-Search query
-Search result
Showing 1 - 50 of 178 items for (author: krammer & f)
EMDB-41874: 
CryoEM structure of A/Solomon Islands/3/2006 H1 HA in complex with 05.GC.w2.3C10-H1_SI06

EMDB-28728: 
Structure of 3A10 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase

EMDB-28729: 
Structure of 1F04 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase

EMDB-28730: 
Structure of 3C08 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase

PDB-8ez3: 
Structure of 3A10 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase

PDB-8ez7: 
Structure of 1F04 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase

PDB-8ez8: 
Structure of 3C08 Fab in complex with A/Moscow/10/1999 (H3N2) influenza virus neuraminidase

EMDB-27920: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

EMDB-27921: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

PDB-8e6j: 
3H03 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

PDB-8e6k: 
2H08 Fab in complex with influenza virus neuraminidase from A/Brevig Mission/1/1918 (H1N1)

EMDB-27112: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (global refinement)

EMDB-27113: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement)

PDB-8d0z: 
S728-1157 IgG in complex with SARS-CoV-2-6P-Mut7 Spike protein (focused refinement)

EMDB-26200: 
Cryo-EM structure of SARS-CoV-2 spike in complex with FSR22, an anti-SARS-CoV-2 DARPin
EMDB-26201: 
Cryo-EM structure of SARS-CoV-2 spike in complex with FSR22, an anti-SARS-CoV-2 DARPin (Local refinement of FSR22 and RBD)
EMDB-27749: 
Cryo-EM structure of SARS-CoV-2 RBD in complex with anti-SARS-CoV-2 DARPin,SR22, and two antibody Fabs, S309 and CR3022
EMDB-27750: 
Cryo-EM structure of SARS-CoV-2 RBD in complex with anti-SARS-CoV-2 DARPin,SR16m, and two antibody Fabs, S309 and CR3022

PDB-7tyz: 
Cryo-EM structure of SARS-CoV-2 spike in complex with FSR22, an anti-SARS-CoV-2 DARPin

PDB-7tz0: 
Cryo-EM structure of SARS-CoV-2 spike in complex with FSR22, an anti-SARS-CoV-2 DARPin (Local refinement of FSR22 and RBD)

PDB-8dw2: 
Cryo-EM structure of SARS-CoV-2 RBD in complex with anti-SARS-CoV-2 DARPin,SR22, and two antibody Fabs, S309 and CR3022

PDB-8dw3: 
Cryo-EM structure of SARS-CoV-2 RBD in complex with anti-SARS-CoV-2 DARPin,SR16m, and two antibody Fabs, S309 and CR3022

EMDB-26928: 
Negative stain map of cH4/3
EMDB-26929: 
Negative stain half map of cH4/3 state 1

EMDB-26930: 
Negative stain map of cH4/3 state 3

EMDB-26931: 
Negative stain map of cH4/3 state 3

EMDB-26932: 
Negative stain map of cH4/3 state 4
EMDB-26933: 
Negative stain map of cH15/3

EMDB-26934: 
Negative stain map of cH15/3 state 1
EMDB-26935: 
Negative stain map of cH15/3 state 2

EMDB-26937: 
Negative stain map of cH15/3 state 3

EMDB-26938: 
Negative stain map of cH15/3 state 4
EMDB-26939: 
Negative stain map of cH15/3 in complex with CR9114 Fab

EMDB-24402: 
SARS-CoV-2 Spike in complex with PVI.V6-14 Fab

EMDB-24403: 
SARS-CoV-2 Spike in complex with PVI.V6-14 Fab

PDB-7rbu: 
SARS-CoV-2 Spike in complex with PVI.V6-14 Fab

PDB-7rbv: 
SARS-CoV-2 Spike in complex with PVI.V6-14 Fab

EMDB-23562: 
Negative stain EM map of 1E01 Fab in complex with N2 Singapore16

EMDB-25634: 
Negative stain map of monoclonal Fab 047-09 4F04 binding the anchor epitope of H1 HA

EMDB-25635: 
Negative stain map of monoclonal Fab 241 IgA 2F04 binding the anchor epitope of H1 HA

EMDB-25636: 
Negative stain map of polyclonal Fab 236.7 binding the anchor and esterase epitopes of H1 HA

EMDB-25637: 
Negative stain map of polyclonal Fab 236.7 binding the RBS epitope of H1 HA

EMDB-25638: 
Negative stain map of polyclonal Fab 236.14 binding an epitope on the top of the head of H1 HA
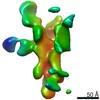
EMDB-25639: 
Negative stain map of polyclonal Fab 236.14 binding the esterase epitope of H1 HA

EMDB-25640: 
Negative stain map of polycolonal Fab 236.14 binding the RBS epitope of H1 HA

EMDB-25641: 
Negative stain map of polyclonal Fab 236.14 binding the anchor epitope of H1 HA
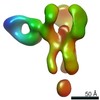
EMDB-25642: 
Negative stain map of polyclonal Fab 241.7 binding the esterase epitope of H1 HA
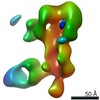
EMDB-25643: 
Negative stain map of polyclonal Fab 241.14 binding the anchor epitope of H1 HA
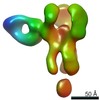
EMDB-25644: 
Negative stain map of polyclonal Fab 241.14 binding the esterase epitope of H1 HA

EMDB-25645: 
Negative stain map of polyclonal Fab 241.14 binding an epitope on the top of the head of H1 HA
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
