4WO3
 
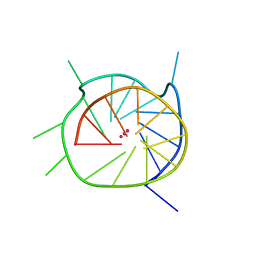 | | THE SECOND C-KIT DNA QUADRUPLEX CRYSTAL STRUCTURE | | Descriptor: | DNA (5'-D(*AP*GP*GP*GP*AP*GP*GP*GP*CP*GP*CP*TP*GP*GP*GP*AP*GP*GP*AP*GP*GP*G)-3'), POTASSIUM ION | | Authors: | Wei, D, Neidle, S. | | Deposit date: | 2014-10-15 | | Release date: | 2014-10-29 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | THE SECOND C-KIT1 DNA QUADRUPLEX CRYSTAL STRUCTURE
TO BE PUBLISHED
|
|
4WO2
 
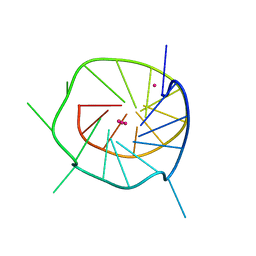 | |
2JTU
 
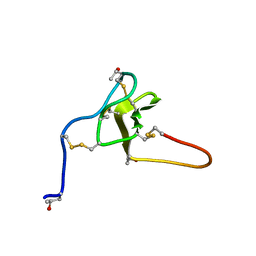 | | NMR structure of iota-RXIA(38) | | Descriptor: | I-superfamily conotoxin r11a | | Authors: | Wei, D, Norton, R. | | Deposit date: | 2007-08-06 | | Release date: | 2008-08-19 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | NMR structure of iota-RXIA(38)
To be Published
|
|
3QXR
 
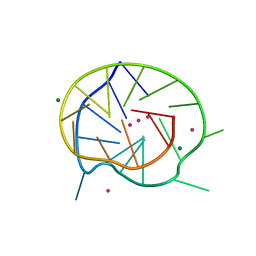 | |
4H29
 
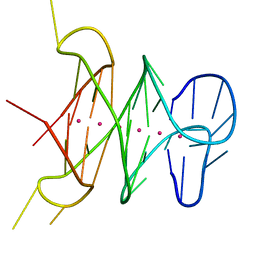 | | B-raf dimer DNA quadruplex | | Descriptor: | DNA (5'-D(*GP*GP*GP*CP*GP*GP*GP*GP*AP*GP*GP*GP*GP*GP*AP*AP*GP*GP*GP*A)-3'), POTASSIUM ION | | Authors: | Wei, D, Parkinson, G, Neidle, S. | | Deposit date: | 2012-09-12 | | Release date: | 2013-07-31 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.991 Å) | | Cite: | Crystal Structure of a Promoter Sequence in the B-raf Gene Reveals an Intertwined Dimer Quadruplex.
J.Am.Chem.Soc., 135, 2013
|
|
1A6X
 
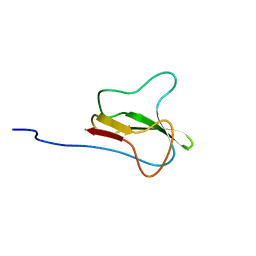 | | STRUCTURE OF THE APO-BIOTIN CARBOXYL CARRIER PROTEIN (APO-BCCP87) OF ESCHERICHIA COLI ACETYL-COA CARBOXYLASE, NMR, 49 STRUCTURES | | Descriptor: | APO-BIOTIN CARBOXYL CARRIER PROTEIN OF ACETYL-COA CARBOXYLASE | | Authors: | Yao, X, Wei, D, Soden Junior, C, Summers, M.F, Beckett, D. | | Deposit date: | 1998-03-04 | | Release date: | 1998-10-14 | | Last modified: | 2022-02-16 | | Method: | SOLUTION NMR | | Cite: | Structure of the carboxy-terminal fragment of the apo-biotin carboxyl carrier subunit of Escherichia coli acetyl-CoA carboxylase.
Biochemistry, 36, 1997
|
|
1WM7
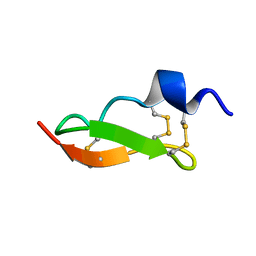 
 | | Solution Structure of BmP01 from the Venom of Scorpion Buthus martensii Karsch, 9 structures | | Descriptor: | Neurotoxin BmP01 | | Authors: | Wu, G, Li, Y, Wei, D, He, F, Jiang, S, Hu, G, Wu, H, Chen, X. | | Deposit date: | 2004-07-05 | | Release date: | 2004-07-27 | | Last modified: | 2022-03-02 | | Method: | SOLUTION NMR | | Cite: | Solution Structure of BmP01 from the Venom of Scorpion Buthus martensii Karsch
Biochem.Biophys.Res.Commun., 276, 2000
|
|
2JRY
 
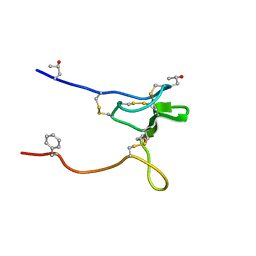 | | Structure and Sodium Channel Activity of an Excitatory I1-Superfamily Conotoxin | | Descriptor: | I-superfamily conotoxin r11a | | Authors: | Buczek, O, Wei, D, Babon, J, Yang, X, Fiedler, B, Chen, P, Yoshikami, D, Olivera, B, Bulaj, G, Norton, R. | | Deposit date: | 2007-06-29 | | Release date: | 2007-10-23 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structure and sodium channel activity of an excitatory I1-superfamily conotoxin.
Biochemistry, 46, 2007
|
|
4EHR
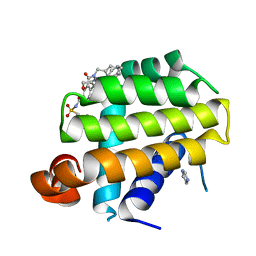 
 | | Crystal structure of Bcl-Xl complex with 4-(5-butyl-3-(hydroxymethyl)-1-phenyl-1h-pyrazol-4-yl)-3-(3,4-dihydro-2(1h)-isoquinolinylcarbonyl)-n-((2-(trimethylsilyl)ethyl)sulfonyl)benzamide | | Descriptor: | 4-[5-butyl-3-(hydroxymethyl)-1-phenyl-1H-pyrazol-4-yl]-3-(3,4-dihydroisoquinolin-2(1H)-ylcarbonyl)-N-{[2-(trimethylsilyl)ethyl]sulfonyl}benzamide, Bcl-2-like protein 1, IMIDAZOLE | | Authors: | Schroeder, G.M, Wei, D, Banfi, P, Cai, Z, Lippy, J, Menichincheri, M, Modugno, M, Naglich, J, Penhallow, B, Perez, H.L, Sack, J, Schmidt, R.J, Tebben, A, Yan, C, Zhang, L, Galvani, A, Lombardo, L.J, Borzilleri, R.M. | | Deposit date: | 2012-04-03 | | Release date: | 2012-06-06 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Pyrazole and pyrimidine phenylacylsulfonamides as dual Bcl-2/Bcl-xL antagonists.
Bioorg.Med.Chem.Lett., 22, 2012
|
|
3VZ1
 
 | |
3VZ2
 
 | |
3VZ3
 
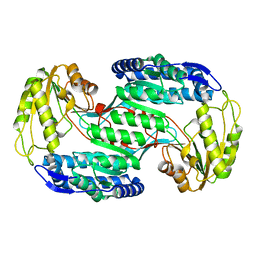
 | | Structural insights into substrate and cofactor selection by sp2771 | | Descriptor: | 4-oxobutanoic acid, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, Succinate-semialdehyde dehydrogenase | | Authors: | Yuan, Y.A, Yuan, Z, Yin, B, Wei, D. | | Deposit date: | 2012-10-09 | | Release date: | 2013-07-10 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structural basis for cofactor and substrate selection by cyanobacterium succinic semialdehyde dehydrogenase
J.Struct.Biol., 182, 2013
|
|
3VZ0
 
 | | Structural insights into cofactor and substrate selection by Gox0499 | | Descriptor: | NONAETHYLENE GLYCOL, Putative NAD-dependent aldehyde dehydrogenase | | Authors: | Yuan, Y.A, Yuan, Z, Yin, B, Wei, D. | | Deposit date: | 2012-10-09 | | Release date: | 2013-07-10 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural basis for cofactor and substrate selection by cyanobacterium succinic semialdehyde dehydrogenase
J.Struct.Biol., 182, 2013
|
|
7YPN
 
 | | Crystal structure of transaminase CC1012 mutant M9 complexed with PLP | | Descriptor: | 1,2-ETHANEDIOL, Aspartate aminotransferase family protein, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Yang, L, Wang, H, Wei, D. | | Deposit date: | 2022-08-03 | | Release date: | 2023-05-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.049 Å) | | Cite: | Mechanism-Guided Computational Design of omega-Transaminase by Reprograming of High-Energy-Barrier Steps.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
7YPM
 
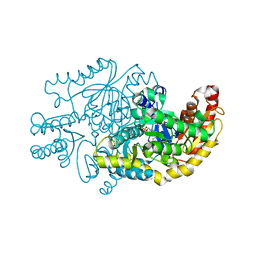 | | Crystal structure of transaminase CC1012 complexed with PLP and L-alanine | | Descriptor: | 1,2-ETHANEDIOL, ALANINE, Aspartate aminotransferase family protein, ... | | Authors: | Yang, L, Wang, H, Wei, D. | | Deposit date: | 2022-08-03 | | Release date: | 2023-05-24 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.984 Å) | | Cite: | Mechanism-Guided Computational Design of omega-Transaminase by Reprograming of High-Energy-Barrier Steps.
Angew.Chem.Int.Ed.Engl., 61, 2022
|
|
3U0U
 
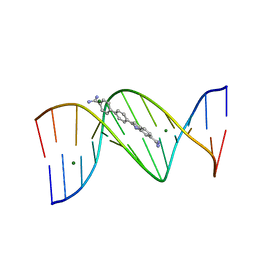 | |
3U2N
 
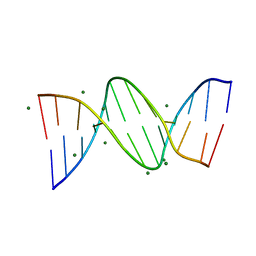 | | Crystal structure of DNA(CGCGAATTCGCG)2 at 1.25 angstroms | | Descriptor: | 5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3', MAGNESIUM ION | | Authors: | Wei, D.G, Neidle, S. | | Deposit date: | 2011-10-04 | | Release date: | 2013-04-17 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Small-molecule binding to the DNA minor groove is mediated by a conserved water cluster.
J.Am.Chem.Soc., 135, 2013
|
|
3U08
 
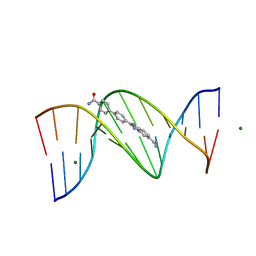 | | Crystal structure of DB1963-D(CGCGAATTCGCG)2 complex at 1.25 A resolution | | Descriptor: | 4'-(5-carbamimidoyl-1H-benzimidazol-2-yl)biphenyl-4-carboxamide, 5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3', MAGNESIUM ION | | Authors: | Wei, D.G, Neidle, S. | | Deposit date: | 2011-09-28 | | Release date: | 2012-09-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Small-molecule binding to the DNA minor groove is mediated by a conserved water cluster.
J.Am.Chem.Soc., 135, 2013
|
|
3U05
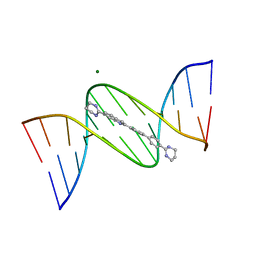 
 | | Crystal structure of DB1804-D(CGCGAATTCGCG)2 complex | | Descriptor: | 5'-D(*CP*GP*CP*GP*AP*AP*TP*TP*CP*GP*CP*G)-3', 6-(1,4,5,6-tetrahydropyrimidin-2-yl)-2-[4'-(1,4,5,6-tetrahydropyrimidin-2-yl)biphenyl-4-yl]-1H-indole, MAGNESIUM ION | | Authors: | Wei, D.G, Neidle, S. | | Deposit date: | 2011-09-28 | | Release date: | 2012-09-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.27 Å) | | Cite: | Small-molecule binding to the DNA minor groove is mediated by a conserved water cluster.
J.Am.Chem.Soc., 135, 2013
|
|
2P4L
 
