[English] 日本語
 Yorodumi
Yorodumi- EMDB-3908: PolyA polymerase module of the cleavage and polyadenylation facto... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3908 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | PolyA polymerase module of the cleavage and polyadenylation factor (CPF) from Saccharomyces cerevisiae | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information: / : / Processing of Intronless Pre-mRNAs / termination of RNA polymerase II transcription, poly(A)-coupled / mRNA cleavage and polyadenylation specificity factor complex / termination of RNA polymerase II transcription / : /  mitochondrion / mitochondrion /  RNA binding / RNA binding /  metal ion binding ...: / : / Processing of Intronless Pre-mRNAs / termination of RNA polymerase II transcription, poly(A)-coupled / mRNA cleavage and polyadenylation specificity factor complex / termination of RNA polymerase II transcription / : / metal ion binding ...: / : / Processing of Intronless Pre-mRNAs / termination of RNA polymerase II transcription, poly(A)-coupled / mRNA cleavage and polyadenylation specificity factor complex / termination of RNA polymerase II transcription / : /  mitochondrion / mitochondrion /  RNA binding / RNA binding /  metal ion binding / metal ion binding /  nucleus / nucleus /  cytosol cytosolSimilarity search - Function | |||||||||
| Biological species |   SaccharSaccharomyces cerevisiae (strain ATCC 204508 / S288c)omyces (yeast) / SaccharSaccharomyces cerevisiae (strain ATCC 204508 / S288c)omyces (yeast) /   Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) / Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) /   Saccharomyces cerevisiae S288c (yeast) Saccharomyces cerevisiae S288c (yeast) | |||||||||
| Method |  single particle reconstruction / single particle reconstruction /  cryo EM / Resolution: 3.55 Å cryo EM / Resolution: 3.55 Å | |||||||||
 Authors Authors | Casanal A / Kumar A / Hill CH / Emsley P / Passmore L | |||||||||
 Citation Citation |  Journal: Science / Year: 2017 Journal: Science / Year: 2017Title: Architecture of eukaryotic mRNA 3'-end processing machinery. Authors: Ana Casañal / Ananthanarayanan Kumar / Chris H Hill / Ashley D Easter / Paul Emsley / Gianluca Degliesposti / Yuliya Gordiyenko / Balaji Santhanam / Jana Wolf / Katrin Wiederhold / Gillian ...Authors: Ana Casañal / Ananthanarayanan Kumar / Chris H Hill / Ashley D Easter / Paul Emsley / Gianluca Degliesposti / Yuliya Gordiyenko / Balaji Santhanam / Jana Wolf / Katrin Wiederhold / Gillian L Dornan / Mark Skehel / Carol V Robinson / Lori A Passmore /  Abstract: Newly transcribed eukaryotic precursor messenger RNAs (pre-mRNAs) are processed at their 3' ends by the ~1-megadalton multiprotein cleavage and polyadenylation factor (CPF). CPF cleaves pre-mRNAs, ...Newly transcribed eukaryotic precursor messenger RNAs (pre-mRNAs) are processed at their 3' ends by the ~1-megadalton multiprotein cleavage and polyadenylation factor (CPF). CPF cleaves pre-mRNAs, adds a polyadenylate tail, and triggers transcription termination, but it is unclear how its various enzymes are coordinated and assembled. Here, we show that the nuclease, polymerase, and phosphatase activities of yeast CPF are organized into three modules. Using electron cryomicroscopy, we determined a 3.5-angstrom-resolution structure of the ~200-kilodalton polymerase module. This revealed four β propellers, in an assembly markedly similar to those of other protein complexes that bind nucleic acid. Combined with in vitro reconstitution experiments, our data show that the polymerase module brings together factors required for specific and efficient polyadenylation, to help coordinate mRNA 3'-end processing. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3908.map.gz emd_3908.map.gz | 14.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3908-v30.xml emd-3908-v30.xml emd-3908.xml emd-3908.xml | 22.9 KB 22.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_3908.png emd_3908.png | 143.2 KB | ||
| Masks |  emd_3908_msk_1.map emd_3908_msk_1.map | 15.6 MB |  Mask map Mask map | |
| Others |  emd_3908_half_map_1.map.gz emd_3908_half_map_1.map.gz emd_3908_half_map_2.map.gz emd_3908_half_map_2.map.gz | 11.9 MB 11.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3908 http://ftp.pdbj.org/pub/emdb/structures/EMD-3908 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3908 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3908 | HTTPS FTP |
-Related structure data
| Related structure data |  6eojMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10299 (Title: PolyA polymerase module of the cleavage and polyadenylation factor (CPF) from Saccharomyces cerevisiae EMPIAR-10299 (Title: PolyA polymerase module of the cleavage and polyadenylation factor (CPF) from Saccharomyces cerevisiaeData size: 15.4 TB Data #1: Polished shiny particles [picked particles - multiframe - processed] Data #2: Unaligned super-resolution movies [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3908.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3908.map.gz / Format: CCP4 / Size: 15.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 1.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
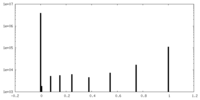
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_3908_msk_1.map emd_3908_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
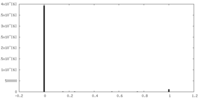
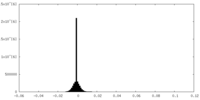
| Density Histograms |
-Half map: #1
| File | emd_3908_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
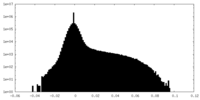
| Density Histograms |
-Half map: #2
| File | emd_3908_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
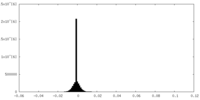
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of Cft1, Yth1, Pfs2 and Fip1
| Entire | Name: Complex of Cft1, Yth1, Pfs2 and Fip1 |
|---|---|
| Components |
|
-Supramolecule #1: Complex of Cft1, Yth1, Pfs2 and Fip1
| Supramolecule | Name: Complex of Cft1, Yth1, Pfs2 and Fip1 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:   SaccharSaccharomyces cerevisiae (strain ATCC 204508 / S288c)omyces (yeast) SaccharSaccharomyces cerevisiae (strain ATCC 204508 / S288c)omyces (yeast) |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Molecular weight | Theoretical: 270 KDa |
-Macromolecule #1: Protein CFT1
| Macromolecule | Name: Protein CFT1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast)Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 153.577156 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MNVYDDVLDA TVVSHSLATH FTTSDYEELL VVRTNILSVY RPTRDGKLYL TDEFKFHGLI TDIGLIPQKD SPLSCLLLCT GVAKISILK FNTLTNSIDT LSLHYYEGKF KGKSLVELAK ISTLRMDPGS SCALLFNNDI IAFLPFHVNK NDDDEEEEDE D ENIDDSEL ...String: MNVYDDVLDA TVVSHSLATH FTTSDYEELL VVRTNILSVY RPTRDGKLYL TDEFKFHGLI TDIGLIPQKD SPLSCLLLCT GVAKISILK FNTLTNSIDT LSLHYYEGKF KGKSLVELAK ISTLRMDPGS SCALLFNNDI IAFLPFHVNK NDDDEEEEDE D ENIDDSEL IHSMNQKSQG TNTFNKRKRT KLGDKFTAPS VVLVASELYE GAKNIIDIQF LKNFTKPTIA LLYQPKLVWA GN TTISKLP TQYVILTLNI QPAESATKIE STTIAFVKEL PWDLHTIVPV SNGAIIVGTN ELAFLDNTGV LQSTVLLNSF ADK ELQKTK IINNSSLEIM FREKNTTSIW IPSSKSKNGG SNNDETLLLM DLKSNIYYIQ MEAEGRLLIK FDIFKLPIVN DLLK ENSNP KCITRLNATN SNKNMDLFIG FGSGNALVLR LNNLKSTIET REAHNPSSGT NSLMDINDDD DEEMDDLYAD EAPEN GLTT NDSKGTVETV QPFDIELLSS LRNVGPITSL TVGKVSSIDD VVKGLPNPNK NEYSLVATSG NGSGSHLTVI QTSVQP EIE LALKFISITQ IWNLKIKGRD RYLITTDSTK SRSDIYESDN NFKLHKGGRL RRDATTVYIS MFGEEKRIIQ VTTNHLY LY DTHFRRLTTI KFDYEVIHVS VMDPYILVTV SRGDIKIFEL EEKNKRKLLK VDLPEILNEM VITSGLILKS NMCNEFLI G LSKSQEEQLL FTFVTADNQI IFFTKDHNDR IFQLNGVDQL NESLYISTYQ LGDEIVPDPS IKQVMINKLG HDNKEEYLT ILTFGGEIYQ YRKLPQRRSR FYRNVTRNDL AITGAPDNAY AKGVSSIERI MHYFPDYNGY SVIFVTGSVP YILIKEDDST PKIFKFGNI PLVSVTPWSE RSVMCVDDIK NARVYTLTTD NMYYGNKLPL KQIKISNVLD DYKTLQKLVY HERAQLFLVS Y CKRVPYEA LGEDGEKVIG YDENVPHAEG FQSGILLINP KSWKVIDKID FPKNSVVNEM RSSMIQINSK TKRKREYIIA GV ANATTED TPPTGAFHIY DVIEVVPEPG KPDTNYKLKE IFQEEVSGTV STVCEVSGRF MISQSQKVLV RDIQEDNSVI PVA FLDIPV FVTDSKSFGN LLIIGDAMQG FQFIGFDAEP YRMISLGRSM SKFQTMSLEF LVNGGDMYFA ATDADRNVHV LKYA PDEPN SLSGQRLVHC SSFTLHSTNS CMMLLPRNEE FGSPQVPSFQ NVGGQVDGSV FKIVPLSEEK YRRLYVIQQQ IIDRE LQLG GLNPRMERLA NDFYQMGHSM RPMLDFNVIR RFCGLAIDRR KSIAQKAGRH AHFEAWRDII NIEFSMRSLC QGK |
-Macromolecule #2: mRNA 3'-end-processing protein YTH1
| Macromolecule | Name: mRNA 3'-end-processing protein YTH1 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast) Saccharomyces cerevisiae (strain ATCC 204508 / S288c) (yeast)Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 24.560416 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: PSLIHPDTAK YPFKFEPFLR QEYSFSLDPD RPICEFYNSR EGPKSCPRGP LCPKKHVLPI FQNKIVCRHW LRGLCKKNDQ CEYLHEYNL RKMPECVFFS KNGYCTQSPD CQYLHIDPAS KIPKCENYEM GFCPLGSSCP RRHIKKVFCQ RYMTGFCPLG K DECDMEHP ...String: PSLIHPDTAK YPFKFEPFLR QEYSFSLDPD RPICEFYNSR EGPKSCPRGP LCPKKHVLPI FQNKIVCRHW LRGLCKKNDQ CEYLHEYNL RKMPECVFFS KNGYCTQSPD CQYLHIDPAS KIPKCENYEM GFCPLGSSCP RRHIKKVFCQ RYMTGFCPLG K DECDMEHP QFIIPDEGSK LRIKRDDEIN TRKMDEEKER RLNAIINGEV |
-Macromolecule #3: Polyadenylation factor subunit 2,Polyadenylation factor subunit 2
| Macromolecule | Name: Polyadenylation factor subunit 2,Polyadenylation factor subunit 2 type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Saccharomyces cerevisiae S288c (yeast) / Strain: ATCC 204508 / S288c Saccharomyces cerevisiae S288c (yeast) / Strain: ATCC 204508 / S288c |
| Molecular weight | Theoretical: 53.636645 KDa |
| Recombinant expression | Organism:   Spodoptera frugiperda (fall armyworm) Spodoptera frugiperda (fall armyworm) |
| Sequence | String: MDGHNQNQYQ NQNQIQQSQQ PPLKKYVTQR RSVDVSSPYI NLYYNRRHGL PNLVVEPETS YTIDIMPPNA YRGRDRVINL PSKFTHLSS NKVKHVIPAI QWTPEGRRLV VATYSGEFSL WNASSFTFET LMQAHDSAVT TMKYSHDSDW MISGDADGMI K IWQPNFSM ...String: MDGHNQNQYQ NQNQIQQSQQ PPLKKYVTQR RSVDVSSPYI NLYYNRRHGL PNLVVEPETS YTIDIMPPNA YRGRDRVINL PSKFTHLSS NKVKHVIPAI QWTPEGRRLV VATYSGEFSL WNASSFTFET LMQAHDSAVT TMKYSHDSDW MISGDADGMI K IWQPNFSM VKEIDAAHTE SIRDMAFSSN DSKFVTCSDD NILKIWNFSN GKQERVLSGH HWDVKSCDWH PEMGLIASAS KD NLVKLWD PRSGNCISSI LKFKHTVLKT RFQPTKGNLL MAISKDKSCR VFDIRYSMKE LMCVRDETDY MTLEWHPINE SMF TLACYD GSLKHFDLLQ NLNEPILTIP YAHDKCITSL SYNPVGHIFA TAAKDRTIRF WTRARPIDPN AYDDPTYNNK KING WFFGI NNDINAVREK SEFGAAPPPP ATLEPHALPN MNGFINKKPR QEIPGIDSNI KSSTLPGLSI (UNK)(UNK)(UNK) (UNK)(UNK) |
-Macromolecule #4: ZINC ION
| Macromolecule | Name: ZINC ION / type: ligand / ID: 4 / Number of copies: 2 / Formula: ZN |
|---|---|
| Molecular weight | Theoretical: 65.409 Da |
-Experimental details
-Structure determination
| Method |  cryo EM cryo EM |
|---|---|
 Processing Processing |  single particle reconstruction single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.1 mg/mL | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.9 Component:
| ||||||||
| Grid | Model: Quantifoil UltrAuFoil / Material: GOLD / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Atmosphere: AIR | ||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: OTHER / Imaging mode: BRIGHT FIELD Bright-field microscopy / Cs: 2.7 mm / Nominal magnification: 81000 Bright-field microscopy / Cs: 2.7 mm / Nominal magnification: 81000 |
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 5 / Number real images: 4227 / Average exposure time: 16.0 sec. / Average electron dose: 45.0 e/Å2 |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Particle selection | Number selected: 460167 |
|---|---|
| CTF correction | Software - Name: RELION (ver. 2.0) / Software - details: GCTF |
| Startup model | Type of model: OTHER Details: 3D model generated from a previous data set collected from a CPF native sample in a Titan Krios equipped with a Falcon II detector. |
| Initial angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION (ver. 2.0) |
| Final 3D classification | Number classes: 10 / Software - Name: RELION (ver. 2.0) |
| Final angle assignment | Type: ANGULAR RECONSTITUTION / Software - Name: RELION (ver. 2.0) |
| Final reconstruction | Number classes used: 1 / Applied symmetry - Point group: C1 (asymmetric) / Algorithm: FOURIER SPACE / Resolution.type: BY AUTHOR / Resolution: 3.55 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: RELION (ver. 2.0) / Number images used: 77197 |
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL |
|---|---|
| Output model |  PDB-6eoj: |
 Movie
Movie Controller
Controller














 Z
Z Y
Y X
X