8BD0
 
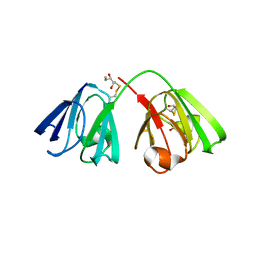 | |
4AOK
 
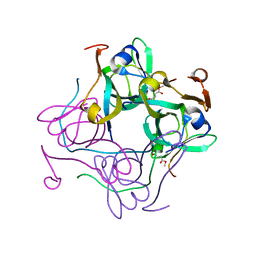 | |
4AON
 
 | |
4C3C
 
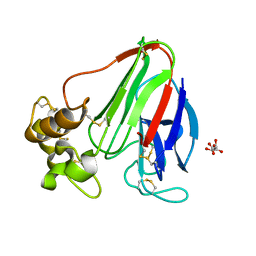 | | Thaumatin refined against hatrx data for time-point 1 | | Descriptor: | L(+)-TARTARIC ACID, THAUMATIN-1 | | Authors: | Yorke, B.A, Beddard, G.S, Owen, R.L, Pearson, A.R. | | Deposit date: | 2013-08-22 | | Release date: | 2014-09-03 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Time-Resolved Crystallography Using the Hadamard Transform.
Nat.Methods, 11, 2014
|
|
7A8Y
 
 | |
4AZD
 
 | | T57V mutant of aspartate decarboxylase | | Descriptor: | ASPARTATE 1-DECARBOXYLASE, MALONATE ION | | Authors: | Webb, M.E, Yorke, B.A, Kershaw, T, Lovelock, S, Lobley, C.M.C, Kilkenny, M.L, Smith, A.G, Blundell, T.L, Pearson, A.R, Abell, C. | | Deposit date: | 2012-06-25 | | Release date: | 2012-07-25 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Threonine 57 is Required for the Post-Translational Activation of Escherichia Coli Aspartate Alpha-Decarboxylase
Acta Crystallogr.,Sect.D, 70, 2014
|
|
8BPI
 
 | |
7VN0
 
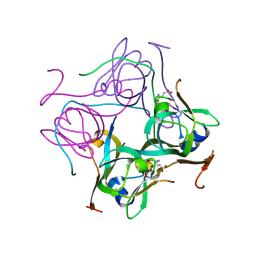
 | | CATPO mutant - T188A | | Descriptor: | ALANINE, CALCIUM ION, CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, ... | | Authors: | Yuzugullu Karakus, Y, Balci Unver, S, Zengin Karatas, M, Goc, G, Pearson, A.R, Yorke, B. | | Deposit date: | 2021-10-10 | | Release date: | 2022-09-14 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Investigation of how gate residues in the main channel affect the catalytic activity of Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
7WCA
 
 | | CATPO mutant - E484A | | Descriptor: | CALCIUM ION, Catalase, PENTAETHYLENE GLYCOL, ... | | Authors: | Yuzugullu Karakus, Y, Balci Unver, S, Zengin Karatas, M, Goc, G, Pearson, A.R, Yorke, B. | | Deposit date: | 2021-12-18 | | Release date: | 2022-12-21 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.78 Å) | | Cite: | Investigation of how gate residues in the main channel affect the catalytic activity of Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
5XVZ
 
 | | CATPO mutant - H246W | | Descriptor: | CALCIUM ION, CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, Catalase, ... | | Authors: | Yuzugullu Karakus, Y, Goc, G, Balci, S, Pearson, A.R, Yorke, B. | | Deposit date: | 2017-06-28 | | Release date: | 2018-07-04 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Identification of the site of oxidase substrate binding in Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
5Y17
 
 | | CATPO mutant - E316F | | Descriptor: | CALCIUM ION, CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, Catalase | | Authors: | Karakus Yuzugullu, Y, Goc, G, Balci, S, Pearson, A.R, Yorke, B. | | Deposit date: | 2017-07-19 | | Release date: | 2018-07-25 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Identification of the site of oxidase substrate binding in Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
5YEM
 
 | | CATPO mutant - T188F | | Descriptor: | CALCIUM ION, Catalase, PROTOPORPHYRIN IX CONTAINING FE | | Authors: | Yuzugullu Karakus, Y, Balci, S, Goc, G, Pearson, A.R, Yorke, B. | | Deposit date: | 2017-09-18 | | Release date: | 2018-07-18 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (1.49 Å) | | Cite: | Investigation of how gate residues in the main channel affect the catalytic activity of Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 80, 2024
|
|
5XY4
 
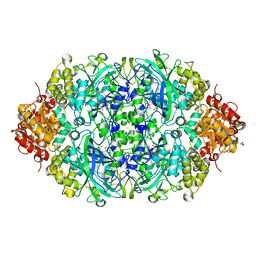 | | CATPO mutant - V536W | | Descriptor: | CALCIUM ION, CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, Catalase, ... | | Authors: | Yuzugullu Karakus, Y, Goc, G, Balci, S, Pearson, A.R, Yorke, B. | | Deposit date: | 2017-07-06 | | Release date: | 2018-07-18 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Identification of the site of oxidase substrate binding in Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
5ZZ1
 
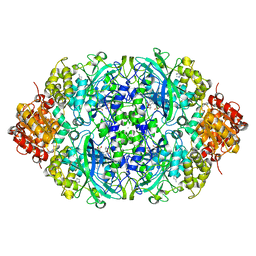 | | Probing the active center of catalase-phenol oxidase from Scytalidium thermophilum | | Descriptor: | 3-AMINO-1,2,4-TRIAZOLE, CALCIUM ION, CIS-HEME D HYDROXYCHLORIN GAMMA-SPIROLACTONE, ... | | Authors: | Yuzugullu Karakus, Y, Trinh, C.H, Pearson, A.R, Ogel, Z.B, McPherson, M.J. | | Deposit date: | 2018-05-29 | | Release date: | 2019-05-29 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Identification of the site of oxidase substrate binding in Scytalidium thermophilum catalase.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
6H01
 
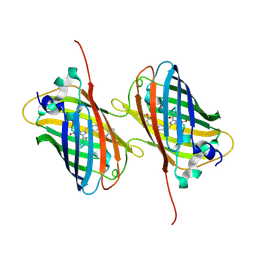 | |
