+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-4258 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of Ryanodine receptor 1 in nanodiscs in the presence of calcium, ATP and ryanodine | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報ATP-gated ion channel activity /  terminal cisterna / terminal cisterna /  ryanodine receptor complex / ryanodine-sensitive calcium-release channel activity / release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / ossification involved in bone maturation / skin development / cellular response to caffeine / intracellularly gated calcium channel activity / outflow tract morphogenesis ...ATP-gated ion channel activity / ryanodine receptor complex / ryanodine-sensitive calcium-release channel activity / release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / ossification involved in bone maturation / skin development / cellular response to caffeine / intracellularly gated calcium channel activity / outflow tract morphogenesis ...ATP-gated ion channel activity /  terminal cisterna / terminal cisterna /  ryanodine receptor complex / ryanodine-sensitive calcium-release channel activity / release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / ossification involved in bone maturation / skin development / cellular response to caffeine / intracellularly gated calcium channel activity / outflow tract morphogenesis / organelle membrane / ryanodine receptor complex / ryanodine-sensitive calcium-release channel activity / release of sequestered calcium ion into cytosol by sarcoplasmic reticulum / ossification involved in bone maturation / skin development / cellular response to caffeine / intracellularly gated calcium channel activity / outflow tract morphogenesis / organelle membrane /  toxic substance binding / toxic substance binding /  voltage-gated calcium channel activity / skeletal muscle fiber development / release of sequestered calcium ion into cytosol / sarcoplasmic reticulum membrane / cellular response to calcium ion / voltage-gated calcium channel activity / skeletal muscle fiber development / release of sequestered calcium ion into cytosol / sarcoplasmic reticulum membrane / cellular response to calcium ion /  筋小胞体 / 筋小胞体 /  muscle contraction / calcium ion transmembrane transport / muscle contraction / calcium ion transmembrane transport /  calcium channel activity / intracellular calcium ion homeostasis / disordered domain specific binding / protein homotetramerization / transmembrane transporter binding / calcium channel activity / intracellular calcium ion homeostasis / disordered domain specific binding / protein homotetramerization / transmembrane transporter binding /  calmodulin binding / calmodulin binding /  calcium ion binding / calcium ion binding /  ATP binding / ATP binding /  生体膜 / identical protein binding 生体膜 / identical protein binding類似検索 - 分子機能 | |||||||||
| 生物種 |   Oryctolagus cuniculus (ウサギ) / Oryctolagus cuniculus (ウサギ) /   Rabbit (ウサギ) Rabbit (ウサギ) | |||||||||
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 7.3 Å クライオ電子顕微鏡法 / 解像度: 7.3 Å | |||||||||
 データ登録者 データ登録者 | Willegems K / Efremov RG | |||||||||
| 資金援助 |  ベルギー, 2件 ベルギー, 2件
| |||||||||
 引用 引用 |  ジャーナル: Structure / 年: 2018 ジャーナル: Structure / 年: 2018タイトル: Influence of Lipid Mimetics on Gating of Ryanodine Receptor. 著者: Katrien Willegems / Rouslan G Efremov /  要旨: Understanding gating principles of ion channels at high resolution is of great importance. Here we investigate the conformational transition from closed to open state in ryanodine receptor 1 (RyR1) ...Understanding gating principles of ion channels at high resolution is of great importance. Here we investigate the conformational transition from closed to open state in ryanodine receptor 1 (RyR1) reconstituted into lipid nanodiscs. RyR1 is a homotetrameric giant ion channel that couples excitation of muscle cells to fast calcium release from the sarcoplasmic reticulum. Using single-particle cryo-EM we show that RyR1 reconstituted into lipid nanodiscs is stabilized in the open conformation when bound to the plant toxin ryanodine, but not in the presence of its physiological activators, calcium and ATP. Further, using ryanodine binding assays we show that membrane mimetics influence RyR1 transition between closed and open-channel conformations. We find that all detergents, including fluorinated detergents added to nanodiscs, stabilize closed state of RyR1. Our biochemical results correlate with available structural data and suggest optimal conditions for structural studies of RyR1 gating. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_4258.map.gz emd_4258.map.gz | 195.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-4258-v30.xml emd-4258-v30.xml emd-4258.xml emd-4258.xml | 25.8 KB 25.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
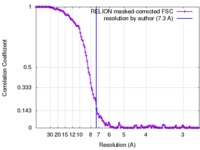
| FSC (解像度算出) |  emd_4258_fsc.xml emd_4258_fsc.xml | 13 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_4258.png emd_4258.png | 99.8 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4258 http://ftp.pdbj.org/pub/emdb/structures/EMD-4258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4258 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4258 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_4258.map.gz / 形式: CCP4 / 大きさ: 209.3 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_4258.map.gz / 形式: CCP4 / 大きさ: 209.3 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ボクセルのサイズ | X=Y=Z: 1.355 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
- 試料の構成要素
試料の構成要素
-全体 : Ryanodine receptor 1 reconstituted in lipid nanodiscs in the pres...
