[English] 日本語
 Yorodumi
Yorodumi- EMDB-6666: The hexamer of full-length Cbln4 in complex with the LNS domain o... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6666 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
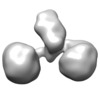
| Title | The hexamer of full-length Cbln4 in complex with the LNS domain of Nrxn1beta | |||||||||
 Map data Map data | ||||||||||
 Sample Sample | Nrxn1beta != Cbln4 Nrxn1beta
| |||||||||
| Biological species |   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  negative staining / Resolution: 20.0 Å negative staining / Resolution: 20.0 Å | |||||||||
 Authors Authors | Zhong C / Li G / Zhang H / Cao L / He Y / Ding J | |||||||||
 Citation Citation |  Journal: Cell Rep / Year: 2017 Journal: Cell Rep / Year: 2017Title: Cbln1 and Cbln4 Are Structurally Similar but Differ in GluD2 Binding Interactions. Authors: Chen Zhong / Jinlong Shen / Huibing Zhang / Guangyi Li / Senlin Shen / Fang Wang / Kuan Hu / Longxing Cao / Yongning He / Jianping Ding /  Abstract: Unlike cerebellin 1 (Cbln1), which bridges neurexin (Nrxn) receptors and δ-type glutamate receptors in a trans-synaptic triad, Cbln4 was reported to have no or weak binding for the receptors ...Unlike cerebellin 1 (Cbln1), which bridges neurexin (Nrxn) receptors and δ-type glutamate receptors in a trans-synaptic triad, Cbln4 was reported to have no or weak binding for the receptors despite sharing ∼70% sequence identity with Cbln1. Here, we report crystal structures of the homotrimers of the C1q domain of Cbln1 and Cbln4 at 2.2 and 2.3 Å resolution, respectively. Comparison of the structures suggests that the difference between Cbln1 and Cbln4 in GluD2 binding might be because of their sequence and structural divergence in loop CD. Surprisingly, we show that Cbln4 binds to Nrxn1β and forms a stable complex with the laminin, nectin, sex-hormone binding globulin (LNS) domain of Nrxn1β. Furthermore, the negative-stain electron microscopy reconstruction of hexameric full-length Cbln1 at 13 Å resolution and that of the Cbln4/Nrxn1β complex at 19 Å resolution suggest that Nrxn1β binds to the N-terminal region of Cbln4, probably through strand β10 of the S4 insert. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6666.map.gz emd_6666.map.gz | 284.3 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6666-v30.xml emd-6666-v30.xml emd-6666.xml emd-6666.xml | 9.6 KB 9.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_6666.png emd_6666.png | 39.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6666 http://ftp.pdbj.org/pub/emdb/structures/EMD-6666 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6666 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6666 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_6666.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6666.map.gz / Format: CCP4 / Size: 1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 3.48 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Nrxn1beta
| Entire | Name: Nrxn1beta |
|---|---|
| Components |
|
-Supramolecule #1: Cbln4
| Supramolecule | Name: Cbln4 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Recombinant expression | Organism:   Homo sapiens (human) / Recombinant plasmid: pCDH Homo sapiens (human) / Recombinant plasmid: pCDH |
-Supramolecule #2: Nrxn1beta
| Supramolecule | Name: Nrxn1beta / type: complex / ID: 2 / Parent: 0 / Macromolecule list: #2 |
|---|
-Macromolecule #1: Cbln4
| Macromolecule | Name: Cbln4 / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LQHHHHHHHH ASQNDTEPIV LEGKCLVVCD SNPATDSKGS SSSPLGISVR AANSKVAFSA VRSTNHEPSE MSNKTRIIYF DQILVNVGNF FTLESVFVAP RKGIYSFSFH VIKVYQSQTI QVNLMLNGKP VISAFAGDKD VTREAATNGV LLYLDKEDKV YLKLEKGNLL GGWQYSTFSG FLVFPL |
-Macromolecule #2: Nrxn1beta
| Macromolecule | Name: Nrxn1beta / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Rattus norvegicus (Norway rat) Rattus norvegicus (Norway rat) |
| Recombinant expression | Organism:   Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: mgshhhhhhh hgsdydiptt enlyfqgssl rgghagttyi fskgggqity kwppndrpst radrlaigfs tvqkeavlvr vdsssglgdy lelhihqgki gvkfnvgtdd iaieesnaii ndgkyhvvrf trsggnatlq vdswpviery pagnndnerl aiarqripyr ...String: mgshhhhhhh hgsdydiptt enlyfqgssl rgghagttyi fskgggqity kwppndrpst radrlaigfs tvqkeavlvr vdsssglgdy lelhihqgki gvkfnvgtdd iaieesnaii ndgkyhvvrf trsggnatlq vdswpviery pagnndnerl aiarqripyr lgrvvdewll dkgrqltifn sqatiiiggk eqgqpfqgql sglyynglkv lnmaaendan iaivgnvrlv gevpssefws hpqfek |
-Experimental details
-Structure determination
| Method |  negative staining negative staining |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Staining | Type: NEGATIVE / Material: 0.75% uranyl formate |
| Vitrification | Cryogen name: OTHER |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: FEI EAGLE (4k x 4k) / Average electron dose: 20.0 e/Å2 |
- Image processing
Image processing
| Initial angle assignment | Type: NOT APPLICABLE |
|---|---|
| Final angle assignment | Type: NOT APPLICABLE |
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 20.0 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 9148 |
 Movie
Movie Controller
Controller