[English] 日本語
 Yorodumi
Yorodumi- PDB-1zg5: NarL complexed to narG-89 promoter palindromic tail-to-tail DNA site -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1zg5 | ||||||
|---|---|---|---|---|---|---|---|
| Title | NarL complexed to narG-89 promoter palindromic tail-to-tail DNA site | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSCRIPTION/DNA /  protein-DNA complex / protein-DNA complex /  helix-turn-helix / two-component response regulator / dna bending / TRANSCRIPTION-DNA COMPLEX helix-turn-helix / two-component response regulator / dna bending / TRANSCRIPTION-DNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationDNA-binding transcription repressor activity / phosphorelay signal transduction system / nitrate assimilation /  protein-DNA complex / transcription cis-regulatory region binding / regulation of DNA-templated transcription / protein-DNA complex / transcription cis-regulatory region binding / regulation of DNA-templated transcription /  DNA binding / DNA binding /  ATP binding / identical protein binding / ATP binding / identical protein binding /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Escherichia coli (E. coli) Escherichia coli (E. coli) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.3 Å MOLECULAR REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Maris, A.E. / Kaczor-Grzeskowiak, M. / Ma, Z. / Kopka, M.L. / Gunsalus, R.P. / Dickerson, R.E. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 2005 Journal: Biochemistry / Year: 2005Title: Primary and Secondary Modes of DNA Recognition by the NarL Two-Component Response Regulator. Authors: Maris, A.E. / Kaczor-Grzeskowiak, M. / Ma, Z. / Kopka, M.L. / Gunsalus, R.P. / Dickerson, R.E. #1:  Journal: Nat.Struct.Biol. / Year: 2002 Journal: Nat.Struct.Biol. / Year: 2002Title: Dimerization allows DNA target site recognition by the NarL response regulator Authors: Maris, A.E. / Sawaya, M.R. / Kaczor-Grzeskowiak, M. / Jarvis, M.R. / Bearson, S.M.D. / Kopka, M.L. / Schroeder, I. / Gunsalus, R.P. / Dickerson, R.E. #2:  Journal: Biochemistry / Year: 1996 Journal: Biochemistry / Year: 1996Title: Structure of the Escherichia coli response regulator NarL Authors: Baikalov, I. / Schroeder, I. / Kaczor-Grzeskowiak, M. / Grzeskowiak, K. / Gunsalus, R.P. / Dickerson, R.E. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1zg5.cif.gz 1zg5.cif.gz | 120.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1zg5.ent.gz pdb1zg5.ent.gz | 89.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1zg5.json.gz 1zg5.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zg/1zg5 https://data.pdbj.org/pub/pdb/validation_reports/zg/1zg5 ftp://data.pdbj.org/pub/pdb/validation_reports/zg/1zg5 ftp://data.pdbj.org/pub/pdb/validation_reports/zg/1zg5 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | 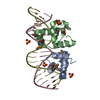 1zg1C  1je8S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The biological unit is a dimer bound to DNA. There are two biological units in the asymmetric unit (chains A, B, C & D and chains E, F, G & H) |
- Components
Components
| #1: DNA chain | Mass: 6134.967 Da / Num. of mol.: 4 / Source method: obtained synthetically / Details: solid phase synthesis #2: Protein | Mass: 9827.909 Da / Num. of mol.: 4 / Fragment: DNA binding domain (residues 147-216) Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Escherichia coli (E. coli) / Gene: NARL / Plasmid: pMJ05 / Production host: Escherichia coli (E. coli) / Gene: NARL / Plasmid: pMJ05 / Production host:   Escherichia coli (E. coli) / Strain (production host): JM109 / References: UniProt: P0AF28 Escherichia coli (E. coli) / Strain (production host): JM109 / References: UniProt: P0AF28#3: Chemical | ChemComp-SO4 /  Sulfate Sulfate#4: Water | ChemComp-HOH / |  Water Water |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.7 Å3/Da / Density % sol: 54.2 % |
|---|---|
Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: Ammonium sulfate, pH 7.5, vapor diffusion, hanging drop, temperature 298K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  NSLS NSLS  / Beamline: X8C / Wavelength: 1.1 Å / Beamline: X8C / Wavelength: 1.1 Å |
| Detector | Type: ADSC QUANTUM 4 / Detector: CCD / Date: Mar 11, 2001 |
| Radiation | Protocol: SINGLE WAVELENGTH / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 1.1 Å / Relative weight: 1 : 1.1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→50 Å / Num. all: 34483 / Num. obs: 34483 / % possible obs: 99.5 % / Observed criterion σ(I): -3 / Redundancy: 4.2 % / Biso Wilson estimate: 52.2 Å2 / Rmerge(I) obs: 0.104 / Rsym value: 0.131 / Χ2: 1.948 / Net I/σ(I): 16.4 |
| Reflection shell | Resolution: 2.2→2.25 Å / % possible obs: 96.4 % / Redundancy: 3.4 % / Rmerge(I) obs: 0.687 / Mean I/σ(I) obs: 2.9 / Num. measured obs: 2194 / Num. unique all: 2194 / Rsym value: 0.954 / Χ2: 4.303 / % possible all: 96.4 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 1JE8 Resolution: 2.3→20 Å / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: Engh & Huber
| |||||||||||||||||||||||||
| Solvent computation | Bsol: 38.791 Å2 | |||||||||||||||||||||||||
| Displacement parameters | Biso mean: 40.426 Å2
| |||||||||||||||||||||||||
| Refine analyze |
| |||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→20 Å
| |||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.3→2.44 Å / Rfactor Rfree error: 0.025
| |||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller














 PDBj
PDBj










































