[English] 日本語
 Yorodumi
Yorodumi- EMDB-7617: Rabbit muscle aldolase at 2.5 A resolution (17sep21j less 25nm ic... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-7617 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Rabbit muscle aldolase at 2.5 A resolution (17sep21j less 25nm ice thickness, 124k particles) | |||||||||
 Map data Map data | Rabbit muscle aldolase at 2.5 A resolution (17sep31j <25nm ice thickness, 124K particles) | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) | |||||||||
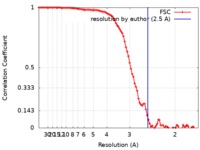
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 2.5 Å cryo EM / Resolution: 2.5 Å | |||||||||
 Authors Authors | Kim LK / Rice WJ / Eng ET / Kopylov M / Cheng A / Raczkowski AM / Jordan KD / Bobe D / Potter CS / Carragher B | |||||||||
 Citation Citation |  Journal: Front Mol Biosci / Year: 2018 Journal: Front Mol Biosci / Year: 2018Title: Benchmarking cryo-EM Single Particle Analysis Workflow. Authors: Laura Y Kim / William J Rice / Edward T Eng / Mykhailo Kopylov / Anchi Cheng / Ashleigh M Raczkowski / Kelsey D Jordan / Daija Bobe / Clinton S Potter / Bridget Carragher /  Abstract: Cryo electron microscopy facilities running multiple instruments and serving users with varying skill levels need a robust and reliable method for benchmarking both the hardware and software ...Cryo electron microscopy facilities running multiple instruments and serving users with varying skill levels need a robust and reliable method for benchmarking both the hardware and software components of their single particle analysis workflow. The workflow is complex, with many bottlenecks existing at the specimen preparation, data collection and image analysis steps; the samples and grid preparation can be of unpredictable quality, there are many different protocols for microscope and camera settings, and there is a myriad of software programs for analysis that can depend on dozens of settings chosen by the user. For this reason, we believe it is important to benchmark the entire workflow, using a standard sample and standard operating procedures, on a regular basis. This provides confidence that all aspects of the pipeline are capable of producing maps to high resolution. Here we describe benchmarking procedures using a test sample, rabbit muscle aldolase. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_7617.map.gz emd_7617.map.gz | 59.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-7617-v30.xml emd-7617-v30.xml emd-7617.xml emd-7617.xml | 15 KB 15 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_7617_fsc.xml emd_7617_fsc.xml | 10.6 KB | Display |  FSC data file FSC data file |
| Images |  emd_7617.png emd_7617.png | 35.5 KB | ||
| Others |  emd_7617_half_map_1.map.gz emd_7617_half_map_1.map.gz emd_7617_half_map_2.map.gz emd_7617_half_map_2.map.gz | 59.2 MB 59.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-7617 http://ftp.pdbj.org/pub/emdb/structures/EMD-7617 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7617 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-7617 | HTTPS FTP |
-Related structure data
| Related structure data |  7528C  7541C  7550C  7551C  7562C  7614C  7615C  7616C C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10182 (Title: Benchmarking cryo-EM single particle analysis workflow EMPIAR-10182 (Title: Benchmarking cryo-EM single particle analysis workflowData size: 37.3 Data #1: aligned micrographs of rabbit muscle aldolase 17sep21j [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_7617.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_7617.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Rabbit muscle aldolase at 2.5 A resolution (17sep31j <25nm ice thickness, 124K particles) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: Rabbit muscle aldolase at 2.5 A resolution (17sep31j...
| File | emd_7617_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Rabbit muscle aldolase at 2.5 A resolution (17sep31j <25nm ice thickness, 124K particles), half map 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
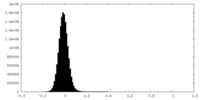
| Density Histograms |
-Half map: Rabbit muscle aldolase at 2.5 A resolution (17sep31j...
| File | emd_7617_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Rabbit muscle aldolase at 2.5 A resolution (17sep31j <25nm ice thickness, 124K particles), half map 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
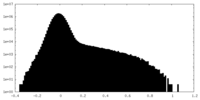
| Density Histograms |
- Sample components
Sample components
-Entire : Rabbit muscle aldolase at 2.5 A resolution (17sep21j <25nm ice th...
| Entire | Name: Rabbit muscle aldolase at 2.5 A resolution (17sep21j <25nm ice thickness, 124K particles) |
|---|---|
| Components |
|
-Supramolecule #1: Rabbit muscle aldolase at 2.5 A resolution (17sep21j <25nm ice th...
| Supramolecule | Name: Rabbit muscle aldolase at 2.5 A resolution (17sep21j <25nm ice thickness, 124K particles) / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Molecular weight | Experimental: 150 KDa |
-Macromolecule #1: Rabbit muscle aldolase
| Macromolecule | Name: Rabbit muscle aldolase / type: protein_or_peptide / ID: 1 / Enantiomer: DEXTRO / EC number:  fructose-bisphosphate aldolase fructose-bisphosphate aldolase |
|---|---|
| Source (natural) | Organism:   Oryctolagus cuniculus (rabbit) Oryctolagus cuniculus (rabbit) |
| Recombinant expression | Organism:   Escherichia coli (E. coli) Escherichia coli (E. coli) |
| Sequence | String: PHSHPALTPE QKKELSDIAH RIVAPGKGIL AADESTGSIA KRLQSIGTEN TEENRRFYRQ LLLTADDRVN PCIGGVILFH ETLYQKADD GRPFPQVIKS KGGVVGIKVD KGVVPLAGTN GETTTQGLDG LSERCAQYKK DGADFAAWRC VLKIGEHTPS A LAIMENAN ...String: PHSHPALTPE QKKELSDIAH RIVAPGKGIL AADESTGSIA KRLQSIGTEN TEENRRFYRQ LLLTADDRVN PCIGGVILFH ETLYQKADD GRPFPQVIKS KGGVVGIKVD KGVVPLAGTN GETTTQGLDG LSERCAQYKK DGADFAAWRC VLKIGEHTPS A LAIMENAN VLARYASICQ QNGIVPIVEP EILPDGDHDL KRCQYVTEKV LAAVYKALSD HHIYLEGTLL KPNMVTPGHA CT QKYSHEE IAMATVTALR RTVPPAVTGV TFLSGGQSEE EASINLNAIN KCPLLKPWAL TFSYGRALQA SALKAWGGKK ENL KAAQEE YVKRALANSL ACQGKYTPSG QAGAAASESL FISNHAY |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Average electron dose: 65.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z
Z Y
Y X
X