[English] 日本語
 Yorodumi
Yorodumi- PDB-6b1c: Macrophage Migration Inhibitory Factor in complex with a Naphthyr... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6b1c | ||||||
|---|---|---|---|---|---|---|---|
| Title | Macrophage Migration Inhibitory Factor in complex with a Naphthyridinone Inhibitor (4a) | ||||||
 Components Components | Macrophage migration inhibitory factor | ||||||
 Keywords Keywords | ISOMERASE/ISOMERASE inhibitor /  CD74 BINDING / TAUTOMERASE / CD74 BINDING / TAUTOMERASE /  INHIBITOR / INHIBITOR /  COMPLEX / ISOMERASE-ISOMERASE INHIBITOR COMPLEX / COMPLEX / ISOMERASE-ISOMERASE INHIBITOR COMPLEX /  ISOMERASE ISOMERASE | ||||||
| Function / homology |  Function and homology information Function and homology informationpositive regulation of prostaglandin secretion involved in immune response / positive regulation of myeloid leukocyte cytokine production involved in immune response / dopachrome isomerase activity /  phenylpyruvate tautomerase / phenylpyruvate tautomerase /  L-dopachrome isomerase / L-dopachrome isomerase /  phenylpyruvate tautomerase activity / phenylpyruvate tautomerase activity /  cytokine receptor binding / positive regulation of lipopolysaccharide-mediated signaling pathway / positive regulation of arachidonic acid secretion / negative regulation of myeloid cell apoptotic process ...positive regulation of prostaglandin secretion involved in immune response / positive regulation of myeloid leukocyte cytokine production involved in immune response / dopachrome isomerase activity / cytokine receptor binding / positive regulation of lipopolysaccharide-mediated signaling pathway / positive regulation of arachidonic acid secretion / negative regulation of myeloid cell apoptotic process ...positive regulation of prostaglandin secretion involved in immune response / positive regulation of myeloid leukocyte cytokine production involved in immune response / dopachrome isomerase activity /  phenylpyruvate tautomerase / phenylpyruvate tautomerase /  L-dopachrome isomerase / L-dopachrome isomerase /  phenylpyruvate tautomerase activity / phenylpyruvate tautomerase activity /  cytokine receptor binding / positive regulation of lipopolysaccharide-mediated signaling pathway / positive regulation of arachidonic acid secretion / negative regulation of myeloid cell apoptotic process / negative regulation of macrophage chemotaxis / negative regulation of mature B cell apoptotic process / positive regulation of chemokine (C-X-C motif) ligand 2 production / negative regulation of protein metabolic process / prostaglandin biosynthetic process / carboxylic acid metabolic process / cytokine receptor binding / positive regulation of lipopolysaccharide-mediated signaling pathway / positive regulation of arachidonic acid secretion / negative regulation of myeloid cell apoptotic process / negative regulation of macrophage chemotaxis / negative regulation of mature B cell apoptotic process / positive regulation of chemokine (C-X-C motif) ligand 2 production / negative regulation of protein metabolic process / prostaglandin biosynthetic process / carboxylic acid metabolic process /  regulation of macrophage activation / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator / regulation of macrophage activation / negative regulation of intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator /  chemoattractant activity / positive regulation of protein kinase A signaling / protein homotrimerization / negative regulation of cellular senescence / negative regulation of DNA damage response, signal transduction by p53 class mediator / DNA damage response, signal transduction by p53 class mediator / positive regulation of B cell proliferation / positive regulation of phosphorylation / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / negative regulation of cell migration / chemoattractant activity / positive regulation of protein kinase A signaling / protein homotrimerization / negative regulation of cellular senescence / negative regulation of DNA damage response, signal transduction by p53 class mediator / DNA damage response, signal transduction by p53 class mediator / positive regulation of B cell proliferation / positive regulation of phosphorylation / Gene and protein expression by JAK-STAT signaling after Interleukin-12 stimulation / negative regulation of cell migration /  cytokine activity / positive regulation of cytokine production / Cell surface interactions at the vascular wall / positive regulation of fibroblast proliferation / positive regulation of peptidyl-tyrosine phosphorylation / cytokine activity / positive regulation of cytokine production / Cell surface interactions at the vascular wall / positive regulation of fibroblast proliferation / positive regulation of peptidyl-tyrosine phosphorylation /  cellular senescence / positive regulation of tumor necrosis factor production / positive regulation of peptidyl-serine phosphorylation / secretory granule lumen / cellular senescence / positive regulation of tumor necrosis factor production / positive regulation of peptidyl-serine phosphorylation / secretory granule lumen /  protease binding / vesicle / ficolin-1-rich granule lumen / positive regulation of ERK1 and ERK2 cascade / cell surface receptor signaling pathway / protease binding / vesicle / ficolin-1-rich granule lumen / positive regulation of ERK1 and ERK2 cascade / cell surface receptor signaling pathway /  inflammatory response / negative regulation of gene expression / inflammatory response / negative regulation of gene expression /  innate immune response / positive regulation of cell population proliferation / Neutrophil degranulation / negative regulation of apoptotic process / innate immune response / positive regulation of cell population proliferation / Neutrophil degranulation / negative regulation of apoptotic process /  cell surface / cell surface /  extracellular space / extracellular exosome / extracellular region / extracellular space / extracellular exosome / extracellular region /  nucleoplasm / identical protein binding / nucleoplasm / identical protein binding /  plasma membrane / plasma membrane /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.163 Å MOLECULAR REPLACEMENT / Resolution: 2.163 Å | ||||||
 Authors Authors | Krimmer, S.G. / Robertson, M.J. / Jorgensen, W.L. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: ACS Med Chem Lett / Year: 2017 Journal: ACS Med Chem Lett / Year: 2017Title: Adding a Hydrogen Bond May Not Help: Naphthyridinone vs Quinoline Inhibitors of Macrophage Migration Inhibitory Factor. Authors: Dawson, T.K. / Dziedzic, P. / Robertson, M.J. / Cisneros, J.A. / Krimmer, S.G. / Newton, A.S. / Tirado-Rives, J. / Jorgensen, W.L. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6b1c.cif.gz 6b1c.cif.gz | 81.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6b1c.ent.gz pdb6b1c.ent.gz | 60.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6b1c.json.gz 6b1c.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b1/6b1c https://data.pdbj.org/pub/pdb/validation_reports/b1/6b1c ftp://data.pdbj.org/pub/pdb/validation_reports/b1/6b1c ftp://data.pdbj.org/pub/pdb/validation_reports/b1/6b1c | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6b1kC  6b2cC 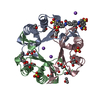 3u18S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  / MIF / Glycosylation-inhibiting factor / GIF / L-dopachrome isomerase / L-dopachrome tautomerase / ...MIF / Glycosylation-inhibiting factor / GIF / L-dopachrome isomerase / L-dopachrome tautomerase / Phenylpyruvate tautomerase / MIF / Glycosylation-inhibiting factor / GIF / L-dopachrome isomerase / L-dopachrome tautomerase / ...MIF / Glycosylation-inhibiting factor / GIF / L-dopachrome isomerase / L-dopachrome tautomerase / Phenylpyruvate tautomeraseMass: 12355.056 Da / Num. of mol.: 3 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: MIF, GLIF, MMIF / Plasmid: PET11B / Cell (production host): BL21 / Production host: Homo sapiens (human) / Gene: MIF, GLIF, MMIF / Plasmid: PET11B / Cell (production host): BL21 / Production host:   Escherichia coli (E. coli) Escherichia coli (E. coli)References: UniProt: P14174,  phenylpyruvate tautomerase, phenylpyruvate tautomerase,  L-dopachrome isomerase L-dopachrome isomerase#2: Chemical | ChemComp-SO4 /  Sulfate Sulfate#3: Chemical | #4: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.73 Å3/Da / Density % sol: 67.04 % |
|---|---|
Crystal grow | Temperature: 293.15 K / Method: vapor diffusion, sitting drop / pH: 7 Details: 2.6 M AMMONIUM SULFATE, 0.1 M TRIS-HCL PH 7, 3% ISOPROPANOL, 2% DMSO |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.54178 Å ROTATING ANODE / Type: RIGAKU MICROMAX-007 HF / Wavelength: 1.54178 Å |
| Detector | Type: RIGAKU SATURN 944+ / Detector: CCD / Date: Jul 27, 2017 |
| Radiation | Monochromator: DOUBLE MIRRORS / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.54178 Å / Relative weight: 1 : 1.54178 Å / Relative weight: 1 |
| Reflection | Resolution: 2.16→50 Å / Num. obs: 28763 / % possible obs: 95.8 % / Redundancy: 5.2 % / Rsym value: 0.077 / Net I/σ(I): 15.7 |
| Reflection shell | Resolution: 2.16→2.2 Å / Redundancy: 3 % / Mean I/σ(I) obs: 2.3 / Num. unique obs: 1067 / Rsym value: 0.458 / % possible all: 71.1 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 3U18 Resolution: 2.163→38.589 Å / SU ML: 0.3 / Cross valid method: FREE R-VALUE / σ(F): 1.33 / Phase error: 27.16
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.163→38.589 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller















 PDBj
PDBj