|
|
eF-site ID
|
1t1e-A |
|
PDB Code
|
1t1e |
|
Chain
|
A |
|
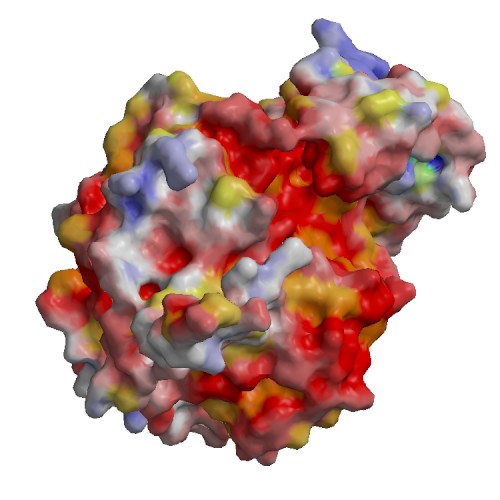
click to enlarge
|
|
|
Title
|
High Resolution Crystal Structure of the Intact Pro-Kumamolisin, a Sedolisin Type Proteinase (previously called Kumamolysin or KSCP) |
|
Classification
|
HYDROLASE |
|
Compound
|
kumamolisin |
|
Source
|
Bacillus sp. MN-32 (Q8RR56_9BACI) |
|
|
Sequence
|
A: |
EKREVLAGHARRQAPQAVDKGPVTGDQRISVTVVLRRQRG
DELEAHVERQAALAPHARVHLEREAFAASHGASLDDFAEI
RKFAEAHGLTLDRAHVAAGTAVLSGPVDAVNQAFGVELRH
FDHPDGSYRSYVGDVRVPASIAPLIEAVLGLDTRPVARPH
FRLRRRAEGEFEARSQSAAPTAYTPLDVAQAYQFPEGLDG
QGQCIAIIELGGGYDETSLAQYFASLGVSAPQVVSVSVDG
ATNQPTGDPNGPDGEVELDIEVAGALAPGAKIAVYFAPNT
DAGFLNAITTAVHDPTHKPSIVSISWGGPEDSWAPASIAA
MNRAFLDAAALGVTVLAAAGDSGSTDGEQDGLYHVDFPAA
SPYVLACGGTRLVASAGRIERETVWNDGPDGGSTGGGVSR
IFPLPSWQERANVPPSANPGAGSGRGVPDVAGNADPATGY
EVVIDGETTVIGGTAAVAPLFAALVARINQKLGKPVGYLN
PTLYQLPPEVFHDITEGNNDIANRARIYQAGPGWDPCTGL
GSPIGIRLLQALLP
|
|
|
Description
|
|
Functional site
|
|
|
1)
|
chain |
A |
| residue |
505 |
| type |
|
| sequence |
D
|
| description |
BINDING SITE FOR RESIDUE CA A 700
|
| source |
: AC1
|
|
|
2)
|
chain |
A |
| residue |
506 |
| type |
|
| sequence |
I
|
| description |
BINDING SITE FOR RESIDUE CA A 700
|
| source |
: AC1
|
|
|
3)
|
chain |
A |
| residue |
523 |
| type |
|
| sequence |
G
|
| description |
BINDING SITE FOR RESIDUE CA A 700
|
| source |
: AC1
|
|
|
4)
|
chain |
A |
| residue |
525 |
| type |
|
| sequence |
G
|
| description |
BINDING SITE FOR RESIDUE CA A 700
|
| source |
: AC1
|
|
|
5)
|
chain |
A |
| residue |
527 |
| type |
|
| sequence |
D
|
| description |
BINDING SITE FOR RESIDUE CA A 700
|
| source |
: AC1
|
|
|
|
