiquitin-associated domain of the mRNA export factor Mex67 to bind its specific targets, the transcription elongation THO complex component Hpr1 and nucleoporin FXFG repeats
J.Biol.Chem., 284, 2009
|
|
3NY9
 
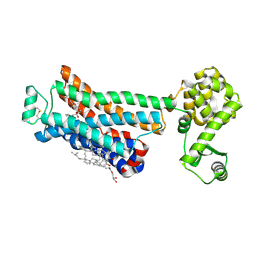 | | Crystal structure of the human beta2 adrenergic receptor in complex with a novel inverse agonist | | Descriptor: | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate, Beta-2 adrenergic receptor, Lysozyme, ... | | Authors: | Fenalti, G, Wacker, D, Brown, M.A, Katritch, V, Abagyan, R, Cherezov, V, Stevens, R.C, Accelerated Technologies Center for Gene to 3D Structure (ATCG3D), GPCR Network (GPCR) | | Deposit date: | 2010-07-14 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.84 Å) | | Cite: | Conserved Binding Mode of Human beta(2) Adrenergic Receptor Inverse Agonists and Antagonist Revealed by X-ray Crystallography
J.Am.Chem.Soc., 132, 2010
|
|
2KAN
 
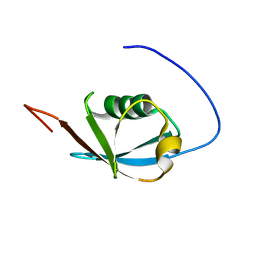 | | Solution NMR structure of ubiquitin-like domain of Arabidopsis thaliana protein At2g32350. Northeast Structural Genomics Consortium target AR3433A | | Descriptor: | uncharacterized protein ar3433a | | Authors: | Eletsky, A, Mills, J.L, Sukumaran, D, Hua, J, Shastry, R, Jiang, M, Ciccosanti, C, Xiao, R, Liu, J, Everret, J.K, Swapna, G.V.T, Acton, T.B, Rost, B, Montelione, G.T, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-11-11 | | Release date: | 2008-11-25 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of ubiquitin-like domain of Arabidopsis thaliana protein At2g32350.
To be Published
|
|
3NV9
 
 | | Crystal structure of Entamoeba histolytica Malic Enzyme | | Descriptor: | ACETATE ION, GLYCEROL, Malic enzyme, ... | | Authors: | Dutta, D, Chakrabarty, A, Ghosh, S.K, Das, A.K. | | Deposit date: | 2010-07-08 | | Release date: | 2011-07-20 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Structural and functional characterization of Entamoeba histolytica Malic enzyme
To be Published
|
|
3O1D
 
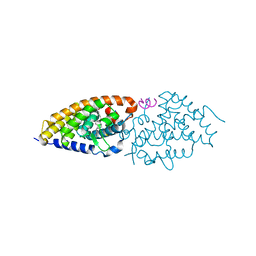 | | Structure-function study of Gemini derivatives with two different side chains at C-20, Gemini-0072 and Gemini-0097. | | Descriptor: | (1R,3R,7E,17beta)-17-[(1S)-6,6,6-trifluoro-5-hydroxy-1-(4-hydroxy-4-methylpentyl)-5-(trifluoromethyl)hex-3-yn-1-yl]-9,10-secoestra-5,7-diene-1,3-diol, Nuclear receptor coactivator 2, Vitamin D3 receptor A | | Authors: | Huet, T, Moras, D, Rochel, N. | | Deposit date: | 2010-07-21 | | Release date: | 2011-07-06 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4007 Å) | | Cite: | Structure-function study of gemini derivatives with two different side chains at C-20, Gemini-0072 and Gemini-0097.
Medchemcomm, 2, 2011
|
|
2KG7
 
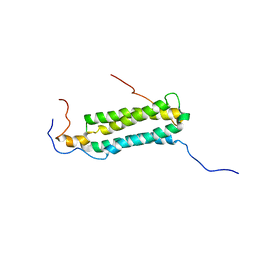 | | Structure and features of the complex formed by the tuberculosis virulence factors Rv0287 and Rv0288 | | Descriptor: | ESAT-6-like protein esxH, Uncharacterized protein esxG (PE family protein) | | Authors: | Ilghari, D, Kirsty, L.L, Waters, L.C, Veverka, V.L, Philip, R.S, Carr, M.D. | | Deposit date: | 2009-03-06 | | Release date: | 2010-03-16 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Solution structure of the M. tuberculosis EsxG-EsxH complex: functional implications and comparisons with other M. tuberculosis Esx family complexes
J.Biol.Chem., 2011
|
|
3O31
 
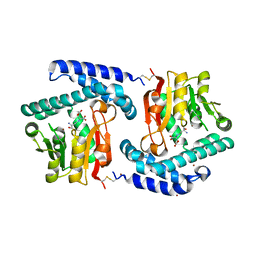 | | E81Q mutant of MtNAS in complex with a reaction intermediate | | Descriptor: | BROMIDE ION, N-[(3S)-3-amino-3-carboxypropyl]-L-glutamic acid, ThermoNicotianamine Synthase | | Authors: | Dreyfus, C, Pignol, D, Arnoux, P. | | Deposit date: | 2010-07-23 | | Release date: | 2011-06-08 | | Last modified: | 2014-11-12 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The crystallographic structure of thermoNicotianamine synthase with a synthetic reaction intermediate highlights the sequential processing mechanism.
Chem.Commun.(Camb.), 47, 2011
|
|
2K68
 
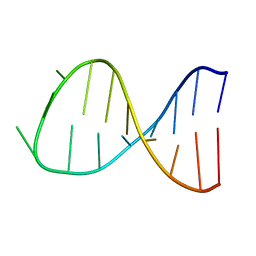 | |
2K7Y
 
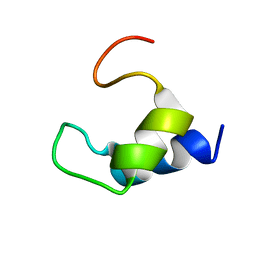 | |
2K5H
 
 | | Solution NMR structure of protein encoded by MTH693 from Methanobacterium thermoautotrophicum: Northeast Structural Genomics Consortium target tt824a | | Descriptor: | Conserved protein | | Authors: | Wu, Y, Singarapu, K, Semesi, A, Sukumaran, D, Yee, A, Garcia, M, Arrowsmith, C, Szyperski, T, Montelione, G.T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-27 | | Release date: | 2008-08-19 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of protein encoded by MTH693 from Methanobacterium thermoautotrophicum: Northeast Structural Genomics Consortium target tt824a
To be Published
|
|
2KKO
 
 | | Solution NMR structure of the homodimeric winged helix-turn-helix DNA-binding domain (fragment 1-100) Mb0332 from Mycobacterium bovis, a possible ArsR-family transcriptional regulator. Northeast Structural Genomics Consortium Target MbR242E. | | Descriptor: | POSSIBLE TRANSCRIPTIONAL REGULATORY PROTEIN (POSSIBLY ARSR-FAMILY) | | Authors: | Ramelot, T.A, Cort, J.R, Wang, D, Ciccosanti, C, Jiang, M, Nair, R, Rost, B, Swapna, G, Acton, T.B, Xiao, R, Everett, J.K, Montelione, G.T, Kennedy, M.A, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2009-06-26 | | Release date: | 2009-08-11 | | Last modified: | 2024-05-08 | | Method: | SOLUTION NMR | | Cite: | Solution NMR structure of the homodimeric winged helix-turn-helix DNA-binding domain (fragment 1-100) Mb0332 from Mycobacterium bovis, a possible ArsR-family transcriptional regulator. Northeast Structural Genomics Consortium Target MbR242E.
To be Published
|
|
2KLZ
 | |
2K4M
 
 | | Solution NMR Structure of M. thermoautotrophicum protein MTH_1000, Northeast Structural Genomics Consortium Target TR8 | | Descriptor: | UPF0146 protein MTH_1000 | | Authors: | Eletsky, A, Sukumaran, D, Semesi, A, Guido, V, Yee, A, Arrowsmith, C, Szyperski, T, Northeast Structural Genomics Consortium (NESG) | | Deposit date: | 2008-06-13 | | Release date: | 2008-08-12 | | Last modified: | 2024-05-29 | | Method: | SOLUTION NMR | | Cite: | Solution NMR Structure of M. thermoautotrophicum protein MTH_1000, Northeast Structural Genomics Consortium Target TR8
To be Published
|
|
