+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-4277 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | CryoEM Structure INO80core Nucleosome complex | ||||||||||||
 マップデータ マップデータ | |||||||||||||
 試料 試料 |
| ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報RMTs methylate histone arginines / DASH complex / heterochromatin formation => GO:0031507 / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / Metalloprotease DUBs / UCH proteinases / Ub-specific processing proteases / attachment of spindle microtubules to kinetochore / chromatin => GO:0000785 ...RMTs methylate histone arginines / DASH complex / heterochromatin formation => GO:0031507 / protein transport along microtubule to mitotic spindle pole body / mitotic sister chromatid biorientation / Metalloprotease DUBs / UCH proteinases / Ub-specific processing proteases / attachment of spindle microtubules to kinetochore / chromatin => GO:0000785 / Ino80 complex / attachment of mitotic spindle microtubules to kinetochore / negative regulation of megakaryocyte differentiation / ATP-dependent activity, acting on DNA / protein localization to CENP-A containing chromatin / Chromatin modifying enzymes / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Deposition of new CENPA-containing nucleosomes at the centromere / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Inhibition of DNA recombination at telomere / Meiotic synapsis / telomere organization / RNA Polymerase I Promoter Opening / Interleukin-7 signaling / SUMOylation of chromatin organization proteins / Assembly of the ORC complex at the origin of replication /  DNAメチル化 / Condensation of Prophase Chromosomes / HCMV Late Events / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / SIRT1 negatively regulates rRNA expression / DNAメチル化 / Condensation of Prophase Chromosomes / HCMV Late Events / Chromatin modifications during the maternal to zygotic transition (MZT) / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / SIRT1 negatively regulates rRNA expression /  innate immune response in mucosa / PRC2 methylates histones and DNA / innate immune response in mucosa / PRC2 methylates histones and DNA /  helicase activity / Defective pyroptosis / HDACs deacetylate histones / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / G2/M DNA damage checkpoint / B-WICH complex positively regulates rRNA expression / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence / Metalloprotease DUBs / helicase activity / Defective pyroptosis / HDACs deacetylate histones / RNA Polymerase I Promoter Escape / Nonhomologous End-Joining (NHEJ) / Transcriptional regulation by small RNAs / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / NoRC negatively regulates rRNA expression / G2/M DNA damage checkpoint / B-WICH complex positively regulates rRNA expression / HDMs demethylate histones / DNA Damage/Telomere Stress Induced Senescence / Metalloprotease DUBs /  紡錘体 / 紡錘体 /  動原体 / PKMTs methylate histone lysines / RMTs methylate histone arginines / 動原体 / PKMTs methylate histone lysines / RMTs methylate histone arginines /  遺伝的組換え / Pre-NOTCH Transcription and Translation / 遺伝的組換え / Pre-NOTCH Transcription and Translation /  nucleosome assembly / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases / nucleosome assembly / Activation of anterior HOX genes in hindbrain development during early embryogenesis / HCMV Early Events / Transcriptional regulation of granulopoiesis / structural constituent of chromatin / UCH proteinases /  ヌクレオソーム / antimicrobial humoral immune response mediated by antimicrobial peptide / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates transcription of genes involved in differentiation of HSCs / chromatin organization / Factors involved in megakaryocyte development and platelet production / Processing of DNA double-strand break ends / HATs acetylate histones / antibacterial humoral response / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence / ヌクレオソーム / antimicrobial humoral immune response mediated by antimicrobial peptide / E3 ubiquitin ligases ubiquitinate target proteins / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / RUNX1 regulates transcription of genes involved in differentiation of HSCs / chromatin organization / Factors involved in megakaryocyte development and platelet production / Processing of DNA double-strand break ends / HATs acetylate histones / antibacterial humoral response / Senescence-Associated Secretory Phenotype (SASP) / Oxidative Stress Induced Senescence /  ヘリカーゼ / Estrogen-dependent gene expression / ヘリカーゼ / Estrogen-dependent gene expression /  chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium / chromosome, telomeric region / Ub-specific processing proteases / defense response to Gram-positive bacterium /  クロマチンリモデリング / Amyloid fiber formation / protein heterodimerization activity / クロマチンリモデリング / Amyloid fiber formation / protein heterodimerization activity /  DNA修復 / DNA修復 /  enzyme binding / enzyme binding /  ATP hydrolysis activity / protein-containing complex / ATP hydrolysis activity / protein-containing complex /  DNA binding / DNA binding /  extracellular space / extracellular space /  RNA binding / extracellular exosome / extracellular region / RNA binding / extracellular exosome / extracellular region /  核質 / 核質 /  ATP binding / ATP binding /  生体膜 / identical protein binding / 生体膜 / identical protein binding /  細胞核 細胞核類似検索 - 分子機能 | ||||||||||||