| 全体 | 名称: Ryanodine receptor 1 reconstituted in lipid nanodiscs in the presence of calcium, ATP and ryanodine |
|---|---|
| 要素 |
|
-超分子 #1: Ryanodine receptor 1 reconstituted in lipid nanodiscs in the pres...
| 超分子 | 名称: Ryanodine receptor 1 reconstituted in lipid nanodiscs in the presence of calcium, ATP and ryanodine タイプ: complex / ID: 1 / 親要素: 0 / 含まれる分子: #1 |
|---|---|
| 由来(天然) | 生物種:   Oryctolagus cuniculus (ウサギ) / 株: New Zealand white rabbit / 組織: skeletal muscle / 細胞中の位置: sarcoplasmic reticulum Oryctolagus cuniculus (ウサギ) / 株: New Zealand white rabbit / 組織: skeletal muscle / 細胞中の位置: sarcoplasmic reticulum |
| 分子量 | 理論値: 2.2 MDa |
-分子 #1: Ryanodine receptor 1
| 分子 | 名称: Ryanodine receptor 1 / タイプ: protein_or_peptide / ID: 1 詳細: Poly Unk stretches correspond to polypeptide regions where the register is not assigned. コピー数: 4 / 光学異性体: LEVO |
|---|---|
| 由来(天然) | 生物種:   Rabbit (ウサギ) / 株: New Zealand white rabbit / 組織: skeletal muscle Rabbit (ウサギ) / 株: New Zealand white rabbit / 組織: skeletal muscle |
| 分子量 | 理論値: 532.033062 KDa |
| 配列 | 文字列: MGDGGEGEDE VQFLRTDDEV VLQCSATVLK EQLKLCLAAE GFGNRLCFLE PTSNAQNVPP DLAICCFTLE QSLSVRALQE MLANTVEAG VESSQGGGHR TLLYGHAILL RHAHSRMYLS CLTTSRSMTD KLAFDVGLQE DATGEACWWT MHPASKQRSE G EKVRVGDD ...文字列: MGDGGEGEDE VQFLRTDDEV VLQCSATVLK EQLKLCLAAE GFGNRLCFLE PTSNAQNVPP DLAICCFTLE QSLSVRALQE MLANTVEAG VESSQGGGHR TLLYGHAILL RHAHSRMYLS CLTTSRSMTD KLAFDVGLQE DATGEACWWT MHPASKQRSE G EKVRVGDD LILVSVSSER YLHLSTASGE LQVDASFMQT LWNMNPICSC CEEGYVTGGH VLRLFHGHMD ECLTISAADS DD QRRLVYY EGGAVCTHAR SLWRLEPLRI SWSGSHLRWG QPLRIRHVTT GRYLALTEDQ GLVVVDACKA HTKATSFCFR VSK EKLDTA PKRDVEGMGP PEIKYGESLC FVQHVASGLW LTYAAPDPKA LRLGVLKKKA ILHQEGHMDD ALFLTRCQQE ESQA ARMIH STAGLYNQFI KGLDSFSGKP RGSGPPAGPA LPIEAVILSL QDLIGYFEPP SEELQHEEKQ SKLRSLRNRQ SLFQE EGML SLVLNCIDRL NVYTTAAHFA EYAGEEAAES WKEIVNLLYE LLASLIRGNR ANCALFSTNL DWVVSKLDRL EASSGI LEV LYCVLIESPE VLNIIQENHI KSIISLLDKH GRNHKVLDVL CSLCVCNGVA VRSNQDLITE NLLPGRELLL QTNLINY VT SIRPNIFVGR AEGSTQYGKW YFEVMVDEVV PFLTAQATHL RVGWALTEGY SPYPGGGEGW GGNGVGDDLY SYGFDGLH L WTGHVARPVT SPGQHLLAPE DVVSCCLDLS VPSISFRING CPVQGVFEAF NLDGLFFPVV SFSAGVKVRF LLGGRHGEF KFLPPPGYAP CHEAVLPRER LRLEPIKEYR REGPRGPHLV GPSRCLSHTD FVPCPVDTVQ IVLPPHLERI REKLAENIHE LWALTRIEQ GWTYGPVRDD NKRLHPCLVN FHSLPEPERN YNLQMSGETL KTLLALGCHV GMADEKAEDN LKKTKLPKTY M MSNGYKPA PLDLSHVRLT