+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1g5v | ||||||
|---|---|---|---|---|---|---|---|
| Title | SOLUTION STRUCTURE OF THE TUDOR DOMAIN OF THE HUMAN SMN PROTEIN | ||||||
 Components Components | SURVIVAL MOTOR NEURON PROTEIN 1 Survival of motor neuron Survival of motor neuron | ||||||
 Keywords Keywords |  TRANSLATION / TRANSLATION /  mRNA processing mRNA processing | ||||||
| Function / homology |  Function and homology information Function and homology informationGemini of coiled bodies / SMN complex / SMN-Sm protein complex / spliceosomal complex assembly /  Cajal body / spliceosomal snRNP assembly / DNA-templated transcription termination / cytoplasmic ribonucleoprotein granule / Z disc / Cajal body / spliceosomal snRNP assembly / DNA-templated transcription termination / cytoplasmic ribonucleoprotein granule / Z disc /  snRNP Assembly ...Gemini of coiled bodies / SMN complex / SMN-Sm protein complex / spliceosomal complex assembly / snRNP Assembly ...Gemini of coiled bodies / SMN complex / SMN-Sm protein complex / spliceosomal complex assembly /  Cajal body / spliceosomal snRNP assembly / DNA-templated transcription termination / cytoplasmic ribonucleoprotein granule / Z disc / Cajal body / spliceosomal snRNP assembly / DNA-templated transcription termination / cytoplasmic ribonucleoprotein granule / Z disc /  snRNP Assembly / snRNP Assembly /  nervous system development / SARS-CoV-2 modulates host translation machinery / nervous system development / SARS-CoV-2 modulates host translation machinery /  perikaryon / perikaryon /  nuclear body / neuron projection / nuclear body / neuron projection /  axon / axon /  RNA binding / RNA binding /  nucleoplasm / identical protein binding / nucleoplasm / identical protein binding /  nucleus / nucleus /  cytosol / cytosol /  cytoplasm cytoplasmSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / mixed torsion, Cartesian angle dynamics simulated annealing protocol SOLUTION NMR / mixed torsion, Cartesian angle dynamics simulated annealing protocol | ||||||
 Authors Authors | Selenko, P. / Sprangers, R. / Stier, G. / Buehler, D. / Fischer, U. / Sattler, M. | ||||||
 Citation Citation |  Journal: Nat.Struct.Biol. / Year: 2001 Journal: Nat.Struct.Biol. / Year: 2001Title: SMN tudor domain structure and its interaction with the Sm proteins. Authors: Selenko, P. / Sprangers, R. / Stier, G. / Buhler, D. / Fischer, U. / Sattler, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1g5v.cif.gz 1g5v.cif.gz | 195.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1g5v.ent.gz pdb1g5v.ent.gz | 169.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1g5v.json.gz 1g5v.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/g5/1g5v https://data.pdbj.org/pub/pdb/validation_reports/g5/1g5v ftp://data.pdbj.org/pub/pdb/validation_reports/g5/1g5v ftp://data.pdbj.org/pub/pdb/validation_reports/g5/1g5v | HTTPS FTP |
|---|
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 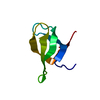
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein |  Survival of motor neuron Survival of motor neuronMass: 9796.644 Da / Num. of mol.: 1 / Fragment: TUDOR DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / Gene: SMN1 / Plasmid: PET9D / Species (production host): Escherichia coli Homo sapiens (human) / Gene: SMN1 / Plasmid: PET9D / Species (production host): Escherichia coliGene (production host): EXPRESSED AS FUSION PROTEIN WITH N-TERMINAL HIS6-GST AND TEV CLEAVAGE SITE Production host:   Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3) / References: UniProt: Q16637 Escherichia coli BL21(DE3) (bacteria) / Strain (production host): BL21(DE3) / References: UniProt: Q16637 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample conditions | Ionic strength: 20mM Phosphate 30mM NaCl / pH: 6.3 / Pressure: ambient / Temperature: 295 K |
-NMR measurement
| NMR spectrometer |
|
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: mixed torsion, Cartesian angle dynamics simulated annealing protocol Software ordinal: 1 Details: 1402 Unambiguous NOE distance restraints 50 Hydrogen bond restraints 44 HN-N dipolar couplings | ||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average,lowest energy | ||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: back calculated data agree with experimental NOESY spectrum,structures with acceptable covalent geometry,structures with favorable non-bond energy,structures with the ...Conformer selection criteria: back calculated data agree with experimental NOESY spectrum,structures with acceptable covalent geometry,structures with favorable non-bond energy,structures with the least restraint violations,structures with the lowest energy Conformers calculated total number: 100 / Conformers submitted total number: 10 |
 Movie
Movie Controller
Controller











 PDBj
PDBj
