+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2g16 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
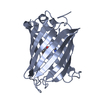
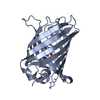
| Title | Structure of S65A Y66S GFP variant after backbone fragmentation | |||||||||
 Components Components | (Green fluorescent protein ) x 2 ) x 2 | |||||||||
 Keywords Keywords |  LUMINESCENT PROTEIN / LUMINESCENT PROTEIN /  Beta barrel / Beta barrel /  chromophore / chromophore /  biosynthesis / fragmentaion / denaturation biosynthesis / fragmentaion / denaturation | |||||||||
| Function / homology |  Function and homology information Function and homology information | |||||||||
| Biological species |   Aequorea victoria (jellyfish) Aequorea victoria (jellyfish) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | |||||||||
 Authors Authors | Barondeau, D.P. | |||||||||
 Citation Citation |  Journal: J.Am.Chem.Soc. / Year: 2006 Journal: J.Am.Chem.Soc. / Year: 2006Title: Understanding GFP Posttranslational Chemistry: Structures of Designed Variants that Achieve Backbone Fragmentation, Hydrolysis, and Decarboxylation. Authors: Barondeau, D.P. / Kassmann, C.J. / Tainer, J.A. / Getzoff, E.D. | |||||||||
| History |
| |||||||||
| Remark 999 | SEQUENCE Ser 65 in the database has been mutated to alanine, Tyr 66 in the database has been ...SEQUENCE Ser 65 in the database has been mutated to alanine, Tyr 66 in the database has been mutated to serine. Residues Ala 65, Ser 66 and Gly 67 consitute the chromophore CRW 66. This ligand underwent denaturation-induced backbone fragmentation between the C1 and CA1 atoms resulting in fragmented CRW ligand and an E1H molecule |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2g16.cif.gz 2g16.cif.gz | 63 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2g16.ent.gz pdb2g16.ent.gz | 44.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2g16.json.gz 2g16.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/g1/2g16 https://data.pdbj.org/pub/pdb/validation_reports/g1/2g16 ftp://data.pdbj.org/pub/pdb/validation_reports/g1/2g16 ftp://data.pdbj.org/pub/pdb/validation_reports/g1/2g16 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2g2sC  2g3dC  2g5zC  2g6eC  1emaS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein |  Mass: 6930.946 Da / Num. of mol.: 1 / Fragment: residues 2-64 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Aequorea victoria (jellyfish) / Gene: GFP / Plasmid: pET11a / Species (production host): Escherichia coli / Production host: Aequorea victoria (jellyfish) / Gene: GFP / Plasmid: pET11a / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21 (DE3) / References: UniProt: P42212 Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21 (DE3) / References: UniProt: P42212 |
|---|---|
| #2: Protein |  Mass: 19818.086 Da / Num. of mol.: 1 / Fragment: residues 65-238 / Mutation: S65A, Y66S, F99S, M153T, V163A Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Aequorea victoria (jellyfish) / Gene: GFP / Plasmid: pET11a / Species (production host): Escherichia coli / Production host: Aequorea victoria (jellyfish) / Gene: GFP / Plasmid: pET11a / Species (production host): Escherichia coli / Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21 (DE3) / References: UniProt: P42212 Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21 (DE3) / References: UniProt: P42212 |
| #3: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.1 Å3/Da / Density % sol: 41.3 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 8 Details: 50 mM MgCl2, 50 mM Hepes, 20% PEG 4000, pH 8.0, VAPOR DIFFUSION, HANGING DROP, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL11-1 / Wavelength: 0.85 Å / Beamline: BL11-1 / Wavelength: 0.85 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Mar 3, 2002 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.85 Å / Relative weight: 1 : 0.85 Å / Relative weight: 1 |
| Reflection | Resolution: 2→20 Å / Num. all: 15928 / Num. obs: 15582 / % possible obs: 97.8 % / Observed criterion σ(I): -3 / Biso Wilson estimate: 19 Å2 / Rsym value: 0.075 / Net I/σ(I): 16 |
| Reflection shell | Resolution: 2→2.07 Å / Mean I/σ(I) obs: 3.9 / Rsym value: 0.254 / % possible all: 97.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1ema Resolution: 2→20 Å / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→20 Å
| ||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||
| LS refinement shell | Resolution: 2→2.03 Å /
|
 Movie
Movie Controller
Controller






















 PDBj
PDBj


