+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-3802 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | The cryo-EM structure of human TFIIH | |||||||||
 マップデータ マップデータ | Postprocessed (b-factor sharpened, low pass filtered) masked map used for atomic coordinate refinement. | |||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報MMXD complex / core TFIIH complex portion of holo TFIIH complex / positive regulation of DNA helicase activity / negative regulation of DNA helicase activity / Cytosolic iron-sulfur cluster assembly / nucleotide-excision repair, DNA duplex unwinding /  central nervous system myelin formation / positive regulation of mitotic recombination / hair cell differentiation / central nervous system myelin formation / positive regulation of mitotic recombination / hair cell differentiation /  ventricular system development ...MMXD complex / core TFIIH complex portion of holo TFIIH complex / positive regulation of DNA helicase activity / negative regulation of DNA helicase activity / Cytosolic iron-sulfur cluster assembly / nucleotide-excision repair, DNA duplex unwinding / ventricular system development ...MMXD complex / core TFIIH complex portion of holo TFIIH complex / positive regulation of DNA helicase activity / negative regulation of DNA helicase activity / Cytosolic iron-sulfur cluster assembly / nucleotide-excision repair, DNA duplex unwinding /  central nervous system myelin formation / positive regulation of mitotic recombination / hair cell differentiation / central nervous system myelin formation / positive regulation of mitotic recombination / hair cell differentiation /  ventricular system development / hair follicle maturation / cyclin-dependent protein kinase activating kinase holoenzyme complex / nucleotide-excision repair factor 3 complex / nucleotide-excision repair, preincision complex assembly / ventricular system development / hair follicle maturation / cyclin-dependent protein kinase activating kinase holoenzyme complex / nucleotide-excision repair factor 3 complex / nucleotide-excision repair, preincision complex assembly /  紫外線 / CAK-ERCC2 complex / transcription factor TFIIK complex / embryonic cleavage / 5'-3' DNA helicase activity / adult heart development / G protein-coupled receptor internalization / transcription factor TFIIH holo complex / transcription factor TFIIH core complex / cyclin-dependent protein serine/threonine kinase activator activity / 3'-5' DNA helicase activity / nuclear thyroid hormone receptor binding / 紫外線 / CAK-ERCC2 complex / transcription factor TFIIK complex / embryonic cleavage / 5'-3' DNA helicase activity / adult heart development / G protein-coupled receptor internalization / transcription factor TFIIH holo complex / transcription factor TFIIH core complex / cyclin-dependent protein serine/threonine kinase activator activity / 3'-5' DNA helicase activity / nuclear thyroid hormone receptor binding /  転写開始前複合体 / RNA Polymerase I Transcription Termination / regulation of mitotic cell cycle phase transition / hematopoietic stem cell proliferation / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / spinal cord development / RNA polymerase II general transcription initiation factor activity / transcription factor TFIID complex / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping / 転写開始前複合体 / RNA Polymerase I Transcription Termination / regulation of mitotic cell cycle phase transition / hematopoietic stem cell proliferation / RNA Pol II CTD phosphorylation and interaction with CE during HIV infection / RNA Pol II CTD phosphorylation and interaction with CE / spinal cord development / RNA polymerase II general transcription initiation factor activity / transcription factor TFIID complex / Formation of the Early Elongation Complex / Formation of the HIV-1 Early Elongation Complex / mRNA Capping /  bone mineralization / HIV Transcription Initiation / RNA Polymerase II HIV Promoter Escape / Transcription of the HIV genome / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / erythrocyte maturation / regulation of cyclin-dependent protein serine/threonine kinase activity / ATPase activator activity / regulation of G1/S transition of mitotic cell cycle / RNA Polymerase I Transcription Initiation / transcription elongation by RNA polymerase I / DNA topological change / intrinsic apoptotic signaling pathway by p53 class mediator / transcription-coupled nucleotide-excision repair / hematopoietic stem cell differentiation / Tat-mediated elongation of the HIV-1 transcript / Formation of HIV-1 elongation complex containing HIV-1 Tat / embryonic organ development / transcription by RNA polymerase I / Formation of HIV elongation complex in the absence of HIV Tat / Cyclin E associated events during G1/S transition / RNA Polymerase II Transcription Elongation / Cyclin A/B1/B2 associated events during G2/M transition / Formation of RNA Pol II elongation complex / response to UV / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Cyclin A:Cdk2-associated events at S phase entry / RNA Polymerase II Pre-transcription Events / bone mineralization / HIV Transcription Initiation / RNA Polymerase II HIV Promoter Escape / Transcription of the HIV genome / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Initiation And Promoter Clearance / erythrocyte maturation / regulation of cyclin-dependent protein serine/threonine kinase activity / ATPase activator activity / regulation of G1/S transition of mitotic cell cycle / RNA Polymerase I Transcription Initiation / transcription elongation by RNA polymerase I / DNA topological change / intrinsic apoptotic signaling pathway by p53 class mediator / transcription-coupled nucleotide-excision repair / hematopoietic stem cell differentiation / Tat-mediated elongation of the HIV-1 transcript / Formation of HIV-1 elongation complex containing HIV-1 Tat / embryonic organ development / transcription by RNA polymerase I / Formation of HIV elongation complex in the absence of HIV Tat / Cyclin E associated events during G1/S transition / RNA Polymerase II Transcription Elongation / Cyclin A/B1/B2 associated events during G2/M transition / Formation of RNA Pol II elongation complex / response to UV / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / Cyclin A:Cdk2-associated events at S phase entry / RNA Polymerase II Pre-transcription Events /  DNA helicase activity / hormone-mediated signaling pathway / extracellular matrix organization / post-embryonic development / insulin-like growth factor receptor signaling pathway / DNA helicase activity / hormone-mediated signaling pathway / extracellular matrix organization / post-embryonic development / insulin-like growth factor receptor signaling pathway /  chromosome segregation / determination of adult lifespan / promoter-specific chromatin binding / transcription elongation by RNA polymerase II / chromosome segregation / determination of adult lifespan / promoter-specific chromatin binding / transcription elongation by RNA polymerase II /  transcription initiation at RNA polymerase II promoter / nucleotide-excision repair / RNA Polymerase I Promoter Escape / TP53 Regulates Transcription of DNA Repair Genes / positive regulation of smooth muscle cell proliferation / G1/S transition of mitotic cell cycle / multicellular organism growth / cellular response to gamma radiation / NoRC negatively regulates rRNA expression / transcription initiation at RNA polymerase II promoter / nucleotide-excision repair / RNA Polymerase I Promoter Escape / TP53 Regulates Transcription of DNA Repair Genes / positive regulation of smooth muscle cell proliferation / G1/S transition of mitotic cell cycle / multicellular organism growth / cellular response to gamma radiation / NoRC negatively regulates rRNA expression /  protein localization / Dual Incision in GG-NER / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / spindle / Formation of TC-NER Pre-Incision Complex / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / response to calcium ion / Gap-filling DNA repair synthesis and ligation in TC-NER / Cyclin D associated events in G1 / protein-macromolecule adaptor activity / RUNX1 regulates transcription of genes involved in differentiation of HSCs / 4 iron, 4 sulfur cluster binding protein localization / Dual Incision in GG-NER / Transcription-Coupled Nucleotide Excision Repair (TC-NER) / spindle / Formation of TC-NER Pre-Incision Complex / Formation of Incision Complex in GG-NER / Dual incision in TC-NER / response to calcium ion / Gap-filling DNA repair synthesis and ligation in TC-NER / Cyclin D associated events in G1 / protein-macromolecule adaptor activity / RUNX1 regulates transcription of genes involved in differentiation of HSCs / 4 iron, 4 sulfur cluster binding類似検索 - 分子機能 | |||||||||
| 生物種 |   Homo sapiens (ヒト) / Homo sapiens (ヒト) /   Human (ヒト) Human (ヒト) | |||||||||
| 手法 |  単粒子再構成法 / 単粒子再構成法 /  クライオ電子顕微鏡法 / 解像度: 4.4 Å クライオ電子顕微鏡法 / 解像度: 4.4 Å | |||||||||
 データ登録者 データ登録者 | Greber BJ / Nguyen THD / Fang J / Afonine PV / Adams PD / Nogales E | |||||||||
| 資金援助 |  米国, 2件 米国, 2件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2017 ジャーナル: Nature / 年: 2017タイトル: The cryo-electron microscopy structure of human transcription factor IIH. 著者: Basil J Greber / Thi Hoang Duong Nguyen / Jie Fang / Pavel V Afonine / Paul D Adams / Eva Nogales /  要旨: Human transcription factor IIH (TFIIH) is part of the general transcriptional machinery required by RNA polymerase II for the initiation of eukaryotic gene transcription. Composed of ten subunits ...Human transcription factor IIH (TFIIH) is part of the general transcriptional machinery required by RNA polymerase II for the initiation of eukaryotic gene transcription. Composed of ten subunits that add up to a molecular mass of about 500 kDa, TFIIH is also essential for nucleotide excision repair. The seven-subunit TFIIH core complex formed by XPB, XPD, p62, p52, p44, p34, and p8 is competent for DNA repair, while the CDK-activating kinase subcomplex, which includes the kinase activity of CDK7 as well as the cyclin H and MAT1 subunits, is additionally required for transcription initiation. Mutations in the TFIIH subunits XPB, XPD, and p8 lead to severe premature ageing and cancer propensity in the genetic diseases xeroderma pigmentosum, Cockayne syndrome, and trichothiodystrophy, highlighting the importance of TFIIH for cellular physiology. Here we present the cryo-electron microscopy structure of human TFIIH at 4.4 Å resolution. The structure reveals the molecular architecture of the TFIIH core complex, the detailed structures of its constituent XPB and XPD ATPases, and how the core and kinase subcomplexes of TFIIH are connected. Additionally, our structure provides insight into the conformational dynamics of TFIIH and the regulation of its activity. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_3802.map.gz emd_3802.map.gz | 4.6 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-3802-v30.xml emd-3802-v30.xml emd-3802.xml emd-3802.xml | 36.8 KB 36.8 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| 画像 |  emd_3802.png emd_3802.png | 140.7 KB | ||
| マスクデータ |  emd_3802_msk_1.map emd_3802_msk_1.map | 64 MB |  マスクマップ マスクマップ | |
| その他 |  emd_3802_half_map_1.map.gz emd_3802_half_map_1.map.gz emd_3802_half_map_2.map.gz emd_3802_half_map_2.map.gz | 49.7 MB 49.7 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3802 http://ftp.pdbj.org/pub/emdb/structures/EMD-3802 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3802 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3802 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_3802.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_3802.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Postprocessed (b-factor sharpened, low pass filtered) masked map used for atomic coordinate refinement. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
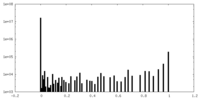
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-マスク #1
| ファイル |  emd_3802_msk_1.map emd_3802_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
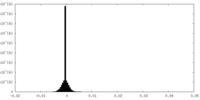
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: None
| ファイル | emd_3802_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | None | ||||||||||||
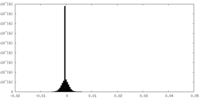
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: None
| ファイル | emd_3802_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | None | ||||||||||||
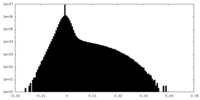
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : TFIIH
+超分子 #1: TFIIH
+超分子 #2: TFIIH core complex
+超分子 #3: TFIIH CDK-activating kinase (CAK) subcomplex
+分子 #1: TFIIH basal transcription factor complex helicase XPB subunit,XPB...
+分子 #2: TFIIH basal transcription factor complex helicase XPD subunit
+分子 #3: General transcription factor IIH subunit 4,p52,General transcript...
+分子 #4: General transcription factor IIH subunit 2
+分子 #5: General transcription factor IIH subunit 3
+分子 #6: General transcription factor IIH subunit 5
+分子 #7: MAT1
+分子 #8: Unassigned secondary structure elements.
+分子 #9: Unassigned secondary structure elements (p52 region)
+分子 #10: Unassigned secondary structure elements (XPB NTE region)
+分子 #11: IRON/SULFUR CLUSTER
-実験情報
-構造解析
| 手法 |  クライオ電子顕微鏡法 クライオ電子顕微鏡法 |
|---|---|
 解析 解析 |  単粒子再構成法 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.0049 mg/mL | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 緩衝液 | pH: 7.8 構成要素:
| ||||||||||||||||||||||||
| グリッド | モデル: C-flat-4/2 / 材質: COPPER / メッシュ: 400 / 支持フィルム - #0 - Film type ID: 1 / 支持フィルム - #0 - 材質: CARBON / 支持フィルム - #0 - トポロジー: HOLEY / 支持フィルム - #1 - Film type ID: 2 / 支持フィルム - #1 - 材質: CARBON / 支持フィルム - #1 - トポロジー: CONTINUOUS / 前処理 - タイプ: PLASMA CLEANING / 前処理 - 雰囲気: AIR | ||||||||||||||||||||||||
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / チャンバー内温度: 278.15 K / 装置: FEI VITROBOT MARK IV / 詳細: 3-4 minute incubation, 2 second blot. | ||||||||||||||||||||||||
| 詳細 | Natively purified complex at approx. 10 nM concentration |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN |
|---|---|
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | C2レンズ絞り径: 50.0 µm / 倍率(補正後): 37879 / 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / 最大 デフォーカス(公称値): 4.5 µm / 最小 デフォーカス(公称値): 2.0 µm Bright-field microscopy / Cs: 2.7 mm / 最大 デフォーカス(公称値): 4.5 µm / 最小 デフォーカス(公称値): 2.0 µm |
| 試料ステージ | 試料ホルダーモデル: GATAN 626 SINGLE TILT LIQUID NITROGEN CRYO TRANSFER HOLDER ホルダー冷却材: NITROGEN |
| 撮影 | フィルム・検出器のモデル: GATAN K2 SUMMIT (4k x 4k) 検出モード: COUNTING / デジタル化 - サイズ - 横: 3840 pixel / デジタル化 - サイズ - 縦: 3710 pixel / デジタル化 - サンプリング間隔: 5.0 µm / デジタル化 - 画像ごとのフレーム数: 1-30 / 撮影したグリッド数: 4 / 実像数: 8300 / 平均露光時間: 8.7 sec. / 平均電子線量: 40.0 e/Å2 詳細: 8300 micrographs collected in four session with identical acquisition settings. Sessions lasted 4, 2, 4, and 4 days and yielded 1200, 1700, 2800, and 2600 micrographs. |
- 画像解析
画像解析
| 粒子像選択 | 選択した数: 1500000 詳細: Initial rounds of particle selection using DogPicker within APPION to generate templates for subsequent RELION autopicking. |
|---|---|
| CTF補正 | ソフトウェア - 名称: CTFFIND (ver. 4) / ソフトウェア - 詳細: run from within RELION 詳細: Correction in RELION based on values determined in CTFFIND4. |
| 初期モデル | モデルのタイプ: EMDB MAP EMDB ID: 詳細: Box and pixel size were adjusted before initial refinement. |
| 初期 角度割当 | タイプ: PROJECTION MATCHING / ソフトウェア - 名称: RELION (ver. 1.4) / 詳細: Maximum-likelihood 3D classification within RELION. |
| 最終 3次元分類 | ソフトウェア - 名称: RELION (ver. 1.4) 詳細: Combination of individually sub-classified datasets (with variable number of classes). |
| 最終 角度割当 | タイプ: PROJECTION MATCHING / ソフトウェア - 名称: RELION (ver. 1.4) / 詳細: Maximum-likelihood 3D refinement within RELION. |
| 最終 再構成 | 使用したクラス数: 3 / 想定した対称性 - 点群: C1 (非対称) / アルゴリズム: FOURIER SPACE / 解像度のタイプ: BY AUTHOR / 解像度: 4.4 Å / 解像度の算出法: FSC 0.143 CUT-OFF / ソフトウェア - 名称: RELION (ver. 1.4) / 詳細: RELION 3D auto-refinement (gold standard). / 使用した粒子像数: 122900 |
| 詳細 | Micrographs were inspected for quality of Thon rings and ice contamination. Poor micrographs were rejected. |
-原子モデル構築 1
| 詳細 | Initial model assembled from high-resolution structures and homology models. Subsequently rebuilt in O and COOT, refined into the cryo-EM map using PHENIX and REFMAC, and fully validated. |
|---|---|
| 精密化 | 空間: REAL / プロトコル: RIGID BODY FIT |
| 得られたモデル |  PDB-5of4:  PDB-6o9m: |
 ムービー
ムービー コントローラー
コントローラー
































 Z
Z Y
Y X
X