-Search query
-Search result
Showing 1 - 50 of 110 items for (author: eek & p)
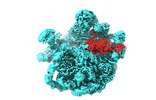
EMDB-37007: 
Mycobacterium smegmatis 50S ribosomal subunit-HflX complex
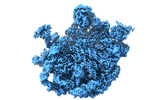
EMDB-38788: 
Mycobacterium smegmatis 50S ribosomal subunit with Erythromycin

PDB-8kab: 
Mycobacterium smegmatis 50S ribosomal subunit-HflX complex

PDB-8xz3: 
Mycobacterium smegmatis 50S ribosomal subunit with Erythromycin

EMDB-18438: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 1

EMDB-18439: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2

EMDB-18440: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 3

EMDB-18443: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 4

EMDB-18460: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1

EMDB-18461: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 2

PDB-8qrk: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 1

PDB-8qrl: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2

PDB-8qrm: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 3

PDB-8qrn: 
mt-SSU in GTPBP8 knock-out cells, state 4

PDB-8qu1: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1

PDB-8qu5: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 2

EMDB-42981: 
Prefusion-stabilized Respirovirus type 3 Fusion protein

PDB-8v5a: 
Prefusion-stabilized Respirovirus type 3 Fusion protein

EMDB-41839: 
Cryo-EM structure of yeast SWR1C subunit Swc5 bound to the nucleosome, 3D class 0

EMDB-41851: 
Cryo-EM structure of yeast SWR1C subunit Swc5 bound to the nucleosome, 3D class 1

EMDB-41852: 
Cryo-EM structure of yeast SWR1C subunit Swc5 bound to the nucleosome, 3D class 2

EMDB-41853: 
Cryo-EM structure of yeast SWR1C subunit Swc5 bound to the nucleosome, 3D class 4

EMDB-41423: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40

EMDB-41424: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1

EMDB-41425: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2

EMDB-41777: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:Pomalidomide:SD40

EMDB-41778: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 1

EMDB-41779: 
Map from local refinement (focused on CRBN) of DDB1dB:CRBN:PT-179:SD40, conformation 2

PDB-8tnp: 
Cryo-EM structure of DDB1dB:CRBN:Pomalidomide:SD40

PDB-8tnq: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 1

PDB-8tnr: 
Cryo-EM structure of DDB1dB:CRBN:PT-179:SD40, conformation 2

EMDB-16001: 
apo PrgB
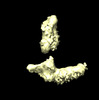
EMDB-16002: 
PrgB bound to DNA

EMDB-16730: 
Cryo-EM structure of complement C5 in complex with nanobodies UNbC5-1 and UNbC5-2

PDB-8cml: 
Cryo-EM structure of complement C5 in complex with nanobodies UNbC5-1 and UNbC5-2

EMDB-16433: 
Cryo-EM structure of NADH bound SLA dehydrogenase RlGabD from Rhizobium leguminosarum bv. trifolii SRD1565

PDB-8c54: 
Cryo-EM structure of NADH bound SLA dehydrogenase RlGabD from Rhizobium leguminosarum bv. trifolii SRD1565

EMDB-40789: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

EMDB-40790: 
Map focused on acidic patch BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

EMDB-40791: 
Overall map of BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

PDB-8svf: 
BAP1/ASXL1 bound to the H2AK119Ub Nucleosome

EMDB-27627: 
Structure of p110 gamma bound to the Ras inhibitory nanobody NB7

PDB-8dp0: 
Structure of p110 gamma bound to the Ras inhibitory nanobody NB7

EMDB-16145: 
Cryo-EM structure of NHEJ supercomplex(trimer)

PDB-8bot: 
Cryo-EM structure of NHEJ supercomplex(trimer)

EMDB-15022: 
CryoEM structure of Ku heterodimer bound to DNA, PAXX and XLF

EMDB-16044: 
DNA-PK Ku80 mediated dimer bound to PAXX

EMDB-16070: 
DNA-PK XLF mediated dimer bound to PAXX

EMDB-16074: 
DNA-PK Ku80 mediated dimer bound to PAXX and XLF

PDB-7zyg: 
CryoEM structure of Ku heterodimer bound to DNA, PAXX and XLF
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
