+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3631 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM Structure of Foot and Mouth Disease Virus O1 Manisa | |||||||||
 Map data Map data | CryoEM Structure of Foot and Mouth Disease Virus O1 Manisa | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationicosahedral viral capsid / modulation by virus of host chromatin organization / RNA-protein covalent cross-linking / ribonucleoside triphosphate phosphatase activity / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / cytoplasmic vesicle membrane /  : / protein complex oligomerization / : / protein complex oligomerization /  regulation of translation ...icosahedral viral capsid / modulation by virus of host chromatin organization / RNA-protein covalent cross-linking / ribonucleoside triphosphate phosphatase activity / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / cytoplasmic vesicle membrane / regulation of translation ...icosahedral viral capsid / modulation by virus of host chromatin organization / RNA-protein covalent cross-linking / ribonucleoside triphosphate phosphatase activity / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / cytoplasmic vesicle membrane /  : / protein complex oligomerization / : / protein complex oligomerization /  regulation of translation / monoatomic ion channel activity / clathrin-dependent endocytosis of virus by host cell / host cell cytoplasm / regulation of translation / monoatomic ion channel activity / clathrin-dependent endocytosis of virus by host cell / host cell cytoplasm /  RNA helicase activity / RNA helicase activity /  viral protein processing / symbiont entry into host cell / viral RNA genome replication / cysteine-type endopeptidase activity / viral protein processing / symbiont entry into host cell / viral RNA genome replication / cysteine-type endopeptidase activity /  RNA-dependent RNA polymerase activity / DNA-templated transcription / structural molecule activity / virion attachment to host cell / RNA-dependent RNA polymerase activity / DNA-templated transcription / structural molecule activity / virion attachment to host cell /  proteolysis / proteolysis /  RNA binding / RNA binding /  ATP binding / ATP binding /  cytoplasm cytoplasmSimilarity search - Function | |||||||||
| Biological species |    Foot-and-mouth disease virus Foot-and-mouth disease virus | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.1 Å cryo EM / Resolution: 3.1 Å | |||||||||
 Authors Authors | Kotecha A / Stuart D | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Rules of engagement between αvβ6 integrin and foot-and-mouth disease virus. Authors: Abhay Kotecha / Quan Wang / Xianchi Dong / Serban L Ilca / Marina Ondiviela / Rao Zihe / Julian Seago / Bryan Charleston / Elizabeth E Fry / Nicola G A Abrescia / Timothy A Springer / Juha T ...Authors: Abhay Kotecha / Quan Wang / Xianchi Dong / Serban L Ilca / Marina Ondiviela / Rao Zihe / Julian Seago / Bryan Charleston / Elizabeth E Fry / Nicola G A Abrescia / Timothy A Springer / Juha T Huiskonen / David I Stuart /     Abstract: Foot-and-mouth disease virus (FMDV) mediates cell entry by attachment to an integrin receptor, generally αvβ6, via a conserved arginine-glycine-aspartic acid (RGD) motif in the exposed, antigenic, ...Foot-and-mouth disease virus (FMDV) mediates cell entry by attachment to an integrin receptor, generally αvβ6, via a conserved arginine-glycine-aspartic acid (RGD) motif in the exposed, antigenic, GH loop of capsid protein VP1. Infection can also occur in tissue culture adapted virus in the absence of integrin via acquired basic mutations interacting with heparin sulphate (HS); this virus is attenuated in natural infections. HS interaction has been visualized at a conserved site in two serotypes suggesting a propensity for sulfated-sugar binding. Here we determined the interaction between αvβ6 and two tissue culture adapted FMDV strains by cryo-electron microscopy. In the preferred mode of engagement, the fully open form of the integrin, hitherto unseen at high resolution, attaches to an extended GH loop via interactions with the RGD motif plus downstream hydrophobic residues. In addition, an N-linked sugar of the integrin attaches to the previously identified HS binding site, suggesting a functional role. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3631.map.gz emd_3631.map.gz | 229 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3631-v30.xml emd-3631-v30.xml emd-3631.xml emd-3631.xml | 18.7 KB 18.7 KB | Display Display |  EMDB header EMDB header |
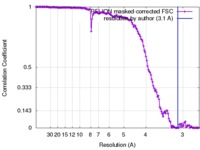
| FSC (resolution estimation) |  emd_3631_fsc.xml emd_3631_fsc.xml | 14 KB | Display |  FSC data file FSC data file |
| Images |  emd_3631.png emd_3631.png | 295.3 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3631 http://ftp.pdbj.org/pub/emdb/structures/EMD-3631 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3631 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3631 | HTTPS FTP |
-Related structure data
| Related structure data |  5nejMC  3630C  3632C  3633C  3634C  3635C  5ne4C  5nedC  5nemC  5nerC  5netC  5neuC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3631.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3631.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM Structure of Foot and Mouth Disease Virus O1 Manisa | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.35 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Foot-and-mouth disease virus
| Entire | Name:    Foot-and-mouth disease virus Foot-and-mouth disease virus |
|---|---|
| Components |
|
-Supramolecule #1: Foot-and-mouth disease virus
| Supramolecule | Name: Foot-and-mouth disease virus / type: virus / ID: 1 / Parent: 0 / Macromolecule list: all / NCBI-ID: 12110 / Sci species name: Foot-and-mouth disease virus / Sci species strain: O1 Manisa / Virus type: VIRION / Virus isolate: SEROTYPE / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:   Bos taurus (cattle) Bos taurus (cattle) |
| Host system | Organism:  Cricetinae gen. sp. (mammal) / Recombinant cell: BHK-21 Cricetinae gen. sp. (mammal) / Recombinant cell: BHK-21 |
| Molecular weight | Theoretical: 9 MDa |
| Virus shell | Shell ID: 1 / Name: Foot and Mouth Disease virus / Diameter: 300.0 Å / T number (triangulation number): 3 |
-Macromolecule #1: O1 Manisa VP1
| Macromolecule | Name: O1 Manisa VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:    Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 23.222348 KDa |
| Recombinant expression | Organism:  Cricetinae gen. sp. (mammal) Cricetinae gen. sp. (mammal) |
| Sequence | String: TTSAGESADP VTATVENYGG ETQVQRRQHT DVSFILDRFV KVTPKDQINV LDLMQTPAHT LVGALLRTAT YYFADLEVAV KHEGNLTWV PNGAPEAALD NTTNPTAYHK APLTRLALPY TAPHRVLATV YNGESKYGDG TVANVRGDLQ VLAQKAARAL P TSFNYGAI ...String: TTSAGESADP VTATVENYGG ETQVQRRQHT DVSFILDRFV KVTPKDQINV LDLMQTPAHT LVGALLRTAT YYFADLEVAV KHEGNLTWV PNGAPEAALD NTTNPTAYHK APLTRLALPY TAPHRVLATV YNGESKYGDG TVANVRGDLQ VLAQKAARAL P TSFNYGAI KATRVTELLY RMKRAETYCP RPLLAIHPDQ ARHKQKIVAP VKQ |
-Macromolecule #2: O1 Manisa VP2
| Macromolecule | Name: O1 Manisa VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:    Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 24.41751 KDa |
| Recombinant expression | Organism:  Cricetinae gen. sp. (mammal) Cricetinae gen. sp. (mammal) |
| Sequence | String: DKKTEETTLL EDRILTTRNG HTTSTTQSSV GVTYGYATAE DFVSGPNTSG LETRVAQAER FFKTHLFDWV TSDPFGRCHL LELPTDHKG VYGYLTDSYA YMRNGWDVEV TAVGNQFNGG CLLVAMVPEL CSIQKRELYQ LTLFPHQFIN PRTNMTAHIT V PFVGVNRY ...String: DKKTEETTLL EDRILTTRNG HTTSTTQSSV GVTYGYATAE DFVSGPNTSG LETRVAQAER FFKTHLFDWV TSDPFGRCHL LELPTDHKG VYGYLTDSYA YMRNGWDVEV TAVGNQFNGG CLLVAMVPEL CSIQKRELYQ LTLFPHQFIN PRTNMTAHIT V PFVGVNRY DQYKVHKPWT LVVMVVAPLT VNSEGAPQIK VYANIAPTNV HVAGEFPSKE |
-Macromolecule #3: O1 Manisa VP3
| Macromolecule | Name: O1 Manisa VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:    Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 23.933879 KDa |
| Recombinant expression | Organism:  Cricetinae gen. sp. (mammal) Cricetinae gen. sp. (mammal) |
| Sequence | String: GIFPVACSDG YGGLVTTDPK TADPAYGKVF NPPRNMLPGR FTNFLDVAEA CPTFLHFEGD VPYVTTKTDS DRVLAQFDLS LAAKHMSNT FLAGLAQYYT QYSGTINLHF MFTGPTDAKA RYMIAYAPPG MEPPKTPEAA AHCIHAEWDT GLNSKFTFSI P YLSAADYT ...String: GIFPVACSDG YGGLVTTDPK TADPAYGKVF NPPRNMLPGR FTNFLDVAEA CPTFLHFEGD VPYVTTKTDS DRVLAQFDLS LAAKHMSNT FLAGLAQYYT QYSGTINLHF MFTGPTDAKA RYMIAYAPPG MEPPKTPEAA AHCIHAEWDT GLNSKFTFSI P YLSAADYT YTASDVAETT NVQGWVCLFQ ITHGKADGDA LVVLASAGKD FELRLPVDAR TQ |
-Macromolecule #4: O1 Manisa VP1
| Macromolecule | Name: O1 Manisa VP1 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:    Foot-and-mouth disease virus Foot-and-mouth disease virus |
| Molecular weight | Theoretical: 8.766075 KDa |
| Recombinant expression | Organism:  Cricetinae gen. sp. (mammal) Cricetinae gen. sp. (mammal) |
| Sequence | String: GAGQSSPATG SQNQSGNTGS IINNYYMQQY QNSMDTQLGD NATSGGSNEG STDTTSTHTT NTQNNDWFSK LASSAFSGLF GALLA |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.5 mg/mL |
|---|---|
| Buffer | pH: 8 / Component - Concentration: 50.0 mM / Component - Name: HEPES |
| Grid | Model: C-flat-2/1 / Material: COPPER / Mesh: 300 / Support film - Material: CARBON / Support film - topology: HOLEY / Support film - Film thickness: 5.0 nm / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 90 % / Chamber temperature: 294 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 37037 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 160000 Bright-field microscopy / Cs: 2.0 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 1.0 µm / Nominal magnification: 160000 |
| Specialist optics | Energy filter - Name: GIF / Energy filter - Lower energy threshold: 0 eV / Energy filter - Upper energy threshold: 20 eV |
| Sample stage | Specimen holder model: GATAN 910 MULTI-SPECIMEN SINGLE TILT CRYO TRANSFER HOLDER Cooling holder cryogen: NITROGEN |
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Frames/image: 2-20 / Number grids imaged: 1 / Number real images: 360 / Average exposure time: 5.0 sec. / Average electron dose: 18.0 e/Å2 |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller