+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2vb1 | ||||||
|---|---|---|---|---|---|---|---|
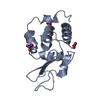
| Title | HEWL at 0.65 angstrom resolution | ||||||
 Components Components | LYSOZYME C | ||||||
 Keywords Keywords |  HYDROLASE / HYDROLASE /  ANTIMICROBIAL / TRICLINIC HEWL / ATOMIC RESOLUTION / ANTIMICROBIAL / TRICLINIC HEWL / ATOMIC RESOLUTION /  LYSOZYME / LYSOZYME /  ALLERGEN / GLYCOSIDASE / BACTERIOLYTIC ENZYME ALLERGEN / GLYCOSIDASE / BACTERIOLYTIC ENZYME | ||||||
| Function / homology |  Function and homology information Function and homology information Antimicrobial peptides / Neutrophil degranulation / Antimicrobial peptides / Neutrophil degranulation /  beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process / beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process /  lysozyme / lysozyme /  lysozyme activity / killing of cells of another organism / defense response to Gram-negative bacterium / defense response to Gram-positive bacterium / defense response to bacterium ... lysozyme activity / killing of cells of another organism / defense response to Gram-negative bacterium / defense response to Gram-positive bacterium / defense response to bacterium ... Antimicrobial peptides / Neutrophil degranulation / Antimicrobial peptides / Neutrophil degranulation /  beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process / beta-N-acetylglucosaminidase activity / cell wall macromolecule catabolic process /  lysozyme / lysozyme /  lysozyme activity / killing of cells of another organism / defense response to Gram-negative bacterium / defense response to Gram-positive bacterium / defense response to bacterium / lysozyme activity / killing of cells of another organism / defense response to Gram-negative bacterium / defense response to Gram-positive bacterium / defense response to bacterium /  endoplasmic reticulum / endoplasmic reticulum /  extracellular space / identical protein binding / extracellular space / identical protein binding /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   GALLUS GALLUS (chicken) GALLUS GALLUS (chicken) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / OTHER / Resolution: 0.65 Å SYNCHROTRON / OTHER / Resolution: 0.65 Å | ||||||
 Authors Authors | Wang, J. / Dauter, M. / Alkire, R. / Joachimiak, A. / Dauter, Z. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.D / Year: 2007 Journal: Acta Crystallogr.,Sect.D / Year: 2007Title: Triclinic Lysozyme at 0.65 A Resolution. Authors: Wang, J. / Dauter, M. / Alkire, R. / Joachimiak, A. / Dauter, Z. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2vb1.cif.gz 2vb1.cif.gz | 113.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2vb1.ent.gz pdb2vb1.ent.gz | 95.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2vb1.json.gz 2vb1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/vb/2vb1 https://data.pdbj.org/pub/pdb/validation_reports/vb/2vb1 ftp://data.pdbj.org/pub/pdb/validation_reports/vb/2vb1 ftp://data.pdbj.org/pub/pdb/validation_reports/vb/2vb1 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14331.160 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)   GALLUS GALLUS (chicken) / Cell: EGG / References: UniProt: P00698, GALLUS GALLUS (chicken) / Cell: EGG / References: UniProt: P00698,  lysozyme lysozyme | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-ACT /  Acetate Acetate | ||||
| #3: Chemical |  Ethylene glycol Ethylene glycol#4: Chemical | ChemComp-NO3 /  Nitrate Nitrate#5: Water | ChemComp-HOH / |  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.69 Å3/Da / Density % sol: 26.9 % / Description: NONE |
|---|---|
Crystal grow | Method: vapor diffusion, sitting drop / pH: 4.7 Details: SITTING DROP METHOD. PROTEIN SOLUTION: 35 MG/ML HEWL, 0.02 M NA ACETATE BUFFER PH 4.7 WELL SOLUTION: 1 M NANO3, 0.1 M NA ACETATE BUFFER PH 4.7, 20 % ETHYLENE GLYCOL PROTEIN AND WELL ...Details: SITTING DROP METHOD. PROTEIN SOLUTION: 35 MG/ML HEWL, 0.02 M NA ACETATE BUFFER PH 4.7 WELL SOLUTION: 1 M NANO3, 0.1 M NA ACETATE BUFFER PH 4.7, 20 % ETHYLENE GLYCOL PROTEIN AND WELL SOLUTIONS MIXED 1:1 IN A DROP AND SEEDED |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.65 / Beamline: 19-ID / Wavelength: 0.65 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Jul 5, 2006 / Details: MIRRORS |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.65 Å / Relative weight: 1 : 0.65 Å / Relative weight: 1 |
| Reflection | Resolution: 0.65→30 Å / Num. obs: 187165 / % possible obs: 97.6 % / Redundancy: 7.1 % / Rmerge(I) obs: 0.04 / Net I/σ(I): 36.2 |
| Reflection shell | Resolution: 0.65→0.67 Å / Redundancy: 2.7 % / Rmerge(I) obs: 0.18 / Mean I/σ(I) obs: 4.2 / % possible all: 67.3 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : OTHER : OTHERStarting model: NONE Resolution: 0.65→30 Å / Num. parameters: 14111 / Num. restraintsaints: 20151 / Cross valid method: FREE R-VALUE / σ(F): 0 / Stereochemistry target values: ENGH AND HUBER
| |||||||||||||||||||||||||||||||||
| Refine analyze | Num. disordered residues: 65 / Occupancy sum hydrogen: 976.69 / Occupancy sum non hydrogen: 1172.99 | |||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 0.65→30 Å
| |||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller















































































































































































































































 PDBj
PDBj