| 生物種 |  Chaetomium thermophilum var. thermophilum DSM 1495 (菌類) / Chaetomium thermophilum var. thermophilum DSM 1495 (菌類) /   Homo sapiens (ヒト) / synthetic construct (人工物) / Homo sapiens (ヒト) / synthetic construct (人工物) /  Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (菌類) Chaetomium thermophilum (strain DSM 1495 / CBS 144.50 / IMI 039719) (菌類) | ||||||||||||
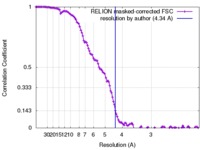
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 4.34 Å クライオ電子顕微鏡法 / 解像度: 4.34 Å | ||||||||||||
 データ登録者 データ登録者 | Eustermann S / Schall K / Kostrewa D / Strauss M / Hopfner K | ||||||||||||
| 資金援助 |  ドイツ, 3件 ドイツ, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2018 ジャーナル: Nature / 年: 2018タイトル: Structural basis for ATP-dependent chromatin remodelling by the INO80 complex. 著者: Sebastian Eustermann / Kevin Schall / Dirk Kostrewa / Kristina Lakomek / Mike Strauss / Manuela Moldt / Karl-Peter Hopfner /  要旨: In the eukaryotic nucleus, DNA is packaged in the form of nucleosomes, each of which comprises about 147 base pairs of DNA wrapped around a histone protein octamer. The position and histone ...In the eukaryotic nucleus, DNA is packaged in the form of nucleosomes, each of which comprises about 147 base pairs of DNA wrapped around a histone protein octamer. The position and histone composition of nucleosomes is governed by ATP-dependent chromatin remodellers such as the 15-subunit INO80 complex . INO80 regulates gene expression, DNA repair and replication by sliding nucleosomes, the exchange of histone H2A.Z with H2A, and the positioning of + 1 and -1 nucleosomes at promoter DNA. The structures and mechanisms of these remodelling reactions are currently unknown. Here we report the cryo-electron microscopy structure of the evolutionarily conserved core of the INO80 complex from the fungus Chaetomium thermophilum bound to a nucleosome, at a global resolution of 4.3 Å and with major parts at 3.7 Å. The INO80 core cradles one entire gyre of the nucleosome through multivalent DNA and histone contacts. An Rvb1/Rvb2 AAA ATPase heterohexamer is an assembly scaffold for the complex and acts as a 'stator' for the motor and nucleosome-gripping subunits. The Swi2/Snf2 ATPase motor binds to nucleosomal DNA at superhelical location -6, unwraps approximately 15 base pairs, disrupts the H2A-DNA contacts and is poised to pump entry DNA into the nucleosome. Arp5 and Ies6 bind superhelical locations -2 and -3 to act as a counter grip for the motor, on the other side of the H2A-H2B dimer. The Arp5 insertion domain forms a grappler element that binds the nucleosome dyad, connects the Arp5 actin-fold and entry DNA over a distance of about 90 Å and packs against histone H2A-H2B near the 'acidic patch'. Our structure together with biochemical data suggests a unified mechanism for nucleosome sliding and histone editing by INO80. The motor is part of a macromolecular ratchet, persistently pumping entry DNA across the H2A-H2B dimer against the Arp5 grip until a large nucleosome translocation step occurs. The transient exposure of H2A-H2B by motor activity as well as differential recognition of H2A.Z and H2A may regulate histone exchange. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_4277.map.gz emd_4277.map.gz | 88.4 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-4277-v30.xml emd-4277-v30.xml emd-4277.xml emd-4277.xml | 34.8 KB 34.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_4277_fsc.xml emd_4277_fsc.xml | 12.2 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_4277.png emd_4277.png | 63 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4277 http://ftp.pdbj.org/pub/emdb/structures/EMD-4277 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4277 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4277 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_4277.map.gz / 形式: CCP4 / 大きさ: 149.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_4277.map.gz / 形式: CCP4 / 大きさ: 149.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : INO80core Nucleosome Complex
+超分子 #1: INO80core Nucleosome Complex
+超分子 #2: INO80core
+超分子 #3: Histone octamer
+超分子 #4: Synthetic deoxyribonucleic acid
+分子 #1: RuvB-like helicase
+分子 #2: RuvB-like helicase
+分子 #3: Ino80
+分子 #4: les2
+分子 #5: Ies6
+分子 #6: Actin related protein 5
+分子 #9: Histone H3.2
+分子 #10: Histone H4
+分子 #11: Histone H2A type 1
+分子 #12: Histone H2B type 1-C/E/F/G/I
+分子 #7: Nucleosomal DNA Strand 1
+分子 #8: Nucleosomal DNA Strand 2
+分子 #13: ADENOSINE-5'-DIPHOSPHATE
+分子 #14: ADENOSINE-5'-TRIPHOSPHATE
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 1 mg/mL |
|---|---|
| 緩衝液 | pH: 8 詳細: 20 mM HEPES pH 8, 60 mM KCl, 0.5% glycerol, 0.25 mM CaCl2, 20 uM ZnCl2, 0.25 mM DTT, 0.05% Octyl-beta-glucoside |
| グリッド | モデル: Quantifoil R2/1 / 材質: COPPER / メッシュ: 200 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 雰囲気: AIR |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 95 % / チャンバー内温度: 281 K / 装置: LEICA EM GP |
| 詳細 | Monodisperse sample: INO80core complex reconstituted with nucleosomal substrate was purified by gelfiltration. Addition of nucleotides or crosslinking was not required. |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 最小 デフォーカス(補正後): 1.3 µm / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy / 最大 デフォーカス(公称値): 3.5 µm Bright-field microscopy / 最大 デフォーカス(公称値): 3.5 µm |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER ホルダー冷却材: NITROGEN |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / デジタル化 - 画像ごとのフレーム数: 1-40 / 撮影したグリッド数: 1 / 実像数: 3992 / 平均電子線量: 59.6 e/Å2 詳細: Images were collected in movie mode with 4 frames per second and 10s total aquisition |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー