- for default pairwise comparison.
$pkcombu -A [molecule_fileA] -B [molecule_fileB] -oam [atom-matching file]
- for Topologically constrained D-MCS (TD-MCS) with max_diff_of_dis = 1
$pkcombu -A [molecule_fileA] -B [molecule_fileB] -con T -mtd 1
- for exact C-MCS
$pkcombu -A [molecule_fileA] -B [molecule_fileB] -con C -alg X
- for showing options in detail
$pkcombu -h
- for showing options for mcs
$pkcombu -hmcs
< input options for 'pkcombu'> -A : molecule A (molA)(*.sdf|*.mol2|*.pdb|*.kcf|*.smi)[] -B : molecule B (molB)(*.sdf|*.mol2|*.pdb|*.kcf|*.smi)[] -fA,-fB : file formats.'P'db,'S'df,'K'cf, '2':MOL2, 'M':smiles [--] -aA,-aB : AtomHetero type. 'A'tom 'H'etatm 'B'oth[BB]The program "pkcombu" can read several file formats of chemical structures: sdf, mol2, pdb, kcf and smi. The file format is distinguished by file exetentions : *.sdf -> SDF, *.mol2 ->MOL2, *.pdb -> PDB, *.kcf -> KCF, *.smi -> SMILES. For molecular files without proper file extensions, users should directly assign the format types by the option '-fA' or '-fB'. For example, if a file "hoge" is for molecule A and in SDF format, users should add the option "-A hoge -fA S". For the molecule in PDB format, a bond connection table is automatically generated from the 3D coordinates of atoms, even if it does not have 'CONNCT' lines.
The options "-oam" and "-oAm" are for outputting detailed descriptions of calcualted atom matching (atom correspondence). The option "-oam" is for outputting the best match, the option "-oAm" is for outputting all the calculated candidate matches. The file format of atom matching file is described in other section.
The option '-sup3' is for superimposing molecules. The pkcombu program can simply superimpose molecule A with smallest RMSD against matched atoms of molecule B. For example, if the user assign "-sup3 T -opA outA.pdb", the structure of superimposed molecule A (for molecule B) is written as "outA.pdb" in PDB format.
The option "-oras" is for showing matching atoms using the molecular visualization program RasMol. If the user assign "-oras out", the pkcombu program generates two files "out-A.ras" and "out-B.ras". To visualize matched atoms in molecule A, a user inputs commands as follows:
$ rasmol -pdb [moleculeA(in PDB)] or $ rasmol -mdl [moleculeA(in SDF)] or $ rasmol -mol2 [moleculeA(in MOL2)] RasMol> script "out-A.ras"
The options "-ops" is for output PostScript file to show corresponding atom pairs.
This option basically assumes that input molecules has two-dimentional structures.
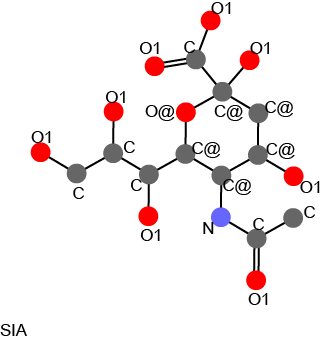
This figure is generated by following commands:
$pkcombu -A SIA.sdf -B G39.sdf -ops out.ps $evince out.ps