+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1sdz | ||||||
|---|---|---|---|---|---|---|---|
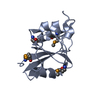
| Title | Crystal structure of DIAP1 BIR1 bound to a Reaper peptide | ||||||
 Components Components |
| ||||||
 Keywords Keywords |  APOPTOSIS / IAP / Reaper / APOPTOSIS / IAP / Reaper /  caspase / interaction caspase / interaction | ||||||
| Function / homology |  Function and homology information Function and homology informationSMAC, XIAP-regulated apoptotic response / DS ligand bound to FT receptor / negative regulation of compound eye retinal cell programmed cell death / antennal morphogenesis / Deactivation of the beta-catenin transactivating complex / Regulation of necroptotic cell death / Regulation of PTEN localization / sensory organ precursor cell division / Regulation of PTEN stability and activity / spermatid nucleus differentiation ...SMAC, XIAP-regulated apoptotic response / DS ligand bound to FT receptor / negative regulation of compound eye retinal cell programmed cell death / antennal morphogenesis / Deactivation of the beta-catenin transactivating complex / Regulation of necroptotic cell death / Regulation of PTEN localization / sensory organ precursor cell division / Regulation of PTEN stability and activity / spermatid nucleus differentiation / positive regulation of Toll signaling pathway / border follicle cell migration / positive regulation of border follicle cell migration / chaeta development /  caspase binding / negative regulation of JNK cascade / protein neddylation / ubiquitin conjugating enzyme binding / ubiquitin-like protein conjugating enzyme binding / NEDD8 ligase activity / cysteine-type endopeptidase inhibitor activity / ubiquitin-specific protease binding / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / protein K48-linked ubiquitination / protein autoubiquitination / positive regulation of protein ubiquitination / RING-type E3 ubiquitin transferase / caspase binding / negative regulation of JNK cascade / protein neddylation / ubiquitin conjugating enzyme binding / ubiquitin-like protein conjugating enzyme binding / NEDD8 ligase activity / cysteine-type endopeptidase inhibitor activity / ubiquitin-specific protease binding / cysteine-type endopeptidase inhibitor activity involved in apoptotic process / protein K48-linked ubiquitination / protein autoubiquitination / positive regulation of protein ubiquitination / RING-type E3 ubiquitin transferase /  Wnt signaling pathway / protein polyubiquitination / ubiquitin-protein transferase activity / positive regulation of canonical Wnt signaling pathway / Wnt signaling pathway / protein polyubiquitination / ubiquitin-protein transferase activity / positive regulation of canonical Wnt signaling pathway /  ubiquitin protein ligase activity / ubiquitin protein ligase activity /  spermatogenesis / spermatogenesis /  regulation of cell cycle / apoptotic process / regulation of cell cycle / apoptotic process /  ubiquitin protein ligase binding / negative regulation of apoptotic process / perinuclear region of cytoplasm / zinc ion binding / ubiquitin protein ligase binding / negative regulation of apoptotic process / perinuclear region of cytoplasm / zinc ion binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Drosophila melanogaster (fruit fly) Drosophila melanogaster (fruit fly) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.78 Å MOLECULAR REPLACEMENT / Resolution: 1.78 Å | ||||||
 Authors Authors | Yan, N. / Wu, J.W. / Shi, Y. | ||||||
 Citation Citation |  Journal: Nat.Struct.Mol.Biol. / Year: 2004 Journal: Nat.Struct.Mol.Biol. / Year: 2004Title: Molecular mechanisms of DrICE inhibition by DIAP1 and removal of inhibition by Reaper, Hid and Grim. Authors: Yan, N. / Wu, J.W. / Chai, J. / Li, W. / Shi, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1sdz.cif.gz 1sdz.cif.gz | 38.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1sdz.ent.gz pdb1sdz.ent.gz | 25.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1sdz.json.gz 1sdz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/sd/1sdz https://data.pdbj.org/pub/pdb/validation_reports/sd/1sdz ftp://data.pdbj.org/pub/pdb/validation_reports/sd/1sdz ftp://data.pdbj.org/pub/pdb/validation_reports/sd/1sdz | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 13552.203 Da / Num. of mol.: 1 / Fragment: DIAP1 BIR1 domain Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Drosophila melanogaster (fruit fly) / Gene: IAP1, TH / Production host: Drosophila melanogaster (fruit fly) / Gene: IAP1, TH / Production host:   Escherichia coli (E. coli) / References: UniProt: Q24306 Escherichia coli (E. coli) / References: UniProt: Q24306 |
|---|---|
| #2: Protein/peptide | Mass: 1094.216 Da / Num. of mol.: 1 / Fragment: Reaper N-terminal peptide / Source method: obtained synthetically Details: The sequence of this peptide occurs naturally in Drosophila (fruit flies). |
| #3: Chemical | ChemComp-ZN / |
| #4: Water | ChemComp-HOH /  Water Water |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.35 Å3/Da / Density % sol: 47.69 % |
|---|---|
Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 9.5 Details: CHES, sodium citrate, pH 9.5, VAPOR DIFFUSION, HANGING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  CHESS CHESS  / Beamline: A1 / Wavelength: 0.961 / Beamline: A1 / Wavelength: 0.961 |
| Detector | Type: FUJI / Detector: IMAGE PLATE / Date: Jul 1, 2003 |
| Radiation | Monochromator: Graphite / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength : 0.961 Å / Relative weight: 1 : 0.961 Å / Relative weight: 1 |
| Reflection | Resolution: 1.78→99 Å / Num. all: 12748 / Num. obs: 12620 / % possible obs: 99 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 9 % / Rmerge(I) obs: 0.056 / Rsym value: 0.059 |
| Reflection shell | Resolution: 1.78→1.85 Å / % possible all: 92.1 |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure : :  MOLECULAR REPLACEMENT / Resolution: 1.78→20 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber MOLECULAR REPLACEMENT / Resolution: 1.78→20 Å / σ(F): 0 / Stereochemistry target values: Engh & Huber
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.78→20 Å
| ||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller













 PDBj
PDBj





