+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qg9 | ||||||
|---|---|---|---|---|---|---|---|
| Title | SECOND REPEAT (IS2MIC) FROM VOLTAGE-GATED SODIUM CHANNEL | ||||||
 Components Components | PROTEIN (SODIUM CHANNEL PROTEIN, BRAIN II ALPHA SUBUNIT) | ||||||
 Keywords Keywords | TRANSMEMBRANE CHANNEL / TRANSMEMBRANE SODIUM CHANNEL | ||||||
| Function / homology |  Function and homology information Function and homology informationPhase 0 - rapid depolarisation / response to pyrethroid / membrane depolarization during action potential /  sodium ion binding / voltage-gated monoatomic ion channel activity / sodium ion binding / voltage-gated monoatomic ion channel activity /  voltage-gated sodium channel complex / voltage-gated sodium channel complex /  voltage-gated sodium channel activity / regulation of monoatomic ion transmembrane transport / cellular response to antibiotic / neuronal action potential ...Phase 0 - rapid depolarisation / response to pyrethroid / membrane depolarization during action potential / voltage-gated sodium channel activity / regulation of monoatomic ion transmembrane transport / cellular response to antibiotic / neuronal action potential ...Phase 0 - rapid depolarisation / response to pyrethroid / membrane depolarization during action potential /  sodium ion binding / voltage-gated monoatomic ion channel activity / sodium ion binding / voltage-gated monoatomic ion channel activity /  voltage-gated sodium channel complex / voltage-gated sodium channel complex /  voltage-gated sodium channel activity / regulation of monoatomic ion transmembrane transport / cellular response to antibiotic / neuronal action potential / sodium ion transmembrane transport / monoatomic cation channel activity / sensory perception of pain / response to wounding / voltage-gated sodium channel activity / regulation of monoatomic ion transmembrane transport / cellular response to antibiotic / neuronal action potential / sodium ion transmembrane transport / monoatomic cation channel activity / sensory perception of pain / response to wounding /  nervous system development / nervous system development /  calmodulin binding / calmodulin binding /  axon / neuronal cell body / axon / neuronal cell body /  membrane / membrane /  plasma membrane / plasma membrane /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Doak, D.J. / Mulvey, D. / Kawaguchi, K. / Villalain, J. / Campbell, I.D. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: Structural studies of synthetic peptides dissected from the voltage-gated sodium channel. Authors: Doak, D.G. / Mulvey, D. / Kawaguchi, K. / Villalain, J. / Campbell, I.D. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qg9.cif.gz 1qg9.cif.gz | 185.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qg9.ent.gz pdb1qg9.ent.gz | 130.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qg9.json.gz 1qg9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qg/1qg9 https://data.pdbj.org/pub/pdb/validation_reports/qg/1qg9 ftp://data.pdbj.org/pub/pdb/validation_reports/qg/1qg9 ftp://data.pdbj.org/pub/pdb/validation_reports/qg/1qg9 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
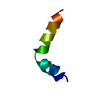
| Deposited unit | 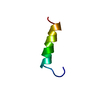
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
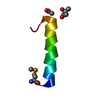
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide | Mass: 2480.853 Da / Num. of mol.: 1 / Fragment: S2 OF REPEAT I / Source method: obtained synthetically Details: SEQUENCE FROM MEMBRANES OF BRAIN CELLS OF RAT (RATTUS NORVEGICUS) References: UniProt: P08104 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: SEE JOURNAL REFERENCE |
- Sample preparation
Sample preparation
| Details | Contents: MICELLE MOLAR RATIO DPC/ IS2 = 68 |
|---|---|
| Sample conditions | pH: 3.3 / Temperature: 303 K |
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: STRUCTURES HAVE BEEN RECALCULATED USING PARAMETERS PARALLHDG5.1.PRO (MICHAEL NILGES) | ||||||||||||
| NMR ensemble | Conformer selection criteria: ENERGY / Conformers calculated total number: 50 / Conformers submitted total number: 30 |
 Movie
Movie Controller
Controller









 PDBj
PDBj