PAQTTLVDRL AENGHNVWAR DRVAQGWSYS AVQDIPARRN PRLVPYRLLD EATKRSNRDS LC QAVRTLL GYGYNIEPPD QEPSQVENQS RWDRVRIFRA EKSYTVQSGR WYFEFEAVTT GEMRVGWARP ELRPDVELGA DEL AYVFNG HRGQRWHLGS EPFGRPWQSG DVVGCMIDLT ENTIIFTLNG EVLMSDSGSE TAFREIEIGD GFLPVCSLGP GQVG HLNLG QDVSSLRFFA ICGLQEGFEP FAINMQRPVT TWFSKSLPQF EPVPPEHPHY EVARMDGTVD TPPCLRLAHR (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)MPLS AAMFLSERKN PAPQCPPRLE VQMLMPVSWS RMPNHF LQV ETRRAGERLG WAVQCQDPLT MMALHIPEEN RCMDILELSE RLDLQRFHSH TLRLYRAVCA LGNNRVAHAL CSHVDQA QL LHALEDAHLP GPLRAGYYDL LISIHLESAC RSRRSMLSEY IVPLTPETRA ITLFPPGRKG GNARRHGLPG VGVTTSLR P PHHFSPPCFV AALPAAGVAE APARLSPAIP LEALRDKALR MLGEAVRDGG QHARDPVGGS VEFQFVPVLK LVSTLLVMG IFGDEDVKQI LKMIEPEVFT EEEEEEEEEE EEEEEEEEDE EEKEEDEEEE EKEDAEKEEE EAPEGEKEDL EEGLLQMKLP ESVKLQMCN LLEYFCDQEL QHRVESLAAF AERYVDKLQA NQRSRYALLM RAFTMSAAET ARRTREFRSP PQEQINMLLH F KDEADEED CPLPEDIRQD LQDFHQDLLA HCGIQLEGEE EEPEEETSLS SRLRSLLETV RLVKKKEEKP EEELPAEEKK PQ SLQELVS HMVVRWAQED YVQSPELVRA MFSLLHRQYD GLGELLRALP RAYTISPSSV EDTMSLLECL GQIRSLLIVQ MGP QEENLM IQSIGNIMNN KVFYQHPNLM RALGMHETVM EVMVNVLGGG ETKEIRFPKM VTSCCRFLCY FCRISRQNQR SMFD HLSYL LENSGIGLGM QGSTPLDVAA ASVIDNNELA LALQEQDLEK VVSYLAGCGL QSCPMLLAKG YPDIGWNPCG GERYL DFLR FAVFVNGESV EENANVVVRL LIRKPECFGP ALRGEGGSGL LAAIEEAIRI SEDPARDGPG VRRDRRREHF GEEPPE ENR VHLGHAIMSF YAALIDLLGR CAPEMHLIQA GKGEALRIRA ILRSLVPLDD LVGIISLPLQ IPTL(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)NFDPRP VETLNVIIPE KLDS FINKF AEYTHEKWAF DKIQNNWSYG ENVDEELKTH PMLRPYKTFS EKDKEIYRWP IKESLKAMIA WEWTIEKARE GEEER TEKK KTRKISQTAQ TYDPREGYNP QPPDLSGVTL SRELQAMAEQ LAENYHNTWG RKKKQELEAK GGGTHPLLVP YDTLTA KEK ARDREKAQEL LKFLQMNGYA VTR(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)TPLY NLPTHRACNM FLESYKAAWI LTEDHSFEDR MIDDLSKAGE QEEEEEEVEE KKPDPLHQLV LHFSRTALTE K SKLDEDYL YMAYADIMAK SCHLEEGGEN GEAEEEEVEV SFEEKEMEKQ RLLYQQSRLH TRGAAEMVLQ MISACKGETG AM VSSTLKL GISILNGGNA EVQQKMLDYL KDKKEVGFFQ SIQALMQTCS VLDLNAFERQ NKAEGLGMVN EDGTVINRQN GEK VMADDE FTQDLFRFLQ LLCEGHNNDF QNYLRTQTGN TTTINIIICT VDYLLRLQES ISDFYWYYSG KDVIEEQGKR NFSK AMSVA KQVFNSLTEY IQGPCTGNQQ SLAHSRLWDA VVGFLHVFAH MMMKLAQDSS QIELLKELLD LQKDMVVMLL SLLEG NVVN GMIARQMVDM LVESSSNVEM ILKFFDMFLK LKDIVGSEAF QDYVTDPRGL ISKKDFQKAM DSQKQFTGPE IQFLLS CSE ADENEMINFE EFANRFQEPA RDIGFNVAVL LTNLSEHVPH DPRLRNFLEL AESILEYFRP