[English] 日本語
 Yorodumi
Yorodumi- PDB-5zyx: Solution NMR structure of K30 peptide in 10 mM dioctanoyl phospha... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5zyx | ||||||
|---|---|---|---|---|---|---|---|
| Title | Solution NMR structure of K30 peptide in 10 mM dioctanoyl phosphatidylglycerol (D8PG) | ||||||
 Components Components | ARG-TRP-LYS-ARG-HIS-ILE-SER-GLU-GLN-LEU-ARG-ARG-ARG-ASP-ARG-LEU-GLN-ARG-GLN-ALA | ||||||
 Keywords Keywords |  ANTIMICROBIAL PROTEIN / ANTIMICROBIAL PROTEIN /  STRUCTURE FROM CYANA 2.1 / K30 / D8PG STRUCTURE FROM CYANA 2.1 / K30 / D8PG | ||||||
| Function / homology |  Function and homology information Function and homology informationAtg12-Atg5-Atg16 complex /  C-terminal protein lipidation / vacuole-isolation membrane contact site / ubiquitin-like protein transferase activity / microautophagy / C-terminal protein lipidation / vacuole-isolation membrane contact site / ubiquitin-like protein transferase activity / microautophagy /  xenophagy / protein localization to phagophore assembly site / phagophore assembly site membrane / corpus callosum development / negative stranded viral RNA replication ...Atg12-Atg5-Atg16 complex / xenophagy / protein localization to phagophore assembly site / phagophore assembly site membrane / corpus callosum development / negative stranded viral RNA replication ...Atg12-Atg5-Atg16 complex /  C-terminal protein lipidation / vacuole-isolation membrane contact site / ubiquitin-like protein transferase activity / microautophagy / C-terminal protein lipidation / vacuole-isolation membrane contact site / ubiquitin-like protein transferase activity / microautophagy /  xenophagy / protein localization to phagophore assembly site / phagophore assembly site membrane / corpus callosum development / negative stranded viral RNA replication / endolysosome membrane / xenophagy / protein localization to phagophore assembly site / phagophore assembly site membrane / corpus callosum development / negative stranded viral RNA replication / endolysosome membrane /  Macroautophagy / Macroautophagy /  axoneme / autophagosome membrane / axoneme / autophagosome membrane /  autophagosome assembly / autophagosome assembly /  autophagosome / positive regulation of autophagy / protein-membrane adaptor activity / sperm midpiece / hippocampus development / autophagosome / positive regulation of autophagy / protein-membrane adaptor activity / sperm midpiece / hippocampus development /  macroautophagy / macroautophagy /  protein transport / protein transport /  GTPase binding / defense response to virus / identical protein binding / GTPase binding / defense response to virus / identical protein binding /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Bhunia, A. / Mohid, A. / Stella, L. / Calligari, P. | ||||||
| Funding support |  India, 1items India, 1items
| ||||||
 Citation Citation |  Journal: Bioconjug.Chem. / Year: 2019 Journal: Bioconjug.Chem. / Year: 2019Title: Design, Synthesis, Antibacterial Potential, and Structural Characterization of N-Acylated Derivatives of the Human Autophagy 16 Polypeptide. Authors: Varnava, K.G. / Mohid, S.A. / Calligari, P. / Stella, L. / Reynison, J. / Bhunia, A. / Sarojini, V. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5zyx.cif.gz 5zyx.cif.gz | 172.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5zyx.ent.gz pdb5zyx.ent.gz | 122.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5zyx.json.gz 5zyx.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zy/5zyx https://data.pdbj.org/pub/pdb/validation_reports/zy/5zyx ftp://data.pdbj.org/pub/pdb/validation_reports/zy/5zyx ftp://data.pdbj.org/pub/pdb/validation_reports/zy/5zyx | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
|---|---|
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 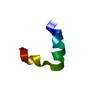
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 2698.125 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)   Homo sapiens (human) / References: UniProt: Q676U5*PLUS Homo sapiens (human) / References: UniProt: Q676U5*PLUS |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Type: solution / Contents: 1 mM 1H K30 Peptide, 90% H2O/10% D2O / Label: 1H / Solvent system: 90% H2O/10% D2O |
|---|---|
| Sample | Conc.: 1 mM / Component: K30 Peptide / Isotopic labeling: 1H |
| Sample conditions | Ionic strength: 0 Not defined / Label: Solution in water / pH: 4.5 / Pressure: 1 bar / Temperature: 298 K |
-NMR measurement
| NMR spectrometer | Type: Bruker AVANCE III / Manufacturer: Bruker / Model : AVANCE III / Field strength: 500 MHz : AVANCE III / Field strength: 500 MHz |
|---|
- Processing
Processing
| NMR software |
| |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 3 simulated annealing / Software ordinal: 3 | |||||||||||||||
| NMR representative | Selection criteria: lowest energy | |||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller


 PDBj
PDBj