+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4pl6 | ||||||
|---|---|---|---|---|---|---|---|
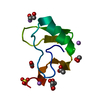
| Title | Structure of the chromodomain of MRG2 in complex with H3K4me3 | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  TRANSCRIPTION / TRANSCRIPTION /  chromo domain / chromo domain /  H3K4me3 H3K4me3 | ||||||
| Function / homology |  Function and homology information Function and homology information euchromatin binding / regulation of long-day photoperiodism, flowering / rDNA protrusion / regulation of timing of transition from vegetative to reproductive phase / epigenetic regulation of gene expression / methylated histone binding / promoter-specific chromatin binding / structural constituent of chromatin / : / euchromatin binding / regulation of long-day photoperiodism, flowering / rDNA protrusion / regulation of timing of transition from vegetative to reproductive phase / epigenetic regulation of gene expression / methylated histone binding / promoter-specific chromatin binding / structural constituent of chromatin / : /  nucleosome ... nucleosome ... euchromatin binding / regulation of long-day photoperiodism, flowering / rDNA protrusion / regulation of timing of transition from vegetative to reproductive phase / epigenetic regulation of gene expression / methylated histone binding / promoter-specific chromatin binding / structural constituent of chromatin / : / euchromatin binding / regulation of long-day photoperiodism, flowering / rDNA protrusion / regulation of timing of transition from vegetative to reproductive phase / epigenetic regulation of gene expression / methylated histone binding / promoter-specific chromatin binding / structural constituent of chromatin / : /  nucleosome / protein heterodimerization activity / nucleosome / protein heterodimerization activity /  nucleolus / regulation of DNA-templated transcription / nucleolus / regulation of DNA-templated transcription /  DNA binding / DNA binding /  nucleus / nucleus /  plasma membrane plasma membraneSimilarity search - Function | ||||||
| Biological species |   Arabidopsis thaliana (thale cress) Arabidopsis thaliana (thale cress) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.681 Å SYNCHROTRON / Resolution: 1.681 Å | ||||||
 Authors Authors | Liu, Y.C. / Huang, Y. | ||||||
 Citation Citation |  Journal: Plos Genet. / Year: 2014 Journal: Plos Genet. / Year: 2014Title: Regulation of arabidopsis flowering by the histone mark readers MRG1/2 via interaction with CONSTANS to modulate FT expression. Authors: Bu, Z. / Yu, Y. / Li, Z. / Liu, Y. / Jiang, W. / Huang, Y. / Dong, A.W. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4pl6.cif.gz 4pl6.cif.gz | 68 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4pl6.ent.gz pdb4pl6.ent.gz | 50.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4pl6.json.gz 4pl6.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/pl/4pl6 https://data.pdbj.org/pub/pdb/validation_reports/pl/4pl6 ftp://data.pdbj.org/pub/pdb/validation_reports/pl/4pl6 ftp://data.pdbj.org/pub/pdb/validation_reports/pl/4pl6 | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 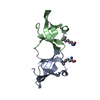
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9118.196 Da / Num. of mol.: 2 / Fragment: UNP residues 51-123 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Arabidopsis thaliana (thale cress) / Gene: At1g02740 / Production host: Arabidopsis thaliana (thale cress) / Gene: At1g02740 / Production host:   Escherichia coli (E. coli) / References: UniProt: Q4V3E2 Escherichia coli (E. coli) / References: UniProt: Q4V3E2#2: Protein/peptide |  Mass: 1293.516 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.)   Arabidopsis thaliana (thale cress) / References: UniProt: P59169*PLUS Arabidopsis thaliana (thale cress) / References: UniProt: P59169*PLUS#3: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.61 Å3/Da / Density % sol: 52.83 % |
|---|---|
Crystal grow | Temperature: 290 K / Method: vapor diffusion, hanging drop / Details: 0.1 M HEPES, PH 7.5, 9% PEG 6K, 5% MPD |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRF SSRF  / Beamline: BL17U / Wavelength: 0.9792 Å / Beamline: BL17U / Wavelength: 0.9792 Å |
| Detector | Type: RAYONIX MX225HE / Detector: CCD |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.9792 Å / Relative weight: 1 : 0.9792 Å / Relative weight: 1 |
| Reflection | Resolution: 1.68→30 Å / Num. obs: 23375 / % possible obs: 96.6 % / Redundancy: 3.3 % / Net I/σ(I): 36.1 |
- Processing
Processing
| Software | Name: PHENIX / Version: (phenix.refine: 1.8.4_1496) / Classification: refinement | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.681→24.965 Å / SU ML: 0.21 / Cross valid method: FREE R-VALUE / σ(F): 1.4 / Phase error: 22.31 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.8 Å / VDW probe radii: 1.1 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.681→24.965 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 3.3538 Å / Origin y: -7.2763 Å / Origin z: -18.5091 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: all |
 Movie
Movie Controller
Controller













 PDBj
PDBj