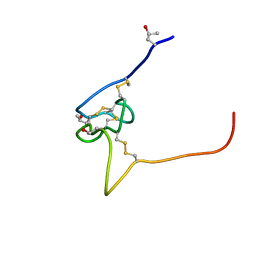 | | Structure and sodium channel activity of an excitatory I1-superfamily conotoxin | | Descriptor: | I-superfamily conotoxin r11a | | Authors: | Buczek, O, Wei, D.X, Babon, J.J, Yang, X.D, Fiedler, B, Yoshikami, D, Olivera, B.M, Bulaj, G, Norton, R.S. | | Deposit date: | 2007-03-12 | | Release date: | 2007-09-25 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Structure and Sodium Channel Activity of an Excitatory I(1)-Superfamily Conotoxin
Biochemistry, 46, 2007
|
|
5IP0
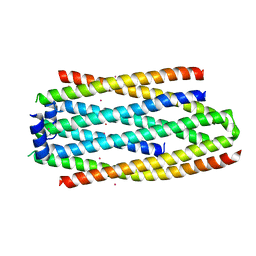 
 | | PHA Binding Protein PhaP (Phasin) | | Descriptor: | CADMIUM ION, PHA granule-associated protein | | Authors: | Chen, G.Q, Wang, X.Q, Zhao, H.Y. | | Deposit date: | 2016-03-09 | | Release date: | 2017-01-25 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3 Å) | | Cite: | Structural Insights on PHA Binding Protein PhaP from Aeromonas hydrophila
Sci Rep, 6, 2016
|
|
6TQI
 
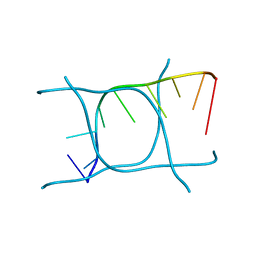 | | I-MOTIF STRUCTURE FORMED FROM THE C STRAND OF A HUMAN TELOMERE FRAGMENT | | Descriptor: | DNA (5'-*TP*AP*AP*CP*CP*CP*TP*AP*A-3') | | Authors: | Parkinson, G.N, Wagner, A, Viladoms-Claverol, J, Duman, R, El-Omari, K. | | Deposit date: | 2019-12-16 | | Release date: | 2020-06-10 | | Last modified: | 2020-11-11 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Native de novo structural determinations of non-canonical nucleic acid motifs by X-ray crystallography at long wavelengths.
Nucleic Acids Res., 48, 2020
|
|
3CE3
 
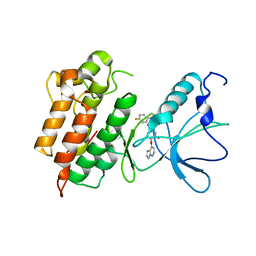 | |
3AWD
 
 | | Crystal structure of gox2181 | | Descriptor: | CADMIUM ION, MAGNESIUM ION, Putative polyol dehydrogenase | | Authors: | Adam Yuan, Y, Yuan, Z. | | Deposit date: | 2011-03-18 | | Release date: | 2012-02-01 | | Last modified: | 2024-03-13 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Biochemical and structural analysis of Gox2181, a new member of the SDR superfamily from Gluconobacter oxydans.
Biochem.Biophys.Res.Commun., 415, 2011
|
|
3WJ7
 
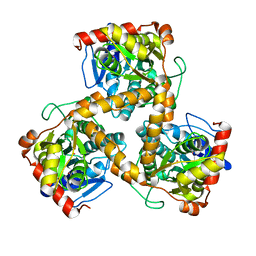 | | Crystal structure of gox2253 | | Descriptor: | MERCURY (II) ION, Putative oxidoreductase | | Authors: | Yuan, Y.A, Yin, B. | | Deposit date: | 2013-10-05 | | Release date: | 2014-06-04 | | Last modified: | 2022-08-24 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural insights into substrate and coenzyme preference by SDR family protein Gox2253 from Gluconobater oxydans.
Proteins, 82, 2014
|
|
