[English] 日本語
 Yorodumi
Yorodumi- PDB-1icn: ESCHERICHIA COLI-DERIVED RAT INTESTINAL FATTY ACID BINDING PROTEI... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1icn | ||||||
|---|---|---|---|---|---|---|---|
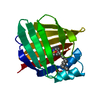
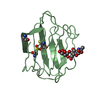
| Title | ESCHERICHIA COLI-DERIVED RAT INTESTINAL FATTY ACID BINDING PROTEIN WITH BOUND MYRISTATE AT 1.5 A RESOLUTION AND I-FABPARG106-->GLN WITH BOUND OLEATE AT 1.74 A RESOLUTION | ||||||
 Components Components | INTESTINAL FATTY ACID BINDING PROTEIN | ||||||
 Keywords Keywords | BINDING PROTEIN(FATTY ACID) | ||||||
| Function / homology |  Function and homology information Function and homology informationTriglyceride catabolism / intestinal lipid absorption / apical cortex / intestinal absorption / long-chain fatty acid transmembrane transporter activity /  long-chain fatty acid binding / long-chain fatty acid binding /  microvillus / long-chain fatty acid transport / fatty acid transport / fatty acid metabolic process ...Triglyceride catabolism / intestinal lipid absorption / apical cortex / intestinal absorption / long-chain fatty acid transmembrane transporter activity / microvillus / long-chain fatty acid transport / fatty acid transport / fatty acid metabolic process ...Triglyceride catabolism / intestinal lipid absorption / apical cortex / intestinal absorption / long-chain fatty acid transmembrane transporter activity /  long-chain fatty acid binding / long-chain fatty acid binding /  microvillus / long-chain fatty acid transport / fatty acid transport / fatty acid metabolic process / microvillus / long-chain fatty acid transport / fatty acid transport / fatty acid metabolic process /  fatty acid binding / fatty acid binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | ||||||
| Biological species |   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.74 Å X-RAY DIFFRACTION / Resolution: 1.74 Å | ||||||
 Authors Authors | Eads, J.C. / Sacchettini, J.C. / Kromminga, A. / Gordon, J.I. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 1993 Journal: J.Biol.Chem. / Year: 1993Title: Escherichia coli-derived rat intestinal fatty acid binding protein with bound myristate at 1.5 A resolution and I-FABPArg106-->Gln with bound oleate at 1.74 A resolution. Authors: Eads, J. / Sacchettini, J.C. / Kromminga, A. / Gordon, J.I. #1:  Journal: J.Biol.Chem. / Year: 1992 Journal: J.Biol.Chem. / Year: 1992Title: Refinement of the Structure of Recombinant Rat Intestinal Fatty Acid-Binding Apoprotein at 1.2 Angstroms Resolution Authors: Scapin, G. / Gordon, J.I. / Sacchettini, J.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1icn.cif.gz 1icn.cif.gz | 41.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1icn.ent.gz pdb1icn.ent.gz | 28.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1icn.json.gz 1icn.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ic/1icn https://data.pdbj.org/pub/pdb/validation_reports/ic/1icn ftp://data.pdbj.org/pub/pdb/validation_reports/ic/1icn ftp://data.pdbj.org/pub/pdb/validation_reports/ic/1icn | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14985.950 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Rattus norvegicus (Norway rat) / References: UniProt: P02693 Rattus norvegicus (Norway rat) / References: UniProt: P02693 |
|---|---|
| #2: Chemical | ChemComp-OLA /  Oleic acid Oleic acid |
| #3: Water | ChemComp-HOH /  Water Water |
| Nonpolymer details | BOUND OLEATE HAS THREE CONFORMATI |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.92 Å3/Da / Density % sol: 36.04 % | |||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Crystal grow | *PLUS Temperature: 17 ℃ / pH: 7.1 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | *PLUS Highest resolution: 1.7 Å / Lowest resolution: 10 Å / Num. obs: 10336 / Num. measured all: 21791 / Rmerge(I) obs: 0.0624 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor obs: 0.161 / Highest resolution: 1.74 Å | ||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 1.74 Å
| ||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 1.74 Å / Rfactor obs: 0.161 | ||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj








