+ Open data
Open data
- Basic information
Basic information
| Entry | 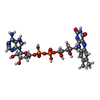 Database: PDB chemical components / ID: FAE Database: PDB chemical components / ID: FAE | ||
|---|---|---|---|
| Name | Name: | Wikipedia |  Wikipedia - Flavin adenine dinucleotide: In biochemistry, flavin adenine dinucleotide is a redox-active coenzyme associated with various proteins, which is involved with several enzymatic reactions in metabolism. A flavoprotein is a protein that contains a flavin group, which may be in the form of FAD or flavin mononucleotide (FMN)... Wikipedia - Flavin adenine dinucleotide: In biochemistry, flavin adenine dinucleotide is a redox-active coenzyme associated with various proteins, which is involved with several enzymatic reactions in metabolism. A flavoprotein is a protein that contains a flavin group, which may be in the form of FAD or flavin mononucleotide (FMN)... |
-Chemical information
| Composition |  | ||||
|---|---|---|---|---|---|
| Others | Type: NON-POLYMER / PDB classification: HETAIN / Three letter code: FAE / Ideal coordinates details: Corina / Model coordinates PDB-ID: 1MXT | ||||
| History |
| ||||
 External links External links |  UniChem / UniChem /  DrugBank / DrugBank /  ChemSpider / ChemSpider /  Wikipedia search / Wikipedia search /  Google search Google search |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
-Details
-SMILES
| ACDLabs 12.01 | | CACTVS 3.370 | OpenEye OEToolkits 1.7.0 | |
|---|
-SMILES CANONICAL
| CACTVS 3.370 | | OpenEye OEToolkits 1.7.0 | |
|---|
-InChI
| InChI 1.03 |
|---|
-InChIKey
| InChI 1.03 |
|---|
-SYSTEMATIC NAME
| ACDLabs 12.01 | | OpenEye OEToolkits 1.7.0 | [[( | |
|---|
-PDB entries
Showing all 4 items

PDB-1mxt: 
Atomic resolution structure of Cholesterol oxidase (Streptomyces sp. SA-COO)

PDB-1n4u: 
CHOLESTEROL OXIDASE FROM STREPTOMYCES @ pH 4.5 (STREPTOMYCES SP. SA-COO)

PDB-3b3r: 
Crystal structure of Streptomyces cholesterol oxidase H447Q/E361Q mutant bound to glycerol (0.98A)

PDB-3b6d: 
Crystal Structure of Streptomyces Cholesterol Oxidase H447Q/E361Q mutant (1.2A)
 Movie
Movie Controller
Controller



