[English] 日本語
 Yorodumi
Yorodumi- EMDB-3699: Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3699 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Rubisco from Rhodobacter sphaeroides in complex with CABP | |||||||||
 Map data Map data | postprocessed Rubisco map; D4 symmetry | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information ribulose-bisphosphate carboxylase / ribulose-bisphosphate carboxylase /  ribulose-bisphosphate carboxylase activity / reductive pentose-phosphate cycle / ribulose-bisphosphate carboxylase activity / reductive pentose-phosphate cycle /  monooxygenase activity / magnesium ion binding monooxygenase activity / magnesium ion bindingSimilarity search - Function | |||||||||
| Biological species |   Rhodobacter sphaeroides (bacteria) Rhodobacter sphaeroides (bacteria) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.39 Å cryo EM / Resolution: 3.39 Å | |||||||||
 Authors Authors | Bracher A / Milicic G / Ciniawsky S / Wendler P / Hayer-Hartl M / Hartl FU | |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2017 Journal: Mol Cell / Year: 2017Title: Mechanism of Enzyme Repair by the AAA Chaperone Rubisco Activase. Authors: Javaid Y Bhat / Goran Miličić / Gabriel Thieulin-Pardo / Andreas Bracher / Andrew Maxwell / Susanne Ciniawsky / Oliver Mueller-Cajar / John R Engen / F Ulrich Hartl / Petra Wendler / Manajit Hayer-Hartl /   Abstract: How AAA+ chaperones conformationally remodel specific target proteins in an ATP-dependent manner is not well understood. Here, we investigated the mechanism of the AAA+ protein Rubisco activase (Rca) ...How AAA+ chaperones conformationally remodel specific target proteins in an ATP-dependent manner is not well understood. Here, we investigated the mechanism of the AAA+ protein Rubisco activase (Rca) in metabolic repair of the photosynthetic enzyme Rubisco, a complex of eight large (RbcL) and eight small (RbcS) subunits containing eight catalytic sites. Rubisco is prone to inhibition by tight-binding sugar phosphates, whose removal is catalyzed by Rca. We engineered a stable Rca hexamer ring and analyzed its functional interaction with Rubisco. Hydrogen/deuterium exchange and chemical crosslinking showed that Rca structurally destabilizes elements of the Rubisco active site with remarkable selectivity. Cryo-electron microscopy revealed that Rca docks onto Rubisco over one active site at a time, positioning the C-terminal strand of RbcL, which stabilizes the catalytic center, for access to the Rca hexamer pore. The pulling force of Rca is fine-tuned to avoid global destabilization and allow for precise enzyme repair. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3699.map.gz emd_3699.map.gz | 10.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3699-v30.xml emd-3699-v30.xml emd-3699.xml emd-3699.xml | 17.4 KB 17.4 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3699.png emd_3699.png | 296.7 KB | ||
| Others |  emd_3699_additional.map.gz emd_3699_additional.map.gz | 94.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3699 http://ftp.pdbj.org/pub/emdb/structures/EMD-3699 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3699 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3699 | HTTPS FTP |
-Related structure data
| Related structure data |  5nv3MC  3700C  3701C  3702C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3699.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3699.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | postprocessed Rubisco map; D4 symmetry | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: unfiltered Rubisco map; D4 symmetry
| File | emd_3699_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unfiltered Rubisco map; D4 symmetry | ||||||||||||
| Projections & Slices |
| ||||||||||||
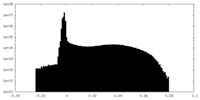
| Density Histograms |
- Sample components
Sample components
-Entire : Rubisco
| Entire | Name: Rubisco |
|---|---|
| Components |
|
-Supramolecule #1: Rubisco
| Supramolecule | Name: Rubisco / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 / Details: Rubisco was treated with the inhibitor CABP. |
|---|---|
| Source (natural) | Organism:   Rhodobacter sphaeroides (bacteria) Rhodobacter sphaeroides (bacteria) |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
-Macromolecule #1: Ribulose bisphosphate carboxylase large chain
