2J9N
 
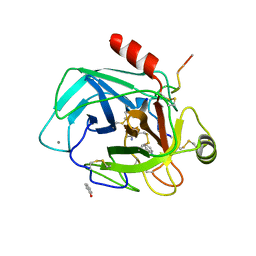 | | Robotically harvested Trypsin complexed with Benzamidine containing polypeptide mediated crystal contacts | | Descriptor: | BENZAMIDINE, CALCIUM ION, CATIONIC TRYPSIN, ... | | Authors: | Viola, R, Carman, P, Walsh, J, Miller, E, Benning, M, Frankel, D, McPherson, A, Cudney, R, Rupp, B. | | Deposit date: | 2006-11-14 | | Release date: | 2007-04-03 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Operator Assisted Harvesting of Protein Crystals Using a Universal Micromanipulation Robot.
J.Appl.Crystallogr., 40, 2007
|
|
4H9K
 
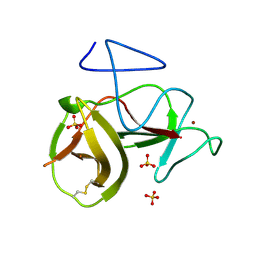 | | Crystal structure of cleavage site mutant of Npro of classical swine fever virus. | | Descriptor: | Hog cholera virus, SULFATE ION, ZINC ION | | Authors: | Gottipati, K, Ruggli, N, Gerber, M, Tratschin, J.-D, Benning, M, Bellamy, H, Choi, K.H. | | Deposit date: | 2012-09-24 | | Release date: | 2013-10-30 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.599 Å) | | Cite: | The Structure of Classical Swine Fever Virus N(pro): A Novel Cysteine Autoprotease and Zinc-Binding Protein Involved in Subversion of Type I Interferon Induction.
Plos Pathog., 9, 2013
|
|
4H9J
 
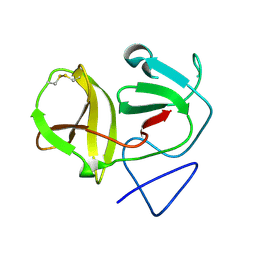 | | Crystal structure of N-terminal protease (Npro) of classical swine fever virus. | | Descriptor: | Hog cholera virus | | Authors: | Gottipati, K, Ruggli, N, Gerber, M, Tratschin, J.-D, Benning, M, Bellamy, H, Choi, K.H. | | Deposit date: | 2012-09-24 | | Release date: | 2013-10-30 | | Last modified: | 2018-01-24 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | The Structure of Classical Swine Fever Virus N(pro): A Novel Cysteine Autoprotease and Zinc-Binding Protein Involved in Subversion of Type I Interferon Induction.
Plos Pathog., 9, 2013
|
|
1PTA
 
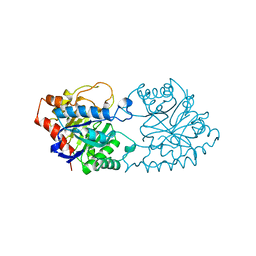 | |
1MDC
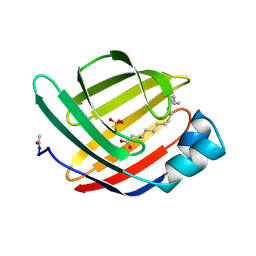 
 | |
1RK3
 
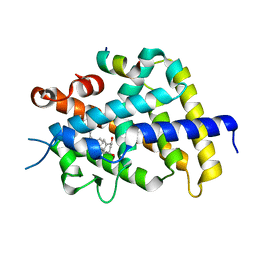 | | crystal structure of the rat vitamin D receptor ligand binding domain complexed with 1,25-dihydroxyvitamin D3 and a synthetic peptide containing the NR2 box of DRIP 205 | | Descriptor: | 5-{2-[1-(5-HYDROXY-1,5-DIMETHYL-HEXYL)-7A-METHYL-OCTAHYDRO-INDEN-4-YLIDENE]-ETHYLIDENE}-4-METHYLENE-CYCLOHEXANE-1,3-DIOL, Peroxisome proliferator-activated receptor binding protein, Vitamin D3 receptor | | Authors: | Vanhooke, J.L, M Benning, M, Bauer, C.B, Pike, J.W, DeLuca, H.F. | | Deposit date: | 2003-11-20 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Molecular Structure of the Rat Vitamin D Receptor Ligand Binding Domain Complexed with 2-Carbon-Substituted Vitamin D(3) Hormone Analogues and a LXXLL-Containing Coactivator Peptide
Biochemistry, 43, 2004
|
|
6B8R
 
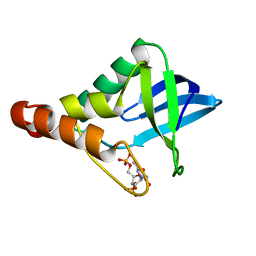 | | Crystal structure of Staphylococcal nuclease variant Delta+PHS V23K/L36Q at cryogenic temperature | | Descriptor: | CALCIUM ION, THYMIDINE-3',5'-DIPHOSPHATE, Thermonuclease | | Authors: | Robinson, A.C, Schlessman, J.L, Garcia-Moreno E, B, Benning, M. | | Deposit date: | 2017-10-09 | | Release date: | 2017-10-18 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Dielectric Properties of a Protein Probed by Reversal of a Buried Ion Pair.
J Phys Chem B, 122, 2018
|
|
3DMU
 
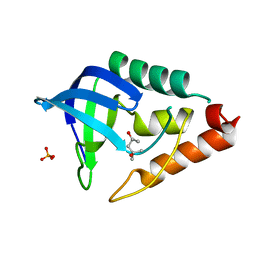 | | Crystal structure of Staphylococcal nuclease variant PHS T62K at cryogenic temperature | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PHOSPHATE ION, Thermonuclease | | Authors: | Khangulov, V.S, Schlessman, J.L, Garcia-Moreno, E.B, Benning, M, Isom, D. | | Deposit date: | 2008-07-01 | | Release date: | 2008-11-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of Staphylococcal nuclease variant PHS T62K at cryogenic temperature
To be Published
|
|
3EJI
 
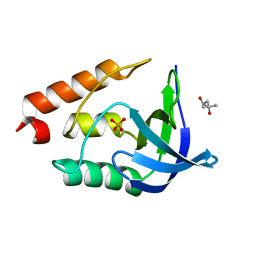 | | Crystal structure of Staphylococcal nuclease variant Delta+PHS L36K at cryogenic temperature | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, PHOSPHATE ION, Thermonuclease | | Authors: | Khangulov, V.S, Schlessman, J.L, Garcia-Moreno, E.B, Benning, M, Isom, D. | | Deposit date: | 2008-09-18 | | Release date: | 2008-11-04 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structural Consequences of the Ionization of Lysines at 25 Internal Positions in a Protein
To be Published
|
|
4HMI
 
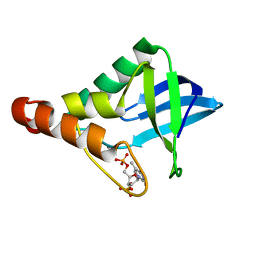 | | Crystal structure of Staphylococcal nuclease variant Delta+PHS V99K at cryogenic temperature | | Descriptor: | CALCIUM ION, THYMIDINE-3',5'-DIPHOSPHATE, Thermonuclease | | Authors: | Robinson, A.C, Schlessman, J.L, Garcia-Moreno E, B, Benning, M. | | Deposit date: | 2012-10-18 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of Staphylococcal nuclease variant Delta+PHS V99K at cryogenic temperature
To be Published
|
|
