2XV6
 | | Crystal structure of the HIV-1 capsid protein C-terminal domain (146- 220) in complex with a camelid VHH. | | Descriptor: | CAMELID VHH 9, CAPSID PROTEIN P24 | | Authors: | Igonet, S, Vaney, M.C, Bartonova, V, Helma, J, Rothbauer, U, Leonhardt, H, Stura, E, Krausslich, H.-G, Rey, F.A. | | Deposit date: | 2010-10-22 | | Release date: | 2011-10-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Targeting HIV-1 Virion Formation with Nanobodies -Implications for the Design of Assembly Inhibitors
To be Published
|
|
2XXM
 
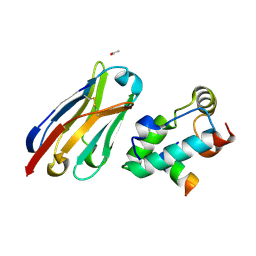 | | Crystal structure of the HIV-1 capsid protein C-terminal domain in complex with a camelid VHH and the CAI peptide. | | Descriptor: | ACETATE ION, CAMELID VHH 9, CAPSID PROTEIN P24, ... | | Authors: | Igonet, S, Vaney, M.C, Bartonova, V, Helma, J, Rothbauer, U, Leonhardt, H, Stura, E, Krausslich, H.-G, Rey, F.A. | | Deposit date: | 2010-11-10 | | Release date: | 2011-10-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Targeting HIV-1 Virion Formation with Nanobodies -Implications for the Design of Assembly Inhibitors
To be Published
|
|
2XT1
 
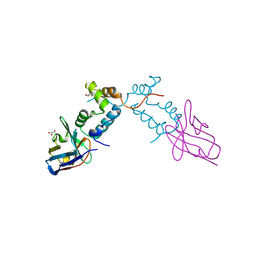 | | Crystal structure of the HIV-1 capsid protein C-terminal domain (146- 231) in complex with a camelid VHH. | | Descriptor: | CAMELID VHH 9, GAG POLYPROTEIN, GLYCEROL | | Authors: | Igonet, S, Vaney, M.C, Bartonova, V, Helma, J, Rothbauer, U, Leonhardt, H, Stura, E, Krausslich, H.-G, Rey, F.A. | | Deposit date: | 2010-10-03 | | Release date: | 2011-10-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.32 Å) | | Cite: | Targeting HIV-1 Virion Formation with Nanobodies -Implications for the Design of Assembly Inhibitors
To be Published
|
|
2XXC
 
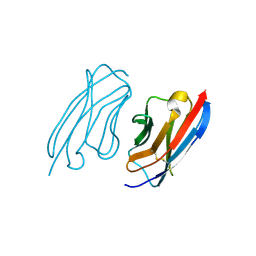 | | Crystal structure of a camelid VHH raised against the HIV-1 capsid protein C-terminal domain. | | Descriptor: | CAMELID VHH 9 | | Authors: | Igonet, S, Vaney, M.C, Bartonova, V, Helma, J, Rothbauer, U, Leonhardt, H, Stura, E, Krausslich, H.-G, Rey, F.A. | | Deposit date: | 2010-11-10 | | Release date: | 2011-10-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.67 Å) | | Cite: | Targeting HIV-1 Virion Formation with Nanobodies -Implications for the Design of Assembly Inhibitors
To be Published
|
|
2HEO
 
