[English] 日本語
 Yorodumi
Yorodumi- PDB-1bh7: A LOW ENERGY STRUCTURE FOR THE FINAL CYTOPLASMIC LOOP OF BAND 3, ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1bh7 | ||||||
|---|---|---|---|---|---|---|---|
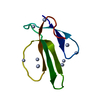
| Title | A LOW ENERGY STRUCTURE FOR THE FINAL CYTOPLASMIC LOOP OF BAND 3, NMR, MINIMIZED AVERAGE STRUCTURE | ||||||
 Components Components | BAND 3 Band 3 anion transport protein Band 3 anion transport protein | ||||||
 Keywords Keywords |  MEMBRANE PROTEIN / CYTOPLASMIC LOOP / MEMBRANE PROTEIN / CYTOPLASMIC LOOP /  ANION EXCHANGE PROTEIN ANION EXCHANGE PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationpH elevation / Defective SLC4A1 causes hereditary spherocytosis type 4 (HSP4), distal renal tubular acidosis (dRTA) and dRTA with hemolytic anemia (dRTA-HA) / negative regulation of urine volume / Bicarbonate transporters / intracellular monoatomic ion homeostasis /  ankyrin-1 complex / plasma membrane phospholipid scrambling / monoatomic anion transmembrane transporter activity / chloride:bicarbonate antiporter activity / solute:inorganic anion antiporter activity ...pH elevation / Defective SLC4A1 causes hereditary spherocytosis type 4 (HSP4), distal renal tubular acidosis (dRTA) and dRTA with hemolytic anemia (dRTA-HA) / negative regulation of urine volume / Bicarbonate transporters / intracellular monoatomic ion homeostasis / ankyrin-1 complex / plasma membrane phospholipid scrambling / monoatomic anion transmembrane transporter activity / chloride:bicarbonate antiporter activity / solute:inorganic anion antiporter activity ...pH elevation / Defective SLC4A1 causes hereditary spherocytosis type 4 (HSP4), distal renal tubular acidosis (dRTA) and dRTA with hemolytic anemia (dRTA-HA) / negative regulation of urine volume / Bicarbonate transporters / intracellular monoatomic ion homeostasis /  ankyrin-1 complex / plasma membrane phospholipid scrambling / monoatomic anion transmembrane transporter activity / chloride:bicarbonate antiporter activity / solute:inorganic anion antiporter activity / bicarbonate transmembrane transporter activity / bicarbonate transport / monoatomic anion transport / chloride transport / chloride transmembrane transporter activity / erythrocyte development / negative regulation of glycolytic process through fructose-6-phosphate / ankyrin-1 complex / plasma membrane phospholipid scrambling / monoatomic anion transmembrane transporter activity / chloride:bicarbonate antiporter activity / solute:inorganic anion antiporter activity / bicarbonate transmembrane transporter activity / bicarbonate transport / monoatomic anion transport / chloride transport / chloride transmembrane transporter activity / erythrocyte development / negative regulation of glycolytic process through fructose-6-phosphate /  ankyrin binding / ankyrin binding /  hemoglobin binding / cortical cytoskeleton / protein-membrane adaptor activity / chloride transmembrane transport / protein localization to plasma membrane / hemoglobin binding / cortical cytoskeleton / protein-membrane adaptor activity / chloride transmembrane transport / protein localization to plasma membrane /  regulation of intracellular pH / transmembrane transport / Erythrocytes take up oxygen and release carbon dioxide / Erythrocytes take up carbon dioxide and release oxygen / cytoplasmic side of plasma membrane / Z disc / regulation of intracellular pH / transmembrane transport / Erythrocytes take up oxygen and release carbon dioxide / Erythrocytes take up carbon dioxide and release oxygen / cytoplasmic side of plasma membrane / Z disc /  blood coagulation / basolateral plasma membrane / blood microparticle / protein homodimerization activity / extracellular exosome / blood coagulation / basolateral plasma membrane / blood microparticle / protein homodimerization activity / extracellular exosome /  membrane / membrane /  plasma membrane plasma membraneSimilarity search - Function | ||||||
| Biological species |   Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  SOLUTION NMR / DISTANCE GEOMETRY, SOLUTION NMR / DISTANCE GEOMETRY,  SIMULATED ANNEALING SIMULATED ANNEALING | ||||||
 Authors Authors | Askin, D. / Bloomberg, G.B. / Chambers, E.J. / Tanner, M.J.A. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1998 Journal: Biochemistry / Year: 1998Title: NMR solution structure of a cytoplasmic surface loop of the human red cell anion transporter, band 3. Authors: Askin, D. / Bloomberg, G.B. / Chambers, E.J. / Tanner, M.J. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1bh7.cif.gz 1bh7.cif.gz | 18.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1bh7.ent.gz pdb1bh7.ent.gz | 13.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1bh7.json.gz 1bh7.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/bh/1bh7 https://data.pdbj.org/pub/pdb/validation_reports/bh/1bh7 ftp://data.pdbj.org/pub/pdb/validation_reports/bh/1bh7 ftp://data.pdbj.org/pub/pdb/validation_reports/bh/1bh7 | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein/peptide |  Band 3 anion transport protein Band 3 anion transport proteinMass: 4158.074 Da / Num. of mol.: 1 / Fragment: FINAL CYTOPLASMIC LOOP Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Homo sapiens (human) / References: UniProt: P02730 Homo sapiens (human) / References: UniProt: P02730 |
|---|
-Experimental details
-Experiment
| Experiment | Method:  SOLUTION NMR SOLUTION NMR | ||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
| ||||||||||||||||
| NMR details | Text: THE STRUCTURE WAS DETERMINED USING 2D SOLUTION-STATE NMR |
- Sample preparation
Sample preparation
| Details | Contents: 30% TFE-D3/H2O |
|---|---|
| Sample conditions | Ionic strength: 12mM / pH: 3.5 / Pressure: ATMOSPHERIC atm / Temperature: 298 K |
Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: JEOL ALPHA 500MHZ / Manufacturer: JEOL / Model : ALPHA 500MHZ / Field strength: 500 MHz : ALPHA 500MHZ / Field strength: 500 MHz |
|---|
- Processing
Processing
| Software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR software |
| ||||||||||||
| Refinement | Method: DISTANCE GEOMETRY,  SIMULATED ANNEALING / Software ordinal: 1 SIMULATED ANNEALING / Software ordinal: 1 Details: 480 DISTANCE RESTRAINTS 10 DIHEDRAL ANGLE RESTRAINTS 4 H-BONDING DISTANCE RESTRAINTS NO NOE VIOLATIONS > 0.05NM NO TORSION ANGLE VIOLATIONS > 5 DEGREES | ||||||||||||
| NMR ensemble | Conformer selection criteria: AVERAGE STRUCTURE- ONLY THE THREE STRUCTURED REGIONS WITHIN THE PEPTIDE ARE GIVEN Conformers calculated total number: 15 / Conformers submitted total number: 1 |
 Movie
Movie Controller
Controller









 PDBj
PDBj