+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qjk | ||||||
|---|---|---|---|---|---|---|---|
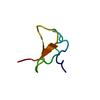
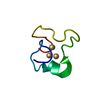
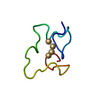
| Title | Metallothionein MTA from sea urchin (alpha domain) | ||||||
 Components Components | METALLOTHIONEIN | ||||||
 Keywords Keywords |  METALLOTHIONEIN / METAL-BINDING / METALLOTHIONEIN / METAL-BINDING /  DETOXIFICATION / DETOXIFICATION /  RADICAL SCAVENGER RADICAL SCAVENGER | ||||||
| Function / homology |  Metallothionein, family 4, echinoidea / Metallothionein, family 4, echinoidea /  Metallothionein, family 4, echinoidea/annelida / Metallothionein, family 4, echinoidea/annelida /  Metallothionein / Metallothionein domain superfamily / Metallothionein / Metallothionein domain superfamily /  metal ion binding / metal ion binding /  : / Metallothionein-A : / Metallothionein-A Function and homology information Function and homology information | ||||||
| Biological species |   STRONGYLOCENTROTUS PURPURATUS (purple sea urchin) STRONGYLOCENTROTUS PURPURATUS (purple sea urchin) | ||||||
| Method |  SOLUTION NMR / torsion angle dynamics SOLUTION NMR / torsion angle dynamics | ||||||
| Model type details | MINIMIZED AVERAGE | ||||||
 Authors Authors | Riek, R. / Precheur, B. / Wang, Y. / Mackay, E.A. / Wider, G. / Guntert, P. / Liu, A. / Kaegi, J.H.R. / Wuthrich, K. | ||||||
 Citation Citation |  Journal: J. Mol. Biol. / Year: 1999 Journal: J. Mol. Biol. / Year: 1999Title: NMR structure of the sea urchin (Strongylocentrotus purpuratus) metallothionein MTA. Authors: Riek, R. / Precheur, B. / Wang, Y. / Mackay, E.A. / Wider, G. / Guntert, P. / Liu, A. / Kagi, J.H. / Wuthrich, K. #1:  Journal: J.Mol.Biol. / Year: 1990 Journal: J.Mol.Biol. / Year: 1990Title: Three-Dimensional Structure of Human [113Cd7]-Metallothionein-2 in Solution Determined by Nuclear Magnetic Resonance Spectroscopy Authors: Messerle, B.A. / Schaffer, A. / Vasak, M. / Kaegi, J.H.R. / Wuthrich, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qjk.cif.gz 1qjk.cif.gz | 16.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qjk.ent.gz pdb1qjk.ent.gz | 11.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qjk.json.gz 1qjk.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qj/1qjk https://data.pdbj.org/pub/pdb/validation_reports/qj/1qjk ftp://data.pdbj.org/pub/pdb/validation_reports/qj/1qjk ftp://data.pdbj.org/pub/pdb/validation_reports/qj/1qjk | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide |  Mass: 3715.345 Da / Num. of mol.: 1 / Fragment: ALPHA DOMAIN Source method: isolated from a genetically manipulated source Details: CADMIUM 4-METAL CLUSTER Source: (gene. exp.)   STRONGYLOCENTROTUS PURPURATUS (purple sea urchin) STRONGYLOCENTROTUS PURPURATUS (purple sea urchin)Production host:   ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P04734 ESCHERICHIA COLI (E. coli) / Strain (production host): BL21(DE3) / References: UniProt: P04734 |
|---|---|
| #2: Chemical | ChemComp-CD / |
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Contents: 90% WATER/10% D2O |
|---|---|
| Sample conditions | Ionic strength: 50 MM NACL / pH: 7.0 / Pressure: 1 atm / Temperature: 288 K |
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: torsion angle dynamics / Software ordinal: 1 | ||||||||||||
| NMR ensemble | Conformer selection criteria: LEAST RESTRAINT VIOLATION / Conformers calculated total number: 20 / Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller










 PDBj
PDBj