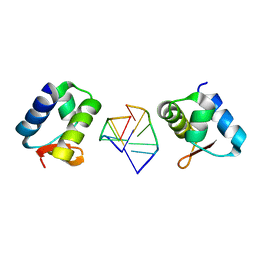 | | General Structure-Based Approach to the Design of Protein Ligands: Application to the Design of Kv1.2 Potassium Channel Blockers. | | Descriptor: | 5'-D(*TP*CP*GP*CP*GP*CP*G)-3', Z-DNA binding protein 1 | | Authors: | Magis, C, Gasparini, S, Charbonnier, J.B, Stura, E, Le Du, M.H, Menez, A, Cuniasse, P. | | Deposit date: | 2006-06-21 | | Release date: | 2006-11-21 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Structure-based secondary structure-independent approach to design protein ligands: Application to the design of Kv1.2 potassium channel blockers.
J.Am.Chem.Soc., 128, 2006
|
|
1DT1
 
 | | THERMUS THERMOPHILUS CYTOCHROME C552 SYNTHESIZED BY ESCHERICHIA COLI | | Descriptor: | CYTOCHROME C552, HEME C | | Authors: | Fee, J.A, Chen, Y, Hill, M.J, Gomez-Moran, E, Loehr, T, Ai, J, Thony-Meyer, L, Williams, P.A, Stura, E, Sridhar, V, McRee, D.E. | | Deposit date: | 2000-01-10 | | Release date: | 2000-02-18 | | Last modified: | 2021-03-03 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Integrity of thermus thermophilus cytochrome c552 synthesized by Escherichia coli cells expressing the host-specific cytochrome c maturation genes, ccmABCDEFGH: biochemical, spectral, and structural characterization of the recombinant protein.
Protein Sci., 9, 2000
|
|
2AS8
 
 | | Crystal structure of mature and fully active Der p 1 allergen | | Descriptor: | MAGNESIUM ION, Major mite fecal allergen Der p 1 | | Authors: | de Halleux, S, Stura, E, VanderElst, L, Carlier, V, Jacquemin, M, Saint-Remy, J.-M. | | Deposit date: | 2005-08-23 | | Release date: | 2005-12-06 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Three-dimensional structure and IgE-binding properties of mature fully active Der p 1, a clinically relevant major allergen
J.Allergy Clin.Immunol., 117, 2006
|
|
8AJ7
 
 | | Kunitz domain of Amblyomin-X | | Descriptor: | 1,2-ETHANEDIOL, Kunitz domain of Amblyomin-X | | Authors: | Ciccone, L, Servent, D, Stura, E.A. | | Deposit date: | 2022-07-27 | | Release date: | 2023-02-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Structural and functional properties of the Kunitz-type and C-terminal domains of Amblyomin-X supporting its antitumor activity.
Front Mol Biosci, 10, 2023
|
|
3F58
 
 | | IGG1 FAB FRAGMENT (58.2) COMPLEX WITH 12-RESIDUE CYCLIC PEPTIDE (INCLUDING RESIDUES 315-324 OF HIV-1 GP120 (MN ISOLATE); H315S MUTATION | | Descriptor: | PROTEIN (CYCLIC PEPTIDE (GP120)), PROTEIN (IMMUNOGLOBULIN GAMMA I (58.2)) | | Authors: | Stanfield, R.L, Cabezas, E, Satterthwait, A.C, Stura, E.A, Profy, A.T, Wilson, I.A. | | Deposit date: | 1998-10-23 | | Release date: | 1999-02-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Dual conformations for the HIV-1 gp120 V3 loop in complexes with different neutralizing fabs.
Structure Fold.Des., 7, 1999
|
|
6R5M
 | | Crystal structure of toxin MT9 from mamba venom | | Descriptor: | ACETYL GROUP, Dendroaspis polylepis MT9, GLYCEROL, ... | | Authors: | Stura, E.A, Tepshi, L, Ciolek, J, Triquigneaux, M, Zoukimian, C, De Waard, M, Beroud, R, Servent, D, Gilles, N, Legrand, P, Ciccone, L. | | Deposit date: | 2019-03-25 | | Release date: | 2020-02-12 | | Last modified: | 2022-05-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | MT9, a natural peptide from black mamba venom antagonizes the muscarinic type 2 receptor and reverses the M2R-agonist-induced relaxation in rat and human arteries
Biomed Pharmacother, 150, 2022
|
|
3LJG
 
 | | Human MMP12 in complex with non-zinc chelating inhibitor | | Descriptor: | ACETOHYDROXAMIC ACID, CALCIUM ION, HEXAETHYLENE GLYCOL, ... | | Authors: | Stura, E.A, Dive, V, Devel, L, Vera, L, Beau, F. | | Deposit date: | 2010-01-26 | | Release date: | 2010-09-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.313 Å) | | Cite: | Insights from selective non-phosphinic inhibitors of MMP-12 tailored to fit with an S1' loop canonical conformation.
J.Biol.Chem., 285, 2010
|
|
3LIR
 
 | | Human MMP12 in complex with non-zinc chelating inhibitor | | Descriptor: | CALCIUM ION, GLYCINE, Macrophage metalloelastase, ... | | Authors: | Stura, E.A, Dive, V, Devel, L, Czarny, B, Vera, L, Beau, F. | | Deposit date: | 2010-01-25 | | Release date: | 2010-09-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Insights from selective non-phosphinic inhibitors of MMP-12 tailored to fit with an S1' loop canonical conformation.
J.Biol.Chem., 285, 2010
|
|
3LIL
 
 | | Human MMP12 in complex with non-zinc chelating inhibitor | | Descriptor: | ACETOHYDROXAMIC ACID, CALCIUM ION, Macrophage metalloelastase, ... | | Authors: | Stura, E.A, Dive, V, Devel, L, Czarny, B, Vera, L, Beau, F. | | Deposit date: | 2010-01-25 | | Release date: | 2010-09-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Insights from selective non-phosphinic inhibitors of MMP-12 tailored to fit with an S1' loop canonical conformation.
J.Biol.Chem., 285, 2010
|
|
3LIK
 
 | | Human MMP12 in complex with non-zinc chelating inhibitor | | Descriptor: | ACETOHYDROXAMIC ACID, CALCIUM ION, HEXAETHYLENE GLYCOL, ... | | Authors: | Stura, E.A, Dive, V, Devel, L, Czarny, B, Vera, L, Beau, F. | | Deposit date: | 2010-01-25 | | Release date: | 2010-09-01 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Insights from selective non-phosphinic inhibitors of MMP-12 tailored to fit with an S1' loop canonical conformation.
J.Biol.Chem., 285, 2010
|
|
2F58
 
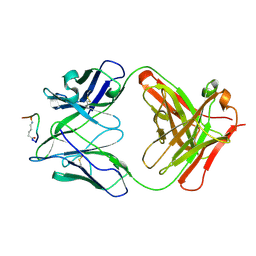 | | IGG1 FAB FRAGMENT (58.2) COMPLEX WITH 12-RESIDUE CYCLIC PEPTIDE (INCLUDING RESIDUES 315-324 OF HIV-1 GP120) (MN ISOLATE) | | Descriptor: | 1-IMINO-5-PENTANONE, PROTEIN (HIV-1 GP120), PROTEIN (IGG1 FAB 58.2 ANTIBODY (HEAVY CHAIN)), ... | | Authors: | Stanfield, R.L, Cabezas, E, Satterthwait, A.C, Stura, E.A, Profy, A.T, Wilson, I.A. | | Deposit date: | 1998-10-23 | | Release date: | 1999-02-09 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Dual conformations for the HIV-1 gp120 V3 loop in complexes with different neutralizing fabs.
Structure Fold.Des., 7, 1999
|
|
2VLW
 
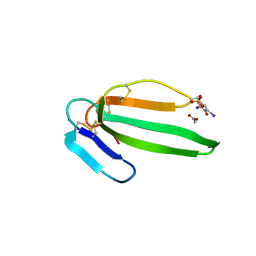 | | Crystal structure of the muscarinic toxin MT7 diiodoTYR51 derivative. | | Descriptor: | ACETATE ION, MUSCARINIC M1-TOXIN1, SULFATE ION | | Authors: | Menez, R, Granata, V, Mourier, G, Fruchart-Gaillard, C, Menez, A, Servant, D, Stura, E.A. | | Deposit date: | 2008-01-17 | | Release date: | 2008-10-14 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (1.39 Å) | | Cite: | Different Interactions between Mt7 Toxin and the Human Muscarinic M1 Receptor in its Free and N-Methylscopolamine-Occupied States.
Mol.Pharmacol., 74, 2008
|
|
2ZCK
 
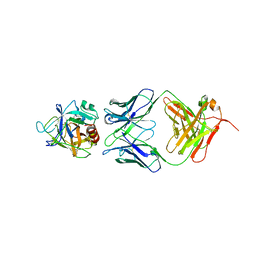 | | Crystal structure of a ternary complex between PSA, a substrat-acyl intermediate and an activating antibody | | Descriptor: | KGISSQY, Prostate-specific antigen, alpha-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Menez, R, Stura, E, Jolivet-Reynaud, C. | | Deposit date: | 2007-11-09 | | Release date: | 2008-01-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Crystal structure of a ternary complex between human prostate-specific antigen, its substrate acyl intermediate and an activating antibody
J.Mol.Biol., 376, 2008
|
|
2ZCH
 
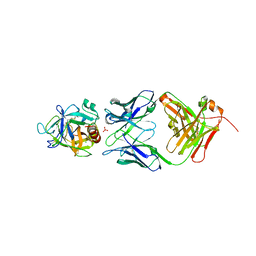 | | Crystal structure of human prostate specific antigen complexed with an activating antibody | | Descriptor: | 2-acetamido-2-deoxy-alpha-D-glucopyranose, CHLORIDE ION, PHOSPHATE ION, ... | | Authors: | Menez, R, Stura, E, Jolivet-Reynaud, C. | | Deposit date: | 2007-11-08 | | Release date: | 2008-01-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.83 Å) | | Cite: | Crystal structure of a ternary complex between human prostate-specific antigen, its substrate acyl intermediate and an activating antibody
J.Mol.Biol., 376, 2008
|
|
2ZCL
 
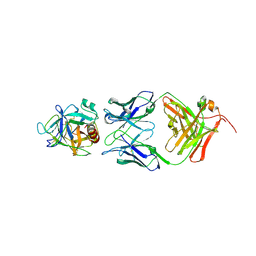 | | Crystal structure of human prostate specific antigen complexed with an activating antibody | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Prostate-specific antigen, monoclonal antibody 8G8F5 Fab | | Authors: | Menez, R, Stura, E, Jolivet-Reynaud, C. | | Deposit date: | 2007-11-09 | | Release date: | 2008-01-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (3.25 Å) | | Cite: | Crystal structure of a ternary complex between human prostate-specific antigen, its substrate acyl intermediate and an activating antibody
J.Mol.Biol., 376, 2008
|
|
3NDS
 | | Crystal structure of engineered Naja Nigricollis toxin alpha | | Descriptor: | (4S)-2-METHYL-2,4-PENTANEDIOL, SULFATE ION, Short neurotoxin 1 | | Authors: | Stura, E.A, Drevet, P, Gregory, G, Gallopin, M, Vandame, M. | | Deposit date: | 2010-06-08 | | Release date: | 2010-08-11 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.2 Å) | | Cite: | Conformational exchange is critical for the productivity of an oxidative folding intermediate with buried free cysteines.
J.Mol.Biol., 403, 2010
|
|
2H5F
 
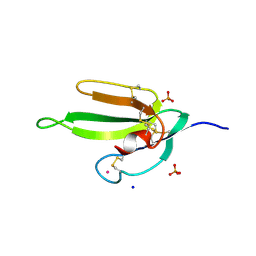 | | Denmotoxin: A the three-finger toxin from colubrid snake Boiga dendrophila with bird-specific activity | | Descriptor: | PHOSPHATE ION, POTASSIUM ION, SODIUM ION, ... | | Authors: | Pawlak, J, Kini, R.M, Stura, E.A. | | Deposit date: | 2006-05-26 | | Release date: | 2006-08-29 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Denmotoxin, a three-finger toxin from the colubrid snake Boiga dendrophila (Mangrove Catsnake) with bird-specific activity.
J.Biol.Chem., 281, 2006
|
|
1F58
 
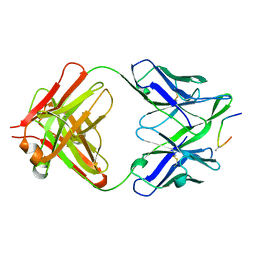 | | IGG1 FAB FRAGMENT (58.2) COMPLEX WITH 24-RESIDUE PEPTIDE (RESIDUES 308-333 OF HIV-1 GP120 (MN ISOLATE) WITH ALA TO AIB SUBSTITUTION AT POSITION 323 | | Descriptor: | Envelope glycoprotein gp120, PROTEIN (IGG1 ANTIBODY 58.2 (HEAVY CHAIN)), PROTEIN (IGG1 ANTIBODY 58.2 (LIGHT CHAIN)) | | Authors: | Stanfield, R.L, Cabezas, E, Satterthwait, A.C, Stura, E.A, Profy, A.T, Wilson, I.A. | | Deposit date: | 1998-10-21 | | Release date: | 1999-02-02 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Dual conformations for the HIV-1 gp120 V3 loop in complexes with different neutralizing fabs.
Structure Fold.Des., 7, 1999
|
|
1N64
 
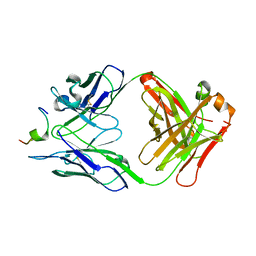 | | Crystal structure analysis of the immunodominant antigenic site on Hepatitis C virus protein bound to mAb 19D9D6 | | Descriptor: | Fab 19D9D6 heavy chain, Fab 19D9D6 light chain, Genome polyprotein Capsid protein C | | Authors: | Menez, R, Bossus, M, Muller, B, Sibai, G, Dalbon, P, Ducancel, F, Jolivet-Reynaud, C, Stura, E. | | Deposit date: | 2002-11-08 | | Release date: | 2003-02-25 | | Last modified: | 2013-09-18 | | Method: | X-RAY DIFFRACTION (2.34 Å) | | Cite: | Crystal structure of a hydrophobic immunodominant
antigenic site on hepatitis C virus core protein
complexed to monoclonal antibody 19D9D6.
J.Immunol., 170, 2003
|
|
1NLB
 
 | | crystal structure of anti-HCV monoclonal antibody 19D9D6 | | Descriptor: | antibody 19D9D6 heavy chain, antibody 19D9D6 light chain | | Authors: | Menez, R, Bossus, M, Muller, B, Sibai, G, Dalbon, P, Ducancel, F, Jolivet-Reynaud, C, Stura, E. | | Deposit date: | 2003-01-07 | | Release date: | 2003-02-25 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Crystal structure of a hydrophobic immunodominant
antigenic site on hepatitis C virus core protein
complexed to monoclonal antibody 19D9D6.
J.Immunol., 170, 2003
|
|
2H7Z
 
 | | Crystal structure of irditoxin | | Descriptor: | Irditoxin subunit A, Irditoxin subunit B | | Authors: | Pawlak, J, Kini, R.M, Stura, E.A, Le Du, M.H. | | Deposit date: | 2006-06-06 | | Release date: | 2006-08-29 | | Last modified: | 2011-10-05 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Irditoxin, a novel covalently linked heterodimeric three-finger toxin with high taxon-specific neurotoxicity.
Faseb J., 23, 2009
|
|
