[English] 日本語
 Yorodumi
Yorodumi- EMDB-4145: Subtomogram average of the ER membrane-associated ribosome from F... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-4145 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
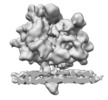
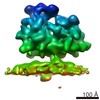
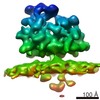
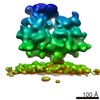
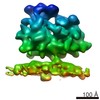
| Title | Subtomogram average of the ER membrane-associated ribosome from FIB-milled C. reinhardtii cells | |||||||||
 Map data Map data | Subtomogram average of the ER membrane-associated ribosome from FIB-milled C. reinhardtii cells | |||||||||
 Sample Sample |
| |||||||||
| Biological species |   Chlamydomonas reinhardtii (plant) Chlamydomonas reinhardtii (plant) | |||||||||
| Method | subtomogram averaging /  cryo EM / Resolution: 19.4 Å cryo EM / Resolution: 19.4 Å | |||||||||
 Authors Authors | Pfeffer S / Schaffer M / Albert S / Engel BD / Foerster F | |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2017 Journal: Nat Commun / Year: 2017Title: Dissecting the molecular organization of the translocon-associated protein complex. Authors: Stefan Pfeffer / Johanna Dudek / Miroslava Schaffer / Bobby G Ng / Sahradha Albert / Jürgen M Plitzko / Wolfgang Baumeister / Richard Zimmermann / Hudson H Freeze / Benjamin D Engel / Friedrich Förster /    Abstract: In eukaryotic cells, one-third of all proteins must be transported across or inserted into the endoplasmic reticulum (ER) membrane by the ER protein translocon. The translocon-associated protein ...In eukaryotic cells, one-third of all proteins must be transported across or inserted into the endoplasmic reticulum (ER) membrane by the ER protein translocon. The translocon-associated protein (TRAP) complex is an integral component of the translocon, assisting the Sec61 protein-conducting channel by regulating signal sequence and transmembrane helix insertion in a substrate-dependent manner. Here we use cryo-electron tomography (CET) to study the structure of the native translocon in evolutionarily divergent organisms and disease-linked TRAP mutant fibroblasts from human patients. The structural differences detected by subtomogram analysis form a basis for dissecting the molecular organization of the TRAP complex. We assign positions to the four TRAP subunits within the complex, providing insights into their individual functions. The revealed molecular architecture of a central translocon component advances our understanding of membrane protein biogenesis and sheds light on the role of TRAP in human congenital disorders of glycosylation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_4145.map.gz emd_4145.map.gz | 25.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-4145-v30.xml emd-4145-v30.xml emd-4145.xml emd-4145.xml | 8 KB 8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_4145.png emd_4145.png | 67.4 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-4145 http://ftp.pdbj.org/pub/emdb/structures/EMD-4145 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4145 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-4145 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_4145.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_4145.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Subtomogram average of the ER membrane-associated ribosome from FIB-milled C. reinhardtii cells | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 3.42 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ER membrane-associated ribosome from FIB-milled C. reinhardtii cells
| Entire | Name: ER membrane-associated ribosome from FIB-milled C. reinhardtii cells |
|---|---|
| Components |
|
-Supramolecule #1: ER membrane-associated ribosome from FIB-milled C. reinhardtii cells
| Supramolecule | Name: ER membrane-associated ribosome from FIB-milled C. reinhardtii cells type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:   Chlamydomonas reinhardtii (plant) Chlamydomonas reinhardtii (plant) |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | cell |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE-PROPANE |
| Details | Whole cells were vitrified by plunge-freezing and thinned by focused ion beam milling. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD Bright-field microscopy Bright-field microscopy |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 1.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Extraction | Number tomograms: 65 / Number images used: 8070 |
|---|---|
| Final angle assignment | Type: OTHER |
| Final reconstruction | Applied symmetry - Point group: C1 (asymmetric) / Resolution.type: BY AUTHOR / Resolution: 19.4 Å / Resolution method: FSC 0.5 CUT-OFF / Software - Name: PyTom / Number subtomograms used: 8070 |
 Movie
Movie Controller
Controller