[English] 日本語
 Yorodumi
Yorodumi- PDB-7acs: The SARS-CoV-2 nucleocapsid phosphoprotein N-terminal domain in c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7acs | ||||||
|---|---|---|---|---|---|---|---|
| Title | The SARS-CoV-2 nucleocapsid phosphoprotein N-terminal domain in complex with 7mer dsRNA | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  VIRAL PROTEIN / protein-RNA complex / docking VIRAL PROTEIN / protein-RNA complex / docking | ||||||
| Function / homology |  Function and homology information Function and homology informationcytoplasmic capsid assembly / viral RNA genome packaging / response to host immune response / negative regulation of interferon-beta production / RNA stem-loop binding / Maturation of nucleoprotein / intracellular non-membrane-bounded organelle / positive regulation of NLRP3 inflammasome complex assembly / CD28 dependent PI3K/Akt signaling / MHC class I protein binding ...cytoplasmic capsid assembly / viral RNA genome packaging / response to host immune response / negative regulation of interferon-beta production / RNA stem-loop binding / Maturation of nucleoprotein / intracellular non-membrane-bounded organelle / positive regulation of NLRP3 inflammasome complex assembly / CD28 dependent PI3K/Akt signaling / MHC class I protein binding / SARS-CoV-2 targets host intracellular signalling and regulatory pathways / molecular condensate scaffold activity / protein sequestering activity / VEGFR2 mediated vascular permeability / TAK1-dependent IKK and NF-kappa-B activation / DDX58/IFIH1-mediated induction of interferon-alpha/beta / NOD1/2 Signaling Pathway / MHC class I protein complex / Interleukin-1 signaling /  viral capsid / Interferon alpha/beta signaling / PIP3 activates AKT signaling / Transcription of SARS-CoV-2 sgRNAs / host cell endoplasmic reticulum-Golgi intermediate compartment / host cell Golgi apparatus / viral nucleocapsid / Translation of Structural Proteins / Virion Assembly and Release / host extracellular space / Induction of Cell-Cell Fusion / Attachment and Entry / host cell perinuclear region of cytoplasm / viral capsid / Interferon alpha/beta signaling / PIP3 activates AKT signaling / Transcription of SARS-CoV-2 sgRNAs / host cell endoplasmic reticulum-Golgi intermediate compartment / host cell Golgi apparatus / viral nucleocapsid / Translation of Structural Proteins / Virion Assembly and Release / host extracellular space / Induction of Cell-Cell Fusion / Attachment and Entry / host cell perinuclear region of cytoplasm /  ribonucleoprotein complex / SARS-CoV-2 activates/modulates innate and adaptive immune responses / protein homodimerization activity / ribonucleoprotein complex / SARS-CoV-2 activates/modulates innate and adaptive immune responses / protein homodimerization activity /  RNA binding / extracellular region / identical protein binding / RNA binding / extracellular region / identical protein binding /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2 | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  molecular dynamics molecular dynamics | ||||||
 Authors Authors | Veverka, V. | ||||||
 Citation Citation |  Journal: Plos Pathog. / Year: 2020 Journal: Plos Pathog. / Year: 2020Title: Structural basis of RNA recognition by the SARS-CoV-2 nucleocapsid phosphoprotein. Authors: Dinesh, D.C. / Chalupska, D. / Silhan, J. / Koutna, E. / Nencka, R. / Veverka, V. / Boura, E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7acs.cif.gz 7acs.cif.gz | 1.6 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7acs.ent.gz pdb7acs.ent.gz | 1.4 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7acs.json.gz 7acs.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ac/7acs https://data.pdbj.org/pub/pdb/validation_reports/ac/7acs ftp://data.pdbj.org/pub/pdb/validation_reports/ac/7acs ftp://data.pdbj.org/pub/pdb/validation_reports/ac/7acs | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 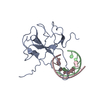
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein |  / Nucleocapsid protein / Protein N / Nucleocapsid protein / Protein NMass: 15129.853 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2Production host:   Escherichia coli (E. coli) / References: UniProt: P0DTC9 Escherichia coli (E. coli) / References: UniProt: P0DTC9 |
|---|---|
| #2: RNA chain | Mass: 2180.371 Da / Num. of mol.: 1 / Source method: obtained synthetically Source: (synth.)   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2 |
| #3: RNA chain | Mass: 2237.379 Da / Num. of mol.: 1 / Source method: obtained synthetically Source: (synth.)   Severe acute respiratory syndrome coronavirus 2 Severe acute respiratory syndrome coronavirus 2 |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR |
|---|---|
| NMR experiment | Sample state: isotropic / Type : 2D 1H-15N : 2D 1H-15N  HSQC HSQC |
- Sample preparation
Sample preparation
| Details | Type: solution Contents: 100 uM [U-13C; U-15N] N-NTD, 100 uM RNA (5'-R(P*CP*AP*CP*UP*GP*AP*C)-3'), 100 uM RNA (5'-R(P*GP*UP*CP*AP*GP*UP*G)-3'), 20 mM sodium phosphate, 50 mM sodium chloride, 95% H2O/5% D2O Label: s1 / Solvent system: 95% H2O/5% D2O | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||
| Sample conditions | Ionic strength: 70 mM / Label: c1 / pH: 5.5 / Pressure: 1 atm / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker AVANCE III HD / Manufacturer: Bruker / Model : AVANCE III HD / Field strength: 850 MHz : AVANCE III HD / Field strength: 850 MHz |
|---|
- Processing
Processing
| NMR software | Name: HADDOCK / Developer: Bonvin / Classification: structure calculation |
|---|---|
| Refinement | Method:  molecular dynamics / Software ordinal: 1 molecular dynamics / Software ordinal: 1 |
| NMR representative | Selection criteria: lowest energy |
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 200 / Conformers submitted total number: 30 |
 Movie
Movie Controller
Controller














 PDBj
PDBj



