1CE4
 
 | |
1QGM
 | |
1I8Y
 | |
1I8X
 | |
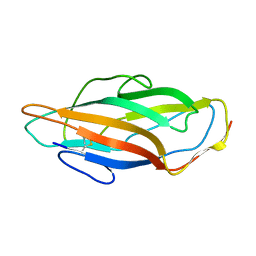
3ZPD
 | |
3ZBE
 
 | |
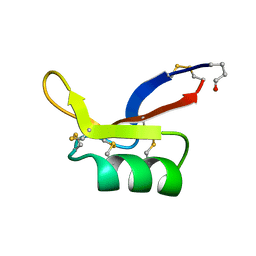
1AYJ
 | |
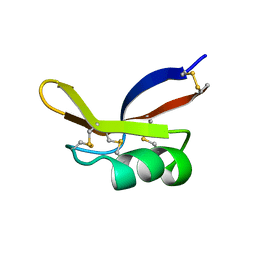
1BK8
 | |
4LOV
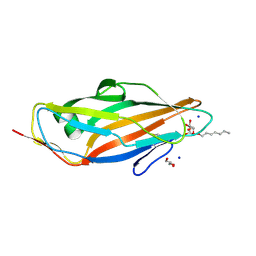 
 | | Crystal structure of FimH in complex with Heptylmannoside | | Descriptor: | GLYCEROL, Protein FimH, SODIUM ION, ... | | Authors: | Garcia-Pino, A, Vanwetswinkel, S, van Nuland, N. | | Deposit date: | 2013-07-13 | | Release date: | 2014-02-12 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Study of the Structural and Dynamic Effects in the FimH Adhesin upon alpha-d-Heptyl Mannose Binding.
J.Med.Chem., 57, 2014
|
|