| Macromolecule | Name: Ribulose bisphosphate carboxylase large chain / type: protein_or_peptide / ID: 1 / Number of copies: 8 / Enantiomer: LEVO / EC number:  ribulose-bisphosphate carboxylase ribulose-bisphosphate carboxylase |
|---|---|
| Source (natural) | Organism:   Rhodobacter sphaeroides (bacteria) Rhodobacter sphaeroides (bacteria) |
| Molecular weight | Theoretical: 51.768852 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: RYKAGVLKYA QMGYWDGDYV PKDTDVLALF RITPQEGVDP VEAAAAVAGE SSTATWTVVW TDRLTACDSY RAKAYRVEPV PGTPGQYFC YVAYDLILFE EGSIANLTAS IIGNVFSFKP LKAARLEDMR FPVAYVKTYK GPPTGIVGER ERLDKFGKPL L GATTKPKL ...String: RYKAGVLKYA QMGYWDGDYV PKDTDVLALF RITPQEGVDP VEAAAAVAGE SSTATWTVVW TDRLTACDSY RAKAYRVEPV PGTPGQYFC YVAYDLILFE EGSIANLTAS IIGNVFSFKP LKAARLEDMR FPVAYVKTYK GPPTGIVGER ERLDKFGKPL L GATTKPKL GLSGKNYGRV VYEGLKGGLD FM(KCX)DDENINS QPFMHWRDRF LYVMEAVNLA SAQTGEVKGH YLNITAGT M EEMYRRAEFA KSLGSVIVMV DLIIGYTAIQ SISEWCRQND MILHMHRAGH GTYTRQKNHG ISFRVIAKWL RLAGVDHLH CGTAVGKLEG DPLTVQGYYN VCREPFNTVD LPRGIFFEQD WADLRKVMPV ASGGIHAGQM HQLLSLFGDD VVLQFGGGTI GHPMGIQAG ATANRVALEA MVLARNEGRN IDVEGPEILR AAAKWCKPLE AALDTWGNIT FNYTSTDTSD FV |
-Macromolecule #2: Ribulose bisphosphate carboxylase small chain 1
| Macromolecule | Name: Ribulose bisphosphate carboxylase small chain 1 / type: protein_or_peptide / ID: 2 / Number of copies: 8 / Enantiomer: LEVO / EC number:  ribulose-bisphosphate carboxylase ribulose-bisphosphate carboxylase |
|---|---|
| Source (natural) | Organism:   Rhodobacter sphaeroides (bacteria) Rhodobacter sphaeroides (bacteria) |
| Molecular weight | Theoretical: 15.183234 KDa |
| Recombinant expression | Organism:   Escherichia coli BL21(DE3) (bacteria) Escherichia coli BL21(DE3) (bacteria) |
| Sequence | String: MRITQGCFSF LPDLTDEQIS AQVDYCLGRG WAVSLEHTDD PHPRNTYWEM WGMPMFDLRD PKGVMIELDE CRKAWPGRYI RINAFDSTR GFETVTMSFI VNRPEVEPSL RMERTEVDGR SIRYTHSIVR |
-Macromolecule #3: 2-CARBOXYARABINITOL-1,5-DIPHOSPHATE
| Macromolecule | Name: 2-CARBOXYARABINITOL-1,5-DIPHOSPHATE / type: ligand / ID: 3 / Number of copies: 8 / Formula: CAP |
|---|---|
| Molecular weight | Theoretical: 356.115 Da |
| Chemical component information |  ChemComp-CAP: |
-Macromolecule #4: MAGNESIUM ION
| Macromolecule | Name: MAGNESIUM ION / type: ligand / ID: 4 / Number of copies: 8 / Formula: MG |
|---|---|
| Molecular weight | Theoretical: 24.305 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.3 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV | ||||||||||||||||||
| Details | RcaCC hexamers (20 micromolar monomer) were mixed with E.C.M-CABP octamers (10 micromolar monomer) in a reaction containing 20 mM HEPES pH 7.5, 50 mM NaCl, 10 mM MgCl2, 10 mM ATP and 1mM RuBP, for 1 min at 25oC prior to addition of 0.125 % of glutaraldehyde (GA). After 10 min the reaction was quenched by addition of 0.1M Tris HCl pH 8 followed by gel filtration on a Superdex 200 PC 3.2/30 column (GE Healthcare).The fractions were eluted in buffer A and analyzed on a 6 % native gel. Fraction 13 containing HMW complexes with the least amount of free Rubisco were chosen for cryo-EM. The crosslinked E.C.M.-CABP-RcaCC complexes were diluted to 0.0030-0.0035 g ml-1 in 20 mM Tris-HCl pH 8.0, 50 mM NaCl, 1 mM ATP, 1 mM ATP-gammaS and 1 mM RuBP |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Details | Cs corrected Krios 1 at NeCEN (June 2016) |
| Image recording | Film or detector model: FEI FALCON II (4k x 4k) / Average exposure time: 1.25 sec. / Average electron dose: 50.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| CTF correction | Software - Name: CTFFIND (ver. 4) |
|---|---|
| Startup model | Type of model: PDB ENTRY PDB model - PDB ID: |
| Initial angle assignment | Type: OTHER |
| Final angle assignment | Type: OTHER / Software - Name: RELION |
| Final reconstruction | Applied symmetry - Point group: D4 (2x4 fold dihedral ) / Resolution.type: BY AUTHOR / Resolution: 3.39 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 333122 ) / Resolution.type: BY AUTHOR / Resolution: 3.39 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION / Number images used: 333122 |
-Atomic model buiding 1
| Refinement | Space: RECIPROCAL / Protocol: OTHER / Target criteria: Maximum likelihood |
|---|---|
| Output model |  PDB-5nv3: |
 Movie
Movie Controller
Controller












 Z
Z Y
Y X
X