[English] 日本語
 Yorodumi
Yorodumi- PDB-1q1v: Structure of the Oncoprotein DEK: a putative DNA-binding Domain R... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1q1v | ||||||
|---|---|---|---|---|---|---|---|
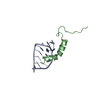
| Title | Structure of the Oncoprotein DEK: a putative DNA-binding Domain Related to the Winged Helix Motif | ||||||
 Components Components | DEK protein | ||||||
 Keywords Keywords |  DNA BINDING PROTEIN / WINGED-HELIX MOTIF DNA BINDING PROTEIN / WINGED-HELIX MOTIF | ||||||
| Function / homology |  Function and homology information Function and homology informationTranscriptional regulation by the AP-2 (TFAP2) family of transcription factors / regulation of double-strand break repair via nonhomologous end joining / contractile muscle fiber / B-WICH complex / regulation of double-strand break repair / : / positive regulation of transcription by RNA polymerase III / positive regulation of transcription by RNA polymerase I / viral genome replication / B-WICH complex positively regulates rRNA expression ...Transcriptional regulation by the AP-2 (TFAP2) family of transcription factors / regulation of double-strand break repair via nonhomologous end joining / contractile muscle fiber / B-WICH complex / regulation of double-strand break repair / : / positive regulation of transcription by RNA polymerase III / positive regulation of transcription by RNA polymerase I / viral genome replication / B-WICH complex positively regulates rRNA expression / Transcriptional regulation of granulopoiesis /  histone binding / transcription by RNA polymerase II / histone binding / transcription by RNA polymerase II /  chromatin remodeling / chromatin remodeling /  nucleolus / regulation of transcription by RNA polymerase II / nucleolus / regulation of transcription by RNA polymerase II /  signal transduction / positive regulation of transcription by RNA polymerase II / signal transduction / positive regulation of transcription by RNA polymerase II /  DNA binding / DNA binding /  RNA binding / RNA binding /  nucleoplasm / nucleoplasm /  nucleus nucleusSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / SOLUTION NMR /  simulated annealing simulated annealing | ||||||
 Authors Authors | Devany, M. / Kotharu, N.P. / Matsuo, H. | ||||||
 Citation Citation |  Journal: PROTEIN SCI. / Year: 2004 Journal: PROTEIN SCI. / Year: 2004Title: Solution NMR structure of the C-terminal domain of the human protein DEK Authors: Devany, M. / Kotharu, N.P. / Matsuo, H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1q1v.cif.gz 1q1v.cif.gz | 233.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1q1v.ent.gz pdb1q1v.ent.gz | 201.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1q1v.json.gz 1q1v.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/q1/1q1v https://data.pdbj.org/pub/pdb/validation_reports/q1/1q1v ftp://data.pdbj.org/pub/pdb/validation_reports/q1/1q1v ftp://data.pdbj.org/pub/pdb/validation_reports/q1/1q1v | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 8265.648 Da / Num. of mol.: 1 / Fragment: RESIDUES 309-378 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: dek / Plasmid: pET28c / Production host: Homo sapiens (human) / Gene: dek / Plasmid: pET28c / Production host:   Escherichia coli (E. coli) / Strain (production host): BL21(DE3)RIL / References: UniProt: P35659 Escherichia coli (E. coli) / Strain (production host): BL21(DE3)RIL / References: UniProt: P35659 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: DISTANCE RESTRAINTS COLLECTED FROM A NOESY SPECTRUM WITH 150MS MIXING TIME. |
- Sample preparation
Sample preparation
| Details | Contents: 1.0MM DEK C-TERMINAL DOMAIN |
|---|---|
| Sample conditions | Ionic strength: 50mM NAPO4, 100mM KCL / pH: 6.75 / Pressure: AMBIENT / Temperature: 293 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method:  simulated annealing / Software ordinal: 1 simulated annealing / Software ordinal: 1 Details: STRUCTURE BASED ON 1456 NON-REDUNDANT NOES, 33 HYDROGEN BOND RESTRAINTS DERIVED FROM DEUTERIUM EXCHANG EXPERIMENTS, AND 46 DIHEDRAL-ANGLE RESTRAINTS DERIVED FROM CA CHEMICAL SHIFTS. | |||||||||
| NMR ensemble | Conformer selection criteria: STRUCTURES WITH THE LOWEST TOTAL ENERGY Conformers calculated total number: 80 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller









 PDBj
PDBj