YLGRIEIMGA SRRIERI YF EISETNRAQW EMPQVKESKR QFIFDVVNEG GEAEKMELFV SFCEDTIFEM QIAAQISEPE GEPEADEDEG MGEAAAEG A EEGAAGAEGA AGTVAAGATA RLAAAAARAL RGLSYRSLRR RVRRLRRLTA REAATALAAL LWAVVARAGA AGAGAAAGA LRLLWGSLFG GGLVEGAKKV TVTELLAGMP DPTSDEVHGE QPAGPGGDAD GAGEGEGEGD AAEGDGDEEV AGHEAGPGGA EGVVAVADG GPFRPEGAGG LGDMGDTTPA EPPTPEGSPI LKRKLGVDGE EEELVPEPEP EPEPEPEKAD EENGEKEEVP E APPEPPKK APPSPPAKKE EAGGAGMEFW GELEVQRVKF LNYLSRNFYT LRFLALFLAF AINFILLFYK VSDSPPGEDD ME GSAAGDL AGAGSGGGSG WGSGAGEEAE GDEDENMVYY FLEESTGYME PALWCLSLLH TLVAFLCIIG YNCLKVPLVI FKR EKELAR KLEFDGLYIT EQPGDDDVKG QWDRLVLNTP SFPSNYWDKF VKRKVLDKHG DIFGRERIAE LLGMDLASLE ITAH NERKP DPPPGLLTWL MSIDVKYQIW KFGVIFTDNS FLYLGWYMVM SLLGHYNNFF FAAHLLDIAM GVKTLRTILS SVTHN GKQL VMTVGLLAVV VYLYTVVAFN FFRKFYNKSE DEDEPDMKCD DMMTCYLFHM YVGVRAGGGI GDEIEDPAGD EYELYR VVF DITFFFFVIV ILLAIIQGLI IDAFGELRDQ QEQVKEDMET KCFICGIGSD YFDTTPHGFE THTLEEHNLA NYMFFLM YL INKDETEHTG QESYVWKMYQ ERCWDFFPAG DCFRKQYEDQ LS |
-分子 #2: ZINC ION
| 分子 | 名称: ZINC ION / タイプ: ligand / ID: 2 / コピー数: 4 / 式: ZN |
|---|---|
| 分子量 | 理論値: 65.409 Da |
-分子 #3: CALCIUM ION
| 分子 | 名称: CALCIUM ION / タイプ: ligand / ID: 3 / コピー数: 4 / 式: CA |
|---|---|
| 分子量 | 理論値: 40.078 Da |
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 3 mg/mL | |||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.4 構成要素:
| |||||||||||||||||||||||||||
| グリッド | モデル: Quantifoil R2/1 / 材質: COPPER / メッシュ: 300 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 前処理 - タイプ: GLOW DISCHARGE / 前処理 - 雰囲気: AIR / 前処理 - 気圧: 0.04 kPa 詳細: Glow discharged in ELMO plasma cleaner at current of 3 mA for 1 min. | |||||||||||||||||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内温度: 293.15 K / 装置: HOMEMADE PLUNGER |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 2.0 µm / 倍率(公称値): 59000 Bright-field microscopy / 最大 デフォーカス(公称値): 3.0 µm / 最小 デフォーカス(公称値): 2.0 µm / 倍率(公称値): 59000 |
| 試料ステージ | ホルダー冷却材: NITROGEN |
| 撮影 | フィルム・検出器のモデル: FEI FALCON II (4k x 4k) 検出モード: COUNTING / デジタル化 - サイズ - 横: 4096 pixel / デジタル化 - サイズ - 縦: 4096 pixel / デジタル化 - サンプリング間隔: 14.0 µm / デジタル化 - 画像ごとのフレーム数: 1-7 / 撮影したグリッド数: 1 / 実像数: 3078 / 平均露光時間: 1.0 sec. / 平均電子線量: 28.0 e/Å2 |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
- 画像解析
画像解析
-原子モデル構築 1
| 精密化 | 空間: REAL / プロトコル: FLEXIBLE FIT |
|---|---|
| 得られたモデル |  PDB-6fg3: |
 ムービー
ムービー コントローラー
コントローラー